

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.10
(1082 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
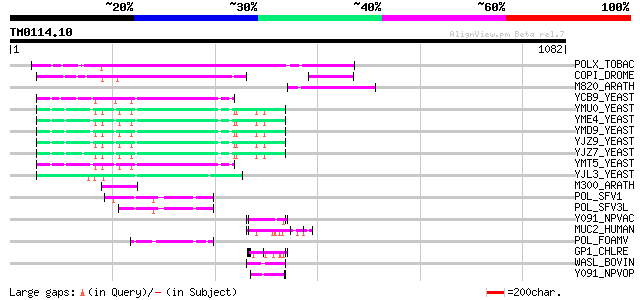
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 270 2e-71
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 186 3e-46
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 127 2e-28
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 109 3e-23
YMU0_YEAST (Q04670) Transposon Ty1 protein B 107 1e-22
YME4_YEAST (Q04711) Transposon Ty1 protein B 107 1e-22
YMD9_YEAST (Q03434) Transposon Ty1 protein B 107 1e-22
YJZ9_YEAST (P47100) Transposon Ty1 protein B 107 1e-22
YJZ7_YEAST (P47098) Transposon Ty1 protein B 105 5e-22
YMT5_YEAST (Q04214) Transposon Ty1 protein B 105 7e-22
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 80 2e-14
M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300... 60 2e-08
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 59 5e-08
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 58 2e-07
Y091_NPVAC (P41479) Hypothetical 24.1 kDa protein in LEF4-P33 in... 53 4e-06
MUC2_HUMAN (Q02817) Mucin 2 precursor (Intestinal mucin 2) 53 5e-06
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 52 1e-05
GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor (H... 51 1e-05
WASL_BOVIN (Q95107) Neural Wiskott-Aldrich syndrome protein (N-W... 50 4e-05
Y091_NPVOP (O10341) Hypothetical 29.3 kDa protein (ORF92) 49 6e-05
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 270 bits (690), Expect = 2e-71
Identities = 198/645 (30%), Positives = 298/645 (45%), Gaps = 31/645 (4%)
Query: 42 MTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPT 101
M L+ P+S W +DT AS H Y K+ G+ I G GD + T
Sbjct: 285 MHLSGPESEWVVDTAASHHATPVRDLFCRYVAGDFGTVKM--GNTSYSKIAGIGDICIKT 342
Query: 102 SHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQT---GIPLMRC 158
+ L L V H P + NLIS L D ++ Y F+ ++ + + +
Sbjct: 343 -NVGCTLVLKDVRHVPDLRMNLISGIALDRDG-----YESY-FANQKWRLTKGSLVIAKG 395
Query: 159 NSLGDLYPVTPSVPFAGL-------AQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPT 211
+ G LY + L + LWH R+GH LQ L I+Y +
Sbjct: 396 VARGTLYRTNAEICQGELNAAQDEISVDLWHKRMGHMSEKGLQILAKKSLISYAKGTTVK 455
Query: 212 VCESCVFGKHVRLPFVSSNNVTVMPFDILHSDL-WTSPVLSSAGHRYYVLFLDDFTDFLW 270
C+ C+FGK R+ F +S+ + D+++SD+ + S G++Y+V F+DD + LW
Sbjct: 456 PCDYCLFGKQHRVSFQTSSERKLNILDLVYSDVCGPMEIESMGGNKYFVTFIDDASRKLW 515
Query: 271 TFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSC 330
+ L K QVF++F + +K L+ DNG E+ + F YC++HG+ +
Sbjct: 516 VYILKTKDQVFQVFQKFHALVERETGRKLKRLRSDNGGEYTSREFEEYCSSHGIRHEKTV 575
Query: 331 PHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPT 390
P T NG AER RTI +R++L A +P SFW A+Q A YL+N P L+ P
Sbjct: 576 PGTPQHNGVAERMNRTIVEKVRSMLRMAKLPKSFWGEAVQTACYLINRSPSVPLAFEIPE 635
Query: 391 QLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRK 450
++ ++ SY+HL+VFGC + VP KL +S PC+F+GY GY+ +D +K
Sbjct: 636 RVWTNKEVSYSHLKVFGCRAFAHVPKEQRTKLDDKSIPCIFIGYGDEEFGYRLWDPVKKK 695
Query: 451 IIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPS 510
+I SR V+F E+ A + + PS + TS + + V P
Sbjct: 696 VIRSRDVVFRESEVRTAADMSEKVKNGIIPNFVTIPSTSNNPTSAESTTDEVSEQGEQPG 755
Query: 511 QLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLS-VTIDDPTIS 569
++ P + P+R RS R + R+ + V I D
Sbjct: 756 EVIEQGEQLDEGVEEVEHPTQGEEQHQPLR----RSERPRVESRRYPSTEYVLISD---D 808
Query: 570 PLPKNPKLALSDP---NWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKAN 626
P++ K LS P AMQ E ++L +N T+ LV P ++C W+F K +
Sbjct: 809 REPESLKEVLSHPEKNQLMKAMQEEMESLQKNGTYKLVELPKGKRPLKCKWVFKLKKDGD 868
Query: 627 GCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
RYKARLV G Q G+D DE FSPVVK +IRT+L++A S
Sbjct: 869 CKLVRYKARLVVKGFEQKKGIDFDEIFSPVVKMTSIRTILSLAAS 913
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 186 bits (472), Expect = 3e-46
Identities = 130/426 (30%), Positives = 210/426 (48%), Gaps = 26/426 (6%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS H+ + T + + G+ I G L H+ +TL
Sbjct: 291 LDSGASDHLINDESLYTDSVEVVPPLKIAVAKQGEFIYATKRGIVRLRNDHE---ITLED 347
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPSVP 172
VL + NL+SV++L + +S+ FD G ++ + G+ +++ + + + PV
Sbjct: 348 VLFCKEAAGNLMSVKRLQ-EAGMSIEFDKSGVTIS--KNGLMVVKNSGMLNNVPVINFQA 404
Query: 173 FAGLAQS-----LWHSRLGHPGSSALQSL-RSNKFITYEHLN----SPTVCESCVFGKHV 222
++ A+ LWH R GH L + R N F LN S +CE C+ GK
Sbjct: 405 YSINAKHKNNFRLWHERFGHISDGKLLEIKRKNMFSDQSLLNNLELSCEICEPCLNGKQA 464
Query: 223 RLPFVSSNNVTVM--PFDILHSDLWTSPV--LSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
RLPF + T + P ++HSD+ P+ ++ Y+V+F+D FT + T+ + KS
Sbjct: 465 RLPFKQLKDKTHIKRPLFVVHSDV-CGPITPVTLDDKNYFVIFVDQFTHYCVTYLIKYKS 523
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
VF MF + HF+ V L DNG E+ + +C G+ + + PHT NG
