

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
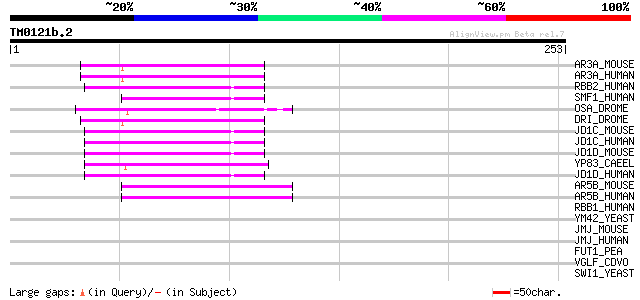
AR3A_MOUSE (Q62431) AT-rich interactive domain-containing protei... 59 9e-09
AR3A_HUMAN (Q99856) AT-rich interactive domain-containing protei... 59 9e-09
RBB2_HUMAN (P29375) Retinoblastoma-binding protein 2 (RBBP-2) 57 3e-08
SMF1_HUMAN (O14497) SWI/SNF-related, matrix-associated, actin-de... 54 5e-07
OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein) 53 8e-07
DRI_DROME (Q24573) Dead ringer protein (Retained protein) 48 2e-05
JD1C_MOUSE (P41230) Jumonji/ARID domain-containing protein 1C (S... 47 3e-05
JD1C_HUMAN (P41229) Jumonji/ARID domain-containing protein 1C (S... 47 3e-05
JD1D_MOUSE (Q62240) Jumonji/ARID domain-containing protein 1D (S... 47 4e-05
YP83_CAEEL (Q09441) Hypothetical protein C08B11.3 in chromosome II 44 3e-04
JD1D_HUMAN (Q9BY66) Jumonji/ARID domain-containing protein 1D (S... 44 3e-04
AR5B_MOUSE (Q8BM75) AT-rich interactive domain-containing protei... 44 3e-04
AR5B_HUMAN (Q14865) AT-rich interactive domain-containing protei... 44 3e-04
RBB1_HUMAN (P29374) Retinoblastoma-binding protein 1 (RBBP-1) 37 0.035
YM42_YEAST (Q03214) Hypothetical 162.7 kDa protein in SIP18-SPT2... 36 0.10
JMJ_MOUSE (Q62315) Jumonji protein (Jumonji/ARID domain-containi... 35 0.13
JMJ_HUMAN (Q92833) Jumonji protein (Jumonji/ARID domain-containi... 35 0.17
FUT1_PEA (Q9M5Q1) Galactoside 2-alpha-L-fucosyltransferase (EC 2... 35 0.23
VGLF_CDVO (P12569) Fusion glycoprotein precursor [Contains: Fusi... 33 0.66
SWI1_YEAST (P09547) Transcription regulatory protein SWI1 (SWI/S... 32 1.5
>AR3A_MOUSE (Q62431) AT-rich interactive domain-containing protein
3A (ARID domain-containing protein 3A) (Dead ringer
like-1 protein) (B-cell regulator of IgH transcription)
(Bright)
Length = 601
Score = 59.3 bits (142), Expect = 9e-09
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 247 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 306
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 307 NLPTSITSAAFTLRTQYMKYLYPYE 331
>AR3A_HUMAN (Q99856) AT-rich interactive domain-containing protein
3A (ARID domain-containing protein 3A) (B-cell regulator
of IgH transcription) (Bright) (E2F binding protein 1)
Length = 593
Score = 59.3 bits (142), Expect = 9e-09
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMKYLYPYE 326
>RBB2_HUMAN (P29375) Retinoblastoma-binding protein 2 (RBBP-2)
Length = 1722
Score = 57.4 bits (137), Expect = 3e-08
Identities = 33/82 (40%), Positives = 45/82 (54%), Gaps = 1/82 (1%)
Query: 35 FLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS 94
FLD L +F GS IPV+ + LDL+ L V + G+E V EKKW +VGS +
Sbjct: 90 FLDQLAKFWELQGSTLKIPVVERKILDLYALSKIVASKGGFEMVTKEKKWSKVGSRLGYL 149
Query: 95 TTTTSASYVLKKHYWNLLYNFE 116
+ S +LK HY +LY +E
Sbjct: 150 PGKGTGS-LLKSHYERILYPYE 170
>SMF1_HUMAN (O14497) SWI/SNF-related, matrix-associated,
actin-dependent regulator of chromatin subfamily F
member 1 (SWI-SNF complex protein p270) (B120)
Length = 1902
Score = 53.5 bits (127), Expect = 5e-07
