

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
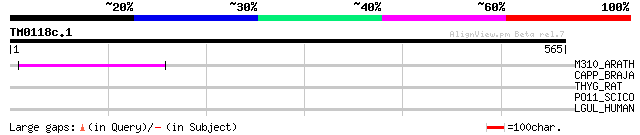
Query= TM0118c.1
(565 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 110 1e-23
CAPP_BRAJA (Q89R17) Phosphoenolpyruvate carboxylase (EC 4.1.1.31... 35 0.40
THYG_RAT (P06882) Thyroglobulin precursor 31 9.9
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 31 9.9
LGUL_HUMAN (Q04760) Lactoylglutathione lyase (EC 4.4.1.5) (Methy... 31 9.9
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 110 bits (274), Expect = 1e-23
Identities = 54/150 (36%), Positives = 85/150 (56%), Gaps = 1/150 (0%)
Query: 10 LAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPK-GCGGIGFR 68
+A+P Y MSCF L LC ++ ++ F+W + R + W+ W LC+ K GG+GFR
Sbjct: 1 MALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFR 60
Query: 69 DFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAW 128
D FN ALLAK +RI+ + ++L+++ ++ YFP S +E G RPSY W S+
Sbjct: 61 DLGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGRE 120
Query: 129 IFQEGGLWRVGNGKNIRA*EDKWVPHGGPL 158
+ G L +G+G + + D+W+ PL
Sbjct: 121 LLSRGLLRTIGDGIHTKVWLDRWIMDETPL 150
>CAPP_BRAJA (Q89R17) Phosphoenolpyruvate carboxylase (EC 4.1.1.31)
(PEPCase) (PEPC)
Length = 932
Score = 35.4 bits (80), Expect = 0.40
Identities = 59/238 (24%), Positives = 96/238 (39%), Gaps = 15/238 (6%)
Query: 262 ADAVPQCCETAWRACLDILPVRSILHRRGVV--GNPCCPRCSEVEETVHHALFSCPLVRM 319
+DA+PQC + + D+L V +L G+V + ET+ S ++
Sbjct: 506 SDAIPQCIISMCKGMSDMLEVAVLLKEVGLVHPSGRSAINIVPLFETIEDLQASSAIMDR 565
Query: 320 IWFASPLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQARNELLFKWKDSSID 379
+ D GSV + L + +D+N G F+ + +++A L+ ++ +
Sbjct: 566 MLSLHDYRRLVDSRGSVQEVMLGY---SDSNKDGGFVTSGWELYKAEINLVDVFERHHVR 622
Query: 380 QLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVTSSKEAGAGMVMRN 439
L R V GP + + + ++G ++ + SSK + A V RN
Sbjct: 623 LRLFHG---RGGSVGRGGGP-SYDAIIAQPGGAVNGQIRITEQGEIISSKYSNA-EVGRN 677
Query: 440 NHGEVLAAATTE---LGPVLSPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTFW 494
N E+LAAAT E L P S E L L +R + ETD Y FW
Sbjct: 678 NL-EILAAATLEASLLHPRQSAPRREYLTAMDELSSLAFRAYRGLVYETDGFVDY-FW 733
>THYG_RAT (P06882) Thyroglobulin precursor
Length = 2768
Score = 30.8 bits (68), Expect = 9.9
Identities = 13/35 (37%), Positives = 16/35 (45%), Gaps = 4/35 (11%)
Query: 182 GCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFF 216
G WD K+ CW P R P P Q +DC +
Sbjct: 2254 GSWDATKLRSSCWQPGTRT----PTPPQISEDCLY 2284
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 30.8 bits (68), Expect = 9.9
Identities = 14/35 (40%), Positives = 17/35 (48%), Gaps = 1/35 (2%)
Query: 282 VRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPL 316
+ LH+RG+ P C C E V H L CPL
Sbjct: 915 MNGFLHKRGLSNTPVC-MCGAPNEDVKHLLGECPL 948
>LGUL_HUMAN (Q04760) Lactoylglutathione lyase (EC 4.4.1.5)
(Methylglyoxalase) (Aldoketomutase) (Glyoxalase I) (Glx
I) (Ketone-aldehyde mutase) (S-D-lactoylglutathione
methylglyoxal lyase)
Length = 183
Score = 30.8 bits (68), Expect = 9.9
Identities = 24/85 (28%), Positives = 31/85 (36%), Gaps = 3/85 (3%)
Query: 468 WALRLATKLGFRRIW-IETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFV 526
WAL L W E D + Y N F H+ V D S F+ FV
Sbjct: 90 WALSRKATLELTHNWGTEDDATQSYHNGNSDPRGFGHIGIAVPDVYSACKRFEELGVKFV 149
Query: 527 RRTSNGAADALARLAFHYGCMIWIE 551
++ +G LA + G WIE
Sbjct: 150 KKPDDGKMKGLAFIQDPDG--YWIE 172
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.139 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,289,549
Number of Sequences: 164201
Number of extensions: 2865843
Number of successful extensions: 6571
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 6568
Number of HSP's gapped (non-prelim): 5
length of query: 565
length of database: 59,974,054
effective HSP length: 115
effective length of query: 450
effective length of database: 41,090,939
effective search space: 18490922550
effective search space used: 18490922550
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0118c.1