Sbjct: 524 DVFSMFQDFVAKSEAHFNLKVVYLYIDNGREYLSNEMRQFCVKKGISYHLTVPHTPQLNG 583
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSL--SNLSPTQLLYRR 396
+ER IRTI RT+++ A + SFW A+ ATYL+N IP ++L S+ +P ++ + +
Sbjct: 584 VSERMIRTITEKARTMVSGAKLDKSFWGEAVLTATYLINRIPSRALVDSSKTPYEMWHNK 643
Query: 397 DPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRH 456
P HLRVFG Y + + K +S +F+GY G+K +D + K I++R
Sbjct: 644 KPYLKHLRVFGATVYVHIKNKQ-GKFDDKSFKSIFVGY--EPNGFKLWDAVNEKFIVARD 700
Query: 457 VIFDET 462
V+ DET
Sbjct: 701 VVVDET 706
Score = 66.2 bits (160), Expect = 4e-10
Identities = 32/88 (36%), Positives = 52/88 (58%)
Query: 583 NWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRS 642
+W+ A+ +E +A N TW + RP + NI+ W+F K G RYKARLV G +
Sbjct: 905 SWEEAINTELNAHKINNTWTITKRPENKNIVDSRWVFSVKYNELGNPIRYKARLVARGFT 964
Query: 643 QIAGVDCDETFSPVVKPATIRTVLTIAL 670
Q +D +ETF+PV + ++ R +L++ +
Sbjct: 965 QKYQIDYEETFAPVARISSFRFILSLVI 992
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 127 bits (319), Expect = 2e-28
Identities = 77/172 (44%), Positives = 100/172 (57%), Gaps = 4/172 (2%)
Query: 542 MATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTW 601
M TRS GI K ++L++T TI PK+ AL DP W AMQ E DAL RNKTW
Sbjct: 1 MLTRSKAGINKLNPKYSLTITT---TIKKEPKSVIFALKDPGWCQAMQEELDALSRNKTW 57
Query: 602 DLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPAT 661
LVP P + NI+ C W+F K ++G +R KARLV G Q G+ ET+SPVV+ AT
Sbjct: 58 ILVPPPVNQNILGCKWVFKTKLHSDGTLDRLKARLVAKGFHQEEGIYFVETYSPVVRTAT 117
Query: 662 IRTVLTIALS-GLGPFTS*MFRMLSSMVICMRQSTCISPWVSVTILILIMCV 712
IRT+L +A +G + MF+M SM I ++ CI+ V + I MCV
Sbjct: 118 IRTILNVAQQLEVGQSINWMFKMHFSMGIFKKKFICINLLVLRILFIHPMCV 169
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 109 bits (273), Expect = 3e-23
Identities = 109/420 (25%), Positives = 177/420 (41%), Gaps = 52/420 (12%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S L + S +N + Q IPI G+ L + ++ T
Sbjct: 456 IDSGASQTLVRSAHYLHHATPNSEIN--IVDAQKQDIPINAIGN--LHFNFQNGTKTSIK 511
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
LHTP I +L+S+ +L + N++ CF D P+++ GD Y
Sbjct: 512 ALHTPNIAYDLLSLSELA-NQNITACFTRNTLERSDGTVLAPIVKH---GDFYWLSKKYL 567
Query: 166 -----------PVTPSVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITY------EHLN 208
V S L H LGH ++Q +TY E N
Sbjct: 568 IPSHISKLTINNVNKSKSVNKYPYPLIHRMLGHANFRSIQKSLKKNAVTYLKESDIEWSN 627
Query: 209 SPTV-CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYVLF 261
+ T C C+ GK + V + + PF LH+D++ PV L + Y++ F
Sbjct: 628 ASTYQCPDCLIGKSTKHRHVKGSRLKYQESYEPFQYLHTDIF-GPVHHLPKSAPSYFISF 686
Query: 262 LDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYC 319
D+ T F W +PL ++ + + +FTS+ I+ F+ V +Q D G E+ N + H +
Sbjct: 687 TDEKTRFQWVYPLHDRREESILNVFTSILAFIKNQFNARVLVIQMDRGSEYTNKTLHKFF 746
Query: 320 AAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNII 379
G+ ++ S +G AER RT+ N RTLL + +P W A++ +T + N +
Sbjct: 747 TNRGITACYTTTADSRAHGVAERLNRTLLNDCRTLLHCSGLPNHLWFSAVEFSTIIRNSL 806
Query: 380 PRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYPLH 437
++ S Q T + FG P++ + +K+ PR P GY LH
Sbjct: 807 VSPK-NDKSARQHAGLAGLDITTILPFG---QPVIVNNHNPDSKIHPRGIP----GYALH 858
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 107 bits (268), Expect = 1e-22
Identities = 131/550 (23%), Positives = 219/550 (39%), Gaps = 83/550 (15%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S ++ S S+ +N + + IPI GD K T
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDIN--VVDAQKRNIPINAIGDLQFHFQDNTK--TSIK 88
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
VLHTP I +L+S+ +L + ++ CF D P+++ GD Y
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD-ITACFTKNVLERSDGTVLAPIVK---YGDFYWVSKKYL 144
Query: 166 -PVTPSVPFAGLAQS----------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-- 212
P SVP + H L H + ++ N ITY N V
Sbjct: 145 LPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITY--FNESDVDW 202
Query: 213 -------CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYV 259
C C+ GK + + + + PF LH+D++ PV L + Y++
Sbjct: 203 SSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF-GPVHNLPKSAPSYFI 261
Query: 260 LFLDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
F D+ T F W +PL ++ + + ++FT++ I+ F +V +Q D G E+ N + H
Sbjct: 262 SFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHK 321
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ +G+ ++ S +G AER RT+ + RT L + +P W A++ +T + N
Sbjct: 322 FLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRN 381
Query: 378 IIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYP 435
+ S S Q + L FG P++ + +K+ PR P GY
Sbjct: 382 SLASPK-SKKSARQHAGLAGLDISTLLPFG---QPVIVNDHNPNSKIHPRGIP----GYA 433
Query: 436 LH----HRGY--------KCFDLSHRKIIISRHVIFDETRFPFATMP---TPPPSTYDCF 480
LH GY K D ++ I+ + D+ + T ++Y F