Identities = 28/65 (43%), Positives = 39/65 (59%), Gaps = 1/65 (1%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNL 111
+P +G + LDL+ LYV V G +V KKWRE+ + N T++++AS LKK Y
Sbjct: 658 LPAVGRKPLDLYRLYVSVKEIGGLTQVNKNKKWRELATNLNVGTSSSAAS-SLKKQYIQC 716
Query: 112 LYNFE 116
LY FE
Sbjct: 717 LYAFE 721
>OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein)
Length = 2716
Score = 52.8 bits (125), Expect = 8e-07
Identities = 34/100 (34%), Positives = 51/100 (51%), Gaps = 5/100 (5%)
Query: 31 DAKLFLDTLRQFHFHMGSKYMI-PVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGS 89
D + +LD LR F + P I + LDL+ LY+ V R G+ +V K W+++
Sbjct: 1002 DRRGWLDKLRAFMEERRTPITACPTISKQPLDLYRLYIYVKERGGFVEVTKSKTWKDIAG 1061
Query: 90 VFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITP 129
+ ++SA+Y L+KHY L FE HF + G I P
Sbjct: 1062 LLGIG-ASSSAAYTLRKHYTKNLLTFE-CHFDR--GDIDP 1097
>DRI_DROME (Q24573) Dead ringer protein (Retained protein)
Length = 911
Score = 48.1 bits (113), Expect = 2e-05
Identities = 28/85 (32%), Positives = 44/85 (50%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ +P++ LDL+ LY V R G V+ +K W+E+
Sbjct: 297 KEFLDDLFSFMQKRGTPINRLPIMAKSVLDLYELYNLVIARGGLVDVINKKLWQEIIKGL 356
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
+ ++ TSA++ L+ Y LY +E
Sbjct: 357 HLPSSITSAAFTLRTQYMKYLYPYE 381
>JD1C_MOUSE (P41230) Jumonji/ARID domain-containing protein 1C (SmcX
protein) (Xe169 protein)
Length = 1554
Score = 47.4 bits (111), Expect = 3e-05
Identities = 25/82 (30%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 35 FLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS 94
+LD + +F GS IP + R LDL++L V GYE + +++W V N+
Sbjct: 85 YLDQIAKFWEIQGSSLKIPNVERRILDLYSLSKIVVEEGGYETICKDRRWARVAQRLNYP 144
Query: 95 TTTTSASYVLKKHYWNLLYNFE 116
S +L+ HY ++Y +E
Sbjct: 145 PGKNIGS-LLRSHYERIVYPYE 165
>JD1C_HUMAN (P41229) Jumonji/ARID domain-containing protein 1C (SmcX
protein) (Xe169 protein)
Length = 1560
Score = 47.4 bits (111), Expect = 3e-05
Identities = 25/82 (30%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 35 FLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS 94
+LD + +F GS IP + R LDL++L V GYE + +++W V N+
Sbjct: 85 YLDQIAKFWEIQGSSLKIPNVERRILDLYSLSKIVVEEGGYEAICKDRRWARVAQRLNYP 144
Query: 95 TTTTSASYVLKKHYWNLLYNFE 116
S +L+ HY ++Y +E
Sbjct: 145 PGKNIGS-LLRSHYERIVYPYE 165
>JD1D_MOUSE (Q62240) Jumonji/ARID domain-containing protein 1D (SmcY
protein) (Histocompatibility Y antigen) (H-Y)
Length = 1548
Score = 47.0 bits (110), Expect = 4e-05
Identities = 24/82 (29%), Positives = 42/82 (50%), Gaps = 1/82 (1%)
Query: 35 FLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS 94
+LD + +F GS IP + + LDL++L V GYE + +++W V N+
Sbjct: 85 YLDQIAKFWEIQGSSLKIPNVERKILDLYSLNKIVMEEGGYEAICKDRRWARVAQRLNYP 144
Query: 95 TTTTSASYVLKKHYWNLLYNFE 116
+ S +L+ HY ++Y +E
Sbjct: 145 SGKNIGS-LLRSHYERIIYPYE 165
>YP83_CAEEL (Q09441) Hypothetical protein C08B11.3 in chromosome II
Length = 1244
Score = 44.3 bits (103), Expect = 3e-04
Identities = 24/85 (28%), Positives = 42/85 (49%), Gaps = 1/85 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