Sbjct: 434 LHPSRNSYGYIIYLPSLKKTVDTTNYVILQGKESRLDQFNYDALTFDEDLNRLTASYQSF 493
Query: 481 --------SDPIHPSIIHQWTSP----DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPS 528
SD ++ H + S P +V+ AV+P+ ST + S S +
Sbjct: 494 IASNEIQQSDDLNIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKT 553
Query: 529 PPSPPQPVPP 538
P+ V P
Sbjct: 554 NIRAPREVDP 563
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 107 bits (268), Expect = 1e-22
Identities = 131/550 (23%), Positives = 219/550 (39%), Gaps = 83/550 (15%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S ++ S S+ +N + + IPI GD K T
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDIN--VVDAQKRNIPINAIGDLQFHFQDNTK--TSIK 88
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
VLHTP I +L+S+ +L + ++ CF D P+++ GD Y
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD-ITACFTKNVLERSDGTVLAPIVK---YGDFYWVSKKYL 144
Query: 166 -PVTPSVPFAGLAQS----------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-- 212
P SVP + H L H + ++ N ITY N V
Sbjct: 145 LPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITY--FNESDVDW 202
Query: 213 -------CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYV 259
C C+ GK + + + + PF LH+D++ PV L + Y++
Sbjct: 203 SSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF-GPVHNLPKSAPSYFI 261
Query: 260 LFLDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
F D+ T F W +PL ++ + + ++FT++ I+ F +V +Q D G E+ N + H
Sbjct: 262 SFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHK 321
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ +G+ ++ S +G AER RT+ + RT L + +P W A++ +T + N
Sbjct: 322 FLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRN 381
Query: 378 IIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYP 435
+ S S Q + L FG P++ + +K+ PR P GY
Sbjct: 382 SLASPK-SKKSARQHAGLAGLDISTLLPFG---QPVIVNDHNPNSKIHPRGIP----GYA 433
Query: 436 LH----HRGY--------KCFDLSHRKIIISRHVIFDETRFPFATMP---TPPPSTYDCF 480
LH GY K D ++ I+ + D+ + T ++Y F
Sbjct: 434 LHPSRNSYGYIIYLPSLKKTVDTTNYVILQGKESRLDQFNYDALTFDEDLNRLTASYQSF 493
Query: 481 --------SDPIHPSIIHQWTSP----DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPS 528
SD ++ H + S P +V+ AV+P+ ST + S S +
Sbjct: 494 IASNEIQQSDDLNIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKT 553
Query: 529 PPSPPQPVPP 538
P+ V P
Sbjct: 554 NIRAPREVDP 563
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 107 bits (268), Expect = 1e-22
Identities = 131/550 (23%), Positives = 219/550 (39%), Gaps = 83/550 (15%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S ++ S S+ +N + + IPI GD K T
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDIN--VVDAQKRNIPINAIGDLQFHFQDNTK--TSIK 88
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
VLHTP I +L+S+ +L + ++ CF D P+++ GD Y
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD-ITACFTKNVLERSDGTVLAPIVK---YGDFYWVSKKYL 144
Query: 166 -PVTPSVPFAGLAQS----------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-- 212
P SVP + H L H + ++ N ITY N V
Sbjct: 145 LPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITY--FNESDVDR 202
Query: 213 -------CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYV 259
C C+ GK + + + + PF LH+D++ PV L + Y++
Sbjct: 203 SSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF-GPVHNLPKSAPSYFI 261
Query: 260 LFLDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
F D+ T F W +PL ++ + + ++FT++ I+ F +V +Q D G E+ N + H
Sbjct: 262 SFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHK 321
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ +G+ ++ S +G AER RT+ + RT L + +P W A++ +T + N
Sbjct: 322 FLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRN 381
Query: 378 IIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYP 435
+ S S Q + L FG P++ + +K+ PR P GY
Sbjct: 382 SLASPK-SKKSARQHAGLAGLDISTLLPFG---QPVIVNDHNPNSKIHPRGIP----GYA 433
Query: 436 LH----HRGY--------KCFDLSHRKIIISRHVIFDETRFPFATMP---TPPPSTYDCF 480
LH GY K D ++ I+ + D+ + T ++Y F
Sbjct: 434 LHPSRNSYGYIIYLPSLKKTVDTTNYVILQGKESRLDQFNYDALTFDEDLNRLTASYQSF 493
Query: 481 --------SDPIHPSIIHQWTSP----DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPS 528
SD ++ H + S P +V+ AV+P+ ST + S S +
Sbjct: 494 IASNEIQQSDDLNIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKT 553
Query: 529 PPSPPQPVPP 538
P+ V P
Sbjct: 554 NIRAPREVDP 563
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 107 bits (268), Expect = 1e-22
Identities = 131/550 (23%), Positives = 219/550 (39%), Gaps = 83/550 (15%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S ++ S S+ +N + + IPI GD K T
Sbjct: 460 LDSGASRTLIRSAHHIHSASSNPDIN--VVDAQKRNIPINAIGDLQFHFQDNTK--TSIK 515
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
VLHTP I +L+S+ +L + ++ CF D P+++ GD Y
Sbjct: 516 VLHTPNIAYDLLSLNELAAVD-ITACFTKNVLERSDGTVLAPIVK---YGDFYWVSKKYL 571
Query: 166 -PVTPSVPFAGLAQS----------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-- 212
P SVP + H L H + ++ N ITY N V
Sbjct: 572 LPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITY--FNESDVDW 629
Query: 213 -------CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYV 259
C C+ GK + + + + PF LH+D++ PV L + Y++
Sbjct: 630 SSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF-GPVHNLPKSAPSYFI 688
Query: 260 LFLDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
F D+ T F W +PL ++ + + ++FT++ I+ F +V +Q D G E+ N + H
Sbjct: 689 SFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHK 748
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ +G+ ++ S +G AER RT+ + RT L + +P W A++ +T + N
Sbjct: 749 FLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRN 808
Query: 378 IIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYP 435
+ S S Q + L FG P++ + +K+ PR P GY
Sbjct: 809 SLASPK-SKKSARQHAGLAGLDISTLLPFG---QPVIVNDHNPNSKIHPRGIP----GYA 860
Query: 436 LH----HRGY--------KCFDLSHRKIIISRHVIFDETRFPFATMP---TPPPSTYDCF 480
LH GY K D ++ I+ + D+ + T ++Y F
Sbjct: 861 LHPSRNSYGYIIYLPSLKKTVDTTNYVILQGKESRLDQFNYDALTFDEDLNRLTASYQSF 920
Query: 481 --------SDPIHPSIIHQWTSP----DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPS 528
SD ++ H + S P +V+ AV+P+ ST + S S +
Sbjct: 921 IASNEIQQSDDLNIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKT 980
Query: 529 PPSPPQPVPP 538
P+ V P
Sbjct: 981 NIRAPREVDP 990
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 105 bits (263), Expect = 5e-22
Identities = 131/550 (23%), Positives = 219/550 (39%), Gaps = 83/550 (15%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S ++ S S+ +N + + IPI GD K T
Sbjct: 460 LDSGASRTLIRSAHHIHSASSNPDIN--VVDAQKRNIPINAIGDLQFHFQDNTK--TSIK 515
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
VLHTP I +L+S+ +L + ++ CF +V + G L GD Y
Sbjct: 516 VLHTPNIAYDLLSLNELAAVD-ITACFTK---NVLERSDGTVLAPIVQYGDFYWVSKRYL 571
Query: 166 -PVTPSVPFAGLAQS----------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-- 212
P SVP + H L H + ++ N ITY N V
Sbjct: 572 LPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITY--FNESDVDW 629
Query: 213 -------CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYV 259
C C+ GK + + + + PF LH+D++ PV L + Y++
Sbjct: 630 SSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF-GPVHNLPKSAPSYFI 688
Query: 260 LFLDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
F D+ T F W +PL ++ + + ++FT++ I+ F +V +Q D G E+ N + H
Sbjct: 689 SFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHK 748
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ +G+ ++ S +G AER RT+ + RT L + +P W A++ +T + N
Sbjct: 749 FLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRN 808
Query: 378 IIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYP 435
+ S S Q + L FG P++ + +K+ PR P GY
Sbjct: 809 SLASPK-SKKSARQHAGLAGLDISTLLPFG---QPVIVNDHNPNSKIHPRGIP----GYA 860
Query: 436 LH----HRGY--------KCFDLSHRKIIISRHVIFDETRFPFATMP---TPPPSTYDCF 480
LH GY K D ++ I+ + D+ + T ++Y F
Sbjct: 861 LHPSRNSYGYIIYLPSLKKTVDTTNYVILQGKESRLDQFNYDALTFDEDLNRLTASYQSF 920
Query: 481 --------SDPIHPSIIHQWTSP----DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPS 528
S+ ++ H + S P +V+ AV+P+ ST + S S +
Sbjct: 921 IASNEIQESNDLNIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKT 980
Query: 529 PPSPPQPVPP 538
P+ V P
Sbjct: 981 NIRAPREVDP 990
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 105 bits (262), Expect = 7e-22
Identities = 106/422 (25%), Positives = 176/422 (41%), Gaps = 56/422 (13%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS + S ++ S S+ +N + + IPI GD K T
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDIN--VVDAQKRNIPINAIGDLQFHFQDNTK--TSIK 88
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY------- 165
VLHTP I +L+S+ +L + ++ CF D P+++ GD Y
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD-ITACFTKNVLERSDGTVLAPIVK---YGDFYWVSKKYL 144
Query: 166 -PVTPSVPFAGLAQS----------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-- 212
P SVP + H L H + ++ N ITY N V
Sbjct: 145 LPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITY--FNESDVDW 202
Query: 213 -------CESCVFGKHVRLPFVSSNNVTVM----PFDILHSDLWTSPV--LSSAGHRYYV 259
C C+ GK + + + + PF LH+D++ PV L + Y++
Sbjct: 203 SSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF-GPVHNLPKSAPSYFI 261
Query: 260 LFLDDFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
F D+ T F W +PL ++ + + ++FT++ I+ F +V +Q D G E+ N + H
Sbjct: 262 SFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHK 321
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ +G+ ++ S +G AER RT+ + RT L + +P W A++ +T + N
Sbjct: 322 FLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRN 381
Query: 378 IIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPS--TTINKLQPRSSPCVFLGYP 435
+ S S Q + L FG P++ + +K+ PR P GY