F ++LR F+ + + +P + G +++L+ LY V G++KV A KW ++ +F
Sbjct: 30 FYNSLRMFYKRRWNATLKLPHVQGVEVNLYRLYDTVMALGGWQKVAASDKWSDIAEMFGC 89
Query: 94 STTTTSASYVLKKHYWNLLYNFEQV 118
+ +K Y L FEQV
Sbjct: 90 KDDILCGDHAIKIIYMRYLSKFEQV 114
>JD1D_HUMAN (Q9BY66) Jumonji/ARID domain-containing protein 1D (SmcY
protein) (Histocompatibility Y antigen) (H-Y)
Length = 1539
Score = 44.3 bits (103), Expect = 3e-04
Identities = 23/82 (28%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 35 FLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS 94
+LD + +F GS IP + + LDL++L V GYE + +++W V ++
Sbjct: 85 YLDQIAKFWEIQGSSLKIPNVERKILDLYSLSKIVIEEGGYEAICKDRRWARVAQRLHYP 144
Query: 95 TTTTSASYVLKKHYWNLLYNFE 116
S +L+ HY ++Y +E
Sbjct: 145 PGKNIGS-LLRSHYERIIYPYE 165
>AR5B_MOUSE (Q8BM75) AT-rich interactive domain-containing protein
5B (ARID domain-containing protein 5B) (Mrf1-like)
(Modulator recognition factor protein 2) (Mrf-2)
(Developmentally and sexually retarded with transient
immune abnormalities protein)
Length = 1188
Score = 44.3 bits (103), Expect = 3e-04
Identities = 19/78 (24%), Positives = 42/78 (53%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNL 111
IP +G ++++L T++ + GYE + A ++W+ + + +TSA+ ++HY L
Sbjct: 343 IPYLGFKQINLWTMFQAAQKLGGYETITARRQWKHIYDELGGNPGSTSAATCTRRHYERL 402
Query: 112 LYNFEQVHFFKVQGPITP 129
+ +E+ + P+ P
Sbjct: 403 ILPYERFIKGEEDKPLPP 420
>AR5B_HUMAN (Q14865) AT-rich interactive domain-containing protein
5B (ARID domain-containing protein 5B) (Mrf1-like)
(Modulator recognition factor 2) (Mrf-2)
Length = 1188
Score = 44.3 bits (103), Expect = 3e-04
Identities = 19/78 (24%), Positives = 42/78 (53%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNL 111
IP +G ++++L T++ + GYE + A ++W+ + + +TSA+ ++HY L
Sbjct: 342 IPYLGFKQINLWTMFQAAQKLGGYETITARRQWKHIYDELGGNPGSTSAATCTRRHYERL 401
Query: 112 LYNFEQVHFFKVQGPITP 129
+ +E+ + P+ P
Sbjct: 402 ILPYERFIKGEEDKPLPP 419
>RBB1_HUMAN (P29374) Retinoblastoma-binding protein 1 (RBBP-1)
Length = 1257
Score = 37.4 bits (85), Expect = 0.035
Identities = 19/65 (29%), Positives = 33/65 (50%)
Query: 53 PVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLL 112
PV+G + L+L L+ V + G + + + W+++ ++ASY LK Y L
Sbjct: 334 PVLGYKDLNLFKLFRLVYHQGGCDNIDSGAVWKQIYMDLGIPILNSAASYNLKTAYRKYL 393
Query: 113 YNFEQ 117
Y FE+
Sbjct: 394 YGFEE 398
>YM42_YEAST (Q03214) Hypothetical 162.7 kDa protein in SIP18-SPT21
intergenic region
Length = 1411
Score = 35.8 bits (81), Expect = 0.10
Identities = 22/66 (33%), Positives = 33/66 (49%), Gaps = 1/66 (1%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS-TTTTSASYVLKKHYWN 110
IP I R LDL+ L V R G+ V +K W ++G +S +S S L+ Y
Sbjct: 210 IPSIDKRTLDLYRLRSCVKLRGGFNAVCEKKLWAQIGRELGYSGRIMSSLSTSLRSAYAK 269
Query: 111 LLYNFE 116
+L +F+
Sbjct: 270 ILLDFD 275
>JMJ_MOUSE (Q62315) Jumonji protein (Jumonji/ARID domain-containing
protein 2)
Length = 1234
Score = 35.4 bits (80), Expect = 0.13
Identities = 26/121 (21%), Positives = 49/121 (40%), Gaps = 21/121 (17%)
Query: 19 PPPLASHEVIVNDAKLFLDTLRQFHF-----------------HMGSKYM----IPVIGG 57