Sbjct: 382 SLASPK-SKKSARQHAGLAGLDISTLLPFG---QPVIVNDHNPNSKIHPRGIP----GYA 433
Query: 436 LH 437
LH
Sbjct: 434 LH 435
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 80.5 bits (197), Expect = 2e-14
Identities = 97/433 (22%), Positives = 174/433 (39%), Gaps = 33/433 (7%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+DTG+ ++ + L +Y + + + +G + ++G G + H +
Sbjct: 414 IDTGSGVNITNDKTLLHNYEDSNRSTRFFGIGKNSSVSVKGYGYIKIKNGHNNTDNKCLL 473
Query: 113 VLHTPQIVKNLISVRQLT--TDNNVSVCFDPYGFSVHDFQT----GIPLMRCNSLGD--- 163
+ P+ +IS L T +S + G + +T G+ ++ N L +
Sbjct: 474 TYYVPEEESTIISCYDLAKKTKMVLSRKYTRLGNKIIKIKTKIVNGVIHVKMNELIERPS 533
Query: 164 ------LYPVTPSVPFAGLAQSLW----HSRLGHPGSSALQ-SLRSNKFI-TYEHLNSPT 211
T S F +S+ H R+GH G ++ S++ N + + + + P
Sbjct: 534 DDSKINAIKPTSSPGFKLNKRSITLEDAHKRMGHTGIQQIENSIKHNHYEESLDLIKEPN 593
Query: 212 V--CESCVFGKHVRLPFV--SSNNVTV--MPFDILHSDLWTSPVLSSAGH--RYYVLFLD 263
C++C K + S NN + P D++ PV SS RY ++ +D
Sbjct: 594 EFWCQTCKISKATKRNHYTGSMNNHSTDHEPGSSWCMDIF-GPVSSSNADTKRYMLIMVD 652
Query: 264 DFTDFLWTFPLSNKSQ--VFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAA 321
+ T + T NK+ + + T F V+ + D G EF N Y +
Sbjct: 653 NNTRYCMTSTHFNKNAETILAQVRKNIQYVETQFDRKVREINSDRGTEFTNDQIEEYFIS 712
Query: 322 HGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPR 381
G+ + + NG+AER IRTI TLL +++ FW +A+ AT + N +
Sbjct: 713 KGIHHILTSTQDHAANGRAERYIRTIITDATTLLRQSNLRVKFWEYAVTSATNIRNYLEH 772
Query: 382 KSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGY 441
KS L P + + R+ + + ++ + KL+P P + L + GY
Sbjct: 773 KSTGKL-PLKAISRQPVTVRLMSFLPFGEKGIIWNHNHKKLKPSGLPSIILCKDPNSYGY 831
Query: 442 KCFDLSHRKIIIS 454
K F S KI+ S
Sbjct: 832 KFFIPSKNKIVTS 844
>M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300
(ORF145a) (ORF1451)
Length = 145
Score = 60.5 bits (145), Expect = 2e-08
Identities = 26/69 (37%), Positives = 38/69 (54%)
Query: 180 LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDI 239
LWHSRL H ++ L F+ ++S CE C++GK R+ F + + T P D
Sbjct: 71 LWHSRLAHMSQRGMELLVKKGFLDSSKVSSLKFCEDCIYGKTHRVNFSTGQHTTKNPLDY 130
Query: 240 LHSDLWTSP 248
+HSDLW +P
Sbjct: 131 VHSDLWGAP 139
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 59.3 bits (142), Expect = 5e-08
Identities = 56/224 (25%), Positives = 92/224 (41%), Gaps = 25/224 (11%)
Query: 185 LGHPGSSALQSLRSNKF----ITYEHLNSPTVCESCVFGKHVRL--PFVSSNNVTVMPFD 238
+ H G A S+K+ + + + S C+ C+ L P + + PFD
Sbjct: 826 IAHTGRDATFLKVSSKYWWPNLRKDVVKSIRQCKQCLVTNATNLTSPPILRPVKPLKPFD 885
Query: 239 ILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS-----QVFEMFTSLTHQIRT 293
+ D + P+ S G+ + ++ +D T F+W +P S + M TS+
Sbjct: 886 KFYID-YIGPLPPSNGYLHVLVVVDSMTGFVWLYPTKAPSTSATVKALNMLTSIAIP--- 941
Query: 294 HFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRT 353
K L D G F +++F + G+ FS P+ +GK ERK I ++
Sbjct: 942 ------KVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTK 995
Query: 354 LLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRD 397
LL P+ W+ L + LN S S +P QLL+ D
Sbjct: 996 LLIGR---PAKWYDLLPVVQLALNNSYSPS-SKYTPHQLLFGVD 1035
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 57.8 bits (138), Expect = 2e-07
Identities = 50/192 (26%), Positives = 78/192 (40%), Gaps = 21/192 (10%)
Query: 213 CESCVFGKHVRL--PFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLW 270
C+ C+ L P + V PFD D + P+ S G+ + ++ +D T F+W
Sbjct: 860 CKQCLVTNAATLAAPPILRPERPVKPFDKFFID-YIGPLPPSNGYLHVLVVVDSMTGFVW 918
Query: 271 TFPLSNKS-----QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLI 325
+P S + M TS+ K + D G F + +F + G+
Sbjct: 919 LYPTKAPSTSATVKALNMLTSIAVP---------KVIHSDQGAAFTSATFADWAKNKGIQ 969
Query: 326 FRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLS 385
FS P+ +GK ERK I ++ LL P+ W+ L + LN S S
Sbjct: 970 LEFSTPYHPQSSGKVERKNSDIKRLLTKLLVGR---PAKWYDLLPVVQLALNNSYSPS-S 1025
Query: 386 NLSPTQLLYRRD 397
+P QLL+ D
Sbjct: 1026 KYTPHQLLFGID 1037
>Y091_NPVAC (P41479) Hypothetical 24.1 kDa protein in LEF4-P33
intergenic region
Length = 224
Score = 53.1 bits (126), Expect = 4e-06
Identities = 33/86 (38%), Positives = 41/86 (47%), Gaps = 12/86 (13%)
Query: 465 PFATMPTPPPSTYDCFSDPI-HPSIIHQWTSPDPSPTSVVP---PAVTPSQLPTTSTSSS 520
P +PTPPP+ P P+ I +P P PT + P P PS +P T T
Sbjct: 47 PIVPLPTPPPTPTPSPPSPTPPPTPIPPTPTPTPPPTPIPPTPTPTPPPSPIPPTPT--- 103
Query: 521 ASPPSSPSPPS-----PPQPVPPIRT 541
SPP SP PP+ PP P+PP T
Sbjct: 104 PSPPPSPIPPTPTPSPPPSPIPPTPT 129
Score = 46.6 bits (109), Expect = 4e-04
Identities = 31/78 (39%), Positives = 37/78 (46%), Gaps = 14/78 (17%)
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVP-PAVTPSQLPTTSTSSS 520
T P PTPPP+ PI P+ +P PSP P P+ PS +P T T
Sbjct: 70 TPIPPTPTPTPPPT-------PIPPT---PTPTPPPSPIPPTPTPSPPPSPIPPTPT--- 116
Query: 521 ASPPSSPSPPSPPQPVPP 538
SPP SP PP+P PP
Sbjct: 117 PSPPPSPIPPTPTPSPPP 134
Score = 44.3 bits (103), Expect = 0.002
Identities = 32/77 (41%), Positives = 35/77 (44%), Gaps = 14/77 (18%)
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVP-PAVTPSQLPTTSTSSS 520
T P PTPPPS PI P+ SP PSP P P+ PS +P T T
Sbjct: 83 TPIPPTPTPTPPPS-------PIPPT---PTPSPPPSPIPPTPTPSPPPSPIPPTPT--- 129
Query: 521 ASPPSSPSPPSPPQPVP 537
SPP SPSP P P
Sbjct: 130 PSPPPSPSPLGEPMYYP 146
Score = 41.2 bits (95), Expect = 0.015
Identities = 35/96 (36%), Positives = 45/96 (46%), Gaps = 16/96 (16%)
Query: 451 IIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPS 510
II+ I D+ + + P PP T S PI P T P PSP S PP P+
Sbjct: 19 IILLTLKILDDIQELYPNPPIVPPPT----SPPIVPLPTPPPT-PTPSPPSPTPP---PT 70
Query: 511 QLPTTSTSSSASPPSSPSPPSP-----PQPVPPIRT 541
+P T T + PP +P PP+P P P+PP T
Sbjct: 71 PIPPTPTPT---PPPTPIPPTPTPTPPPSPIPPTPT 103
>MUC2_HUMAN (Q02817) Mucin 2 precursor (Intestinal mucin 2)
Length = 5179
Score = 52.8 bits (125), Expect = 5e-06
Identities = 42/134 (31%), Positives = 57/134 (42%), Gaps = 15/134 (11%)
Query: 465 PFATMPTPPPSTYDCF---SDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSA 521
P T TPPP+T + P P+ SP +P+S + TPS ST++ +
Sbjct: 1668 PITTTTTPPPTTTPSSPITTTPSPPTTTMTTPSPTTTPSSPITTTTTPS-----STTTPS 1722
Query: 522 SPPSSPSPPSPPQ-PVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPL----PKNPK 576
PP++ + PSP P PP TM T P L +I PT SP P P
Sbjct: 1723 PPPTTMTTPSPTTTPSPPTTTMTTLPPTTTSSPLTTTPLPPSITPPTFSPFSTTTPTTPC 1782
Query: 577 LALSDPNWKSAMQS 590
+ L NW + S
Sbjct: 1783 VPLC--NWTGWLDS 1794
Score = 50.1 bits (118), Expect = 3e-05
Identities = 32/96 (33%), Positives = 44/96 (45%), Gaps = 14/96 (14%)
Query: 465 PFATMPTPPPSTYDC------FSDPIHPSIIHQWTSPDPSPTSVV--PPAVTPSQLPTT- 515
P T TPPP+T + P + + T+P P PT+ PP TPS TT
Sbjct: 1605 PTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTT 1664
Query: 516 -----STSSSASPPSSPSPPSPPQPVPPIRTMATRS 546
+T+++ P ++PS P P PP TM T S
Sbjct: 1665 PSPPITTTTTPPPTTTPSSPITTTPSPPTTTMTTPS 1700
Score = 49.3 bits (116), Expect = 6e-05
Identities = 36/112 (32%), Positives = 48/112 (42%), Gaps = 7/112 (6%)
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWT-SPDPSPTSVVPPAVTPSQLPTTSTSSS 520
T P T P+PPP+T PS T SP T+ PP TPS P T+T+++
Sbjct: 1554 TTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTTPSPPTITTTTPPPTTTPS--PPTTTTTT 1611
Query: 521 ASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLP 572
P ++PSPP+ PP T P + T PT +P P
Sbjct: 1612 PPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPP----PTTTTTPPPTTTPSP 1659
Score = 48.1 bits (113), Expect = 1e-04
Identities = 42/128 (32%), Positives = 54/128 (41%), Gaps = 17/128 (13%)
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPP-AVTPSQLPTT----- 515
T P T P+PPP+T PS T+ P T+ PP T + LPTT
Sbjct: 1398 TPSPPTTTPSPPPTTTTTLPPTTTPSPPTTTTTTPPPTTTPSPPITTTTTPLPTTTPSPP 1457
Query: 516 -STSSSASPPSSPSP----PSPP--QPVPPIRTMATRSMRGIYKPRKLFNL----SVTID 564
ST+++ P ++PSP PSPP P PP T T P + S T
Sbjct: 1458 ISTTTTPPPTTTPSPPTTTPSPPTTTPSPPTTTTTTPPPTTTPSPPMTTPITPPASTTTL 1517
Query: 565 DPTISPLP 572
PT +P P
Sbjct: 1518 PPTTTPSP 1525
Score = 48.1 bits (113), Expect = 1e-04
Identities = 34/110 (30%), Positives = 48/110 (42%), Gaps = 18/110 (16%)
Query: 465 PFATMPTPPPSTYDC--FSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSAS 522
P T TPPP+T + PI P P+ T+ +PP TPS P T+T+++
Sbjct: 1487 PTTTTTTPPPTTTPSPPMTTPITP----------PASTTTLPPTTTPS--PPTTTTTTPP 1534
Query: 523 PPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLP 572
P ++PSPP+ PP T P + T PT +P P
Sbjct: 1535 PTTTPSPPTTTPITPPTSTTTLPPTTTPSPP----PTTTTTPPPTTTPSP 1580
Score = 47.8 bits (112), Expect = 2e-04
Identities = 36/122 (29%), Positives = 50/122 (40%), Gaps = 18/122 (14%)
Query: 465 PFATMPTPPPSTYDC------FSDPIHPSIIHQWTSPDPSPTSVV--PPAVTPS------ 510
P T TPPP+T + P + + T+P P PT+ PP TPS
Sbjct: 1526 PTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTT 1585
Query: 511 QLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISP 570
P T T+++ P ++PSPP+ PP T + P S T PT +P
Sbjct: 1586 PSPPTITTTTPPPTTTPSPPTTTTTTPPPTTTPSPPTTTPITP----PTSTTTLPPTTTP 1641
Query: 571 LP 572
P
Sbjct: 1642 SP 1643
Score = 47.4 bits (111), Expect = 2e-04
Identities = 32/108 (29%), Positives = 44/108 (40%), Gaps = 23/108 (21%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P +T TPPP+T +P P T+ PP TPS P T+T+++ P
Sbjct: 1457 PISTTTTPPPTT-----------------TPSPPTTTPSPPTTTPS--PPTTTTTTPPPT 1497
Query: 525 SSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLP 572
++PSPP PP T P + T PT +P P
Sbjct: 1498 TTPSPPMTTPITPPASTTTLPPTTTPSPP----TTTTTTPPPTTTPSP 1541
Score = 45.4 bits (106), Expect = 8e-04
Identities = 36/125 (28%), Positives = 50/125 (39%), Gaps = 17/125 (13%)
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTST---- 517
T P T P PP++ PS T+ P T+ PP TP PT++T