PPP E +ND F+ ++ H H+ S+ + +P+IGG
Sbjct: 589 PPPDWRPECKLNDEMRFVTQIQHIHKLGRRWGPNVQRLACIKKHLRSQGITMDELPLIGG 648
Query: 58 RKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQ 117
+LDL + + G ++V KKW ++ + T L++ Y L +++
Sbjct: 649 CELDLACFFRLINEMGGMQQVTDLKKWNKLADMLRIPKTAQDRLAKLQEAYCQYLLSYDS 708
Query: 118 V 118
+
Sbjct: 709 L 709
>JMJ_HUMAN (Q92833) Jumonji protein (Jumonji/ARID domain-containing
protein 2)
Length = 1266
Score = 35.0 bits (79), Expect = 0.17
Identities = 25/121 (20%), Positives = 49/121 (39%), Gaps = 21/121 (17%)
Query: 19 PPPLASHEVIVNDAKLFLDTLRQFHF-----------------HMGSKYM----IPVIGG 57
PPP E +ND F+ ++ H H+ S+ + +P+IGG
Sbjct: 591 PPPDWRPECKLNDEMRFVTQIQHIHKLGRRWGPNVQRLACIKKHLKSQGITMDELPLIGG 650
Query: 58 RKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQ 117
+LDL + + G ++V KKW ++ + T L++ Y + +++
Sbjct: 651 CELDLACFFRLINEMGGMQQVTELKKWNKLSDMLRIPKTAQERLAKLQEAYCQYILSYDS 710
Query: 118 V 118
+
Sbjct: 711 L 711
>FUT1_PEA (Q9M5Q1) Galactoside 2-alpha-L-fucosyltransferase (EC
2.4.1.69) (Xyloglucan alpha-(1,2)-fucosyltransferase)
(PsFT1)
Length = 565
Score = 34.7 bits (78), Expect = 0.23
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 7/67 (10%)
Query: 170 GSSSSGKPDLALV----EYAPKHANNGPESNVEAKGYPRYGEGRIEEKFDCGYLVSVK-- 223
G SGKP L+ +Y +H GP + K G G+ E DC Y+V +
Sbjct: 126 GKGLSGKPSSYLISRLRKYEARHKQCGPYTESYNKTVKELGSGQFSESVDCKYVVWISFS 185
Query: 224 -LGSEVL 229
LG+ +L
Sbjct: 186 GLGNRIL 192
>VGLF_CDVO (P12569) Fusion glycoprotein precursor [Contains: Fusion
glycoprotein F2; Fusion glycoprotein F1]
Length = 662
Score = 33.1 bits (74), Expect = 0.66
Identities = 23/78 (29%), Positives = 40/78 (50%), Gaps = 4/78 (5%)
Query: 26 EVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWR 85
+V+++ + L+T+R+ F+ GS +P++ L L L RR Y++ + K+
Sbjct: 586 KVLIDSSNQILETVRRSSFNFGSLLSVPILSCTALALLLLIYCCKRR--YQQTL--KQHT 641
Query: 86 EVGSVFNFSTTTTSASYV 103
+V F T TS SYV
Sbjct: 642 KVDPAFKPDLTGTSKSYV 659
>SWI1_YEAST (P09547) Transcription regulatory protein SWI1 (SWI/SNF
complex component SWI1) (Transcription regulatory
protein ADR6) (Regulatory protein GAM3)
Length = 1314
Score = 32.0 bits (71), Expect = 1.5
Identities = 21/74 (28%), Positives = 35/74 (46%), Gaps = 6/74 (8%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNL 111
IP IG RK++L LY+ V + G ++V ++W V S L+ Y+ +
Sbjct: 430 IPEIGNRKINLFYLYMLVQKFGGADQVTRTQQWSMVAQRLQISDYQQ-----LESIYFRI 484
Query: 112 LYNFEQVHFFKVQG 125
L +E+ H +G
Sbjct: 485 LLPYER-HMISQEG 497
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,741,058
Number of Sequences: 164201
Number of extensions: 1394493
Number of successful extensions: 2971
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2947
Number of HSP's gapped (non-prelim): 29
length of query: 253
length of database: 59,974,054
effective HSP length: 108
effective length of query: 145
effective length of database: 42,240,346
effective search space: 6124850170
effective search space used: 6124850170
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0121b.2