Sbjct: 1499 TPSPPMTTPITPPASTTTLPPTTTPSPPTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPP 1558
Query: 518 SSSASPP----------SSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPT 567
+++ SPP ++PSPP+ P PP T T P + T PT
Sbjct: 1559 TTTPSPPPTTTTTPPPTTTPSPPTTTTPSPPTITTTTPPPTTTPSPP---TTTTTTPPPT 1615
Query: 568 ISPLP 572
+P P
Sbjct: 1616 TTPSP 1620
Score = 42.0 bits (97), Expect = 0.009
Identities = 33/125 (26%), Positives = 50/125 (39%), Gaps = 11/125 (8%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS----SS 520
P T P+PP T PS T+ P T+ PP TP PT++T+ ++
Sbjct: 1581 PTTTTPSPPTITTTTPPPTTTPSPPTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTT 1640
Query: 521 ASPP--SSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA 578
SPP ++ +PP P PP T + + P T I+ P P
Sbjct: 1641 PSPPPTTTTTPPPTTTPSPPTTTTPSPPITTTTTPP-----PTTTPSSPITTTPSPPTTT 1695
Query: 579 LSDPN 583
++ P+
Sbjct: 1696 MTTPS 1700
Score = 39.7 bits (91), Expect = 0.044
Identities = 22/54 (40%), Positives = 29/54 (52%), Gaps = 5/54 (9%)
Query: 493 TSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP--SSPSPPSPPQPVPPIRTMAT 544
T+P P T+ PP T + LP T+T SPP ++ +PP P PPI T T
Sbjct: 1397 TTPSPPTTTPSPPPTTTTTLPPTTT---PSPPTTTTTTPPPTTTPSPPITTTTT 1447
Score = 38.9 bits (89), Expect = 0.076
Identities = 26/88 (29%), Positives = 36/88 (40%), Gaps = 2/88 (2%)
Query: 496 DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRK 555
D T+ PP TPS PTT+T+ P ++PSPP+ PP T + + P
Sbjct: 1393 DKCITTPSPPTTTPSPPPTTTTT--LPPTTTPSPPTTTTTTPPPTTTPSPPITTTTTPLP 1450
Query: 556 LFNLSVTIDDPTISPLPKNPKLALSDPN 583
S I T P P + P+
Sbjct: 1451 TTTPSPPISTTTTPPPTTTPSPPTTTPS 1478
Score = 32.0 bits (71), Expect = 9.2
Identities = 18/70 (25%), Positives = 25/70 (35%)
Query: 468 TMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSP 527
T TP P+ + P +P P+PT P TP TT T + +
Sbjct: 4136 TTVTPTPTPTGTQTPTTTPITTTTTVTPTPTPTGTQTPTTTPITTTTTVTPTPTPTGTQT 4195
Query: 528 SPPSPPQPVP 537
PP+ P
Sbjct: 4196 GPPTHTSTAP 4205
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 51.6 bits (122), Expect = 1e-05
Identities = 44/164 (26%), Positives = 67/164 (40%), Gaps = 13/164 (7%)
Query: 236 PFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHF 295
PFD D + P+ S G+ Y ++ +D T F W +P S TS T +
Sbjct: 675 PFDKFFID-YIGPLPPSQGYLYVLVVVDGMTGFTWLYPTKAPS------TSATVKSLNVL 727
Query: 296 SHNV--KCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRT 353
+ K + D G F +++F + G+ FS P+ K ERK I ++
Sbjct: 728 TSIAIPKVIHSDQGAAFTSSTFAEWAKERGIHLEFSTPYHPQSGSKVERKNSDIKRLLTK 787
Query: 354 LLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRD 397
LL P+ W+ L + LN L +P QLL+ D
Sbjct: 788 LLVGR---PTKWYDLLPVVQLALNNTYSPVL-KYTPHQLLFGID 827
>GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor
(Hydroxyproline-rich glycoprotein 1)
Length = 555
Score = 51.2 bits (121), Expect = 1e-05
Identities = 33/76 (43%), Positives = 39/76 (50%), Gaps = 4/76 (5%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P + P PP + + P+ PS SP P P S PP V PS P + T S SPP
Sbjct: 127 PSPSPPAPPSPSPPSPAPPLPPSPAPPSPSP-PVPPSPSPP-VPPSPAPPSPTPPSPSPP 184
Query: 525 --SSPSPPSPPQPVPP 538
SP+PPSP PVPP
Sbjct: 185 VPPSPAPPSPAPPVPP 200
Score = 51.2 bits (121), Expect = 1e-05
Identities = 30/75 (40%), Positives = 36/75 (48%), Gaps = 2/75 (2%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASP- 523
P + P PP + P PS SP P P S PP+ P P+ + S A P
Sbjct: 153 PSPSPPVPPSPSPPVPPSPAPPSPTPPSPSP-PVPPSPAPPSPAPPVPPSPAPPSPAPPV 211
Query: 524 PSSPSPPSPPQPVPP 538
P SP+PPSPP P PP
Sbjct: 212 PPSPAPPSPPSPAPP 226
Score = 50.4 bits (119), Expect = 3e-05
Identities = 32/76 (42%), Positives = 37/76 (48%), Gaps = 8/76 (10%)
Query: 464 FPFAT-MPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSAS 522
FP T MP PPS + P P T P PSP S VPP+ P S + S
Sbjct: 280 FPANTPMPPSPPSPPPSPAPPTPP------TPPSPSPPSPVPPSPAPVPPSPAPPSPAPS 333
Query: 523 PPSSPSPPSP-PQPVP 537
PP SP+PP+P P P P
Sbjct: 334 PPPSPAPPTPSPSPSP 349
Score = 49.7 bits (117), Expect = 4e-05
Identities = 31/75 (41%), Positives = 38/75 (50%), Gaps = 8/75 (10%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASP- 523
P P+P P + P PS +P PSP S PP+ +P P+ S S A P
Sbjct: 94 PSPAPPSPAPPS------PAPPSPAPPSPAP-PSPPSPAPPSPSPPAPPSPSPPSPAPPL 146
Query: 524 PSSPSPPSPPQPVPP 538
P SP+PPSP PVPP
Sbjct: 147 PPSPAPPSPSPPVPP 161
Score = 48.9 bits (115), Expect = 7e-05
Identities = 32/87 (36%), Positives = 38/87 (42%), Gaps = 13/87 (14%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPD-------PSPTSVVPPAVTPSQLPTTST 517
P P PP T S P+ PS +P PSP VPP+ P P+ +
Sbjct: 166 PVPPSPAPPSPTPPSPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAP 225
Query: 518 SSSASP----PSSPSPPSP--PQPVPP 538
S SP PS P+PPSP P P PP
Sbjct: 226 PSPPSPAPPSPSPPAPPSPVPPSPAPP 252
Score = 48.9 bits (115), Expect = 7e-05
Identities = 30/78 (38%), Positives = 36/78 (45%), Gaps = 2/78 (2%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASP- 523
P P PP + P PS SP P+P S PP+ P P+ + S + P
Sbjct: 101 PAPPSPAPPSPAPPSPAPPSPPSPAPPSPSP-PAPPSPSPPSPAPPLPPSPAPPSPSPPV 159
Query: 524 PSSPSPPSPPQPVPPIRT 541
P SPSPP PP P PP T
Sbjct: 160 PPSPSPPVPPSPAPPSPT 177
Score = 48.5 bits (114), Expect = 1e-04
Identities = 29/75 (38%), Positives = 35/75 (46%), Gaps = 6/75 (8%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPA-VTPSQLPTTSTSSSASP 523
P P PP + P PS P P+P S PP+ PS P + S +P
Sbjct: 58 PAPPSPAPPSPGPPSPAPPSPPSP----APPSPAPPSPAPPSPAPPSPAPPSPAPPSPAP 113
Query: 524 PSSPSPPSPPQPVPP 538
PS P+PPSPP P PP
Sbjct: 114 PS-PAPPSPPSPAPP 127
Score = 48.5 bits (114), Expect = 1e-04
Identities = 31/74 (41%), Positives = 35/74 (46%), Gaps = 5/74 (6%)
Query: 468 TMPTPP----PSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASP 523
T PTPP PS P+ PS +P P P S PP +PS P+ S S S SP
Sbjct: 301 TPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPP-SPAPPTPSPSPSPSPSPSPSPSP 359
Query: 524 PSSPSPPSPPQPVP 537
SPSP P P P
Sbjct: 360 SPSPSPSPSPIPSP 373
Score = 48.1 bits (113), Expect = 1e-04
Identities = 30/75 (40%), Positives = 36/75 (48%), Gaps = 2/75 (2%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASP- 523
P P+PP S P PS +P P P S PP+ +P P+ S SP
Sbjct: 114 PSPAPPSPPSPAPPSPSPPAPPSPSPPSPAP-PLPPSPAPPSPSPPVPPSPSPPVPPSPA 172
Query: 524 PSSPSPPSPPQPVPP 538
P SP+PPSP PVPP
Sbjct: 173 PPSPTPPSPSPPVPP 187
Score = 47.4 bits (111), Expect = 2e-04
Identities = 29/78 (37%), Positives = 34/78 (43%), Gaps = 9/78 (11%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P P PP P PS P P+P S PP+ P P + + SPP
Sbjct: 68 PGPPSPAPPSPPSPAPPSPAPPSP----APPSPAPPSPAPPSPAPPS-PAPPSPAPPSPP 122
Query: 525 S----SPSPPSPPQPVPP 538
S SPSPP+PP P PP
Sbjct: 123 SPAPPSPSPPAPPSPSPP 140
Score = 47.0 bits (110), Expect = 3e-04
Identities = 28/75 (37%), Positives = 35/75 (46%), Gaps = 2/75 (2%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPA-VTPSQLPTTSTSSSASP 523
P P+P P S P+ PS +P PSP+ VPP+ PS P S +
Sbjct: 148 PSPAPPSPSPPVPPSPSPPVPPSPAPPSPTP-PSPSPPVPPSPAPPSPAPPVPPSPAPPS 206
Query: 524 PSSPSPPSPPQPVPP 538
P+ P PPSP P PP
Sbjct: 207 PAPPVPPSPAPPSPP 221
Score = 47.0 bits (110), Expect = 3e-04
Identities = 25/72 (34%), Positives = 32/72 (43%)
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
P+P P + + P PS SP P+ V P PS P + + PP SP P
Sbjct: 213 PSPAPPSPPSPAPPSPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPP 272
Query: 530 PSPPQPVPPIRT 541
P PP+P P T
Sbjct: 273 PPPPRPPFPANT 284
Score = 46.2 bits (108), Expect = 5e-04
Identities = 22/45 (48%), Positives = 26/45 (56%), Gaps = 2/45 (4%)
Query: 495 PDPSPTSVVPPA-VTPSQLPTTSTSSSASPPSSPSPPSPPQPVPP 538
P P+P S PP+ PS P + S PPS P+PPSPP P PP
Sbjct: 41 PSPAPPSPAPPSPAPPSPAPPSPAPPSPGPPS-PAPPSPPSPAPP 84
Score = 45.4 bits (106), Expect = 8e-04
Identities = 29/79 (36%), Positives = 34/79 (42%), Gaps = 9/79 (11%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P + P PPP + P+ PS SP PSP PP P+ S A P
Sbjct: 267 PPSPPPPPPPRPPFPANTPMPPSP----PSPPPSPAPPTPPTPPSPSPPSPVPPSPAPVP 322
Query: 525 SSPSPPS-----PPQPVPP 538
SP+PPS PP P PP
Sbjct: 323 PSPAPPSPAPSPPPSPAPP 341
Score = 45.4 bits (106), Expect = 8e-04
Identities = 29/87 (33%), Positives = 34/87 (38%), Gaps = 17/87 (19%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLP----------- 513
P P PP + P+ PS P P+P S PPA P P
Sbjct: 226 PSPPSPAPPSPSPPAPPSPVPPSP----APPSPAPPSPKPPAPPPPPSPPPPPPPRPPFP 281
Query: 514 --TTSTSSSASPPSSPSPPSPPQPVPP 538
T S SPP SP+PP+PP P P
Sbjct: 282 ANTPMPPSPPSPPPSPAPPTPPTPPSP 308
Score = 44.7 bits (104), Expect = 0.001
Identities = 34/80 (42%), Positives = 38/80 (47%), Gaps = 9/80 (11%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLP-----TTSTSS 519
P + P+P P T P PS + SP P P S PP+ PS P T S S
Sbjct: 290 PPSPPPSPAPPTPPTPPSPSPPSPVPP--SPAPVPPSPAPPSPAPSPPPSPAPPTPSPSP 347
Query: 520 SASPPSSPSP-PSP-PQPVP 537
S SP SPSP PSP P P P
Sbjct: 348 SPSPSPSPSPSPSPSPSPSP 367
Score = 44.7 bits (104), Expect = 0.001
Identities = 28/75 (37%), Positives = 33/75 (43%), Gaps = 6/75 (8%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPA-VTPSQLPTTSTSSSASP 523
P P PP P PS P P+P S PP+ PS P + S +P
Sbjct: 63 PAPPSPGPPSPAPPSPPSPAPPSP----APPSPAPPSPAPPSPAPPSPAPPSPAPPSPAP 118
Query: 524 PSSPSPPSPPQPVPP 538
PS PSP +PP P PP
Sbjct: 119 PSPPSP-APPSPSPP 132
Score = 44.3 bits (103), Expect = 0.002
Identities = 29/75 (38%), Positives = 33/75 (43%), Gaps = 2/75 (2%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSP-DPSPTSVVPPA-VTPSQLPTTSTSSSAS 522
P PTPP P+ PS SP PSP PP+ P+ P+ S S S S
Sbjct: 295 PSPAPPTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPSPS 354
Query: 523 PPSSPSPPSPPQPVP 537
P SPSP P P P
Sbjct: 355 PSPSPSPSPSPSPSP 369
Score = 43.5 bits (101), Expect = 0.003
Identities = 23/67 (34%), Positives = 30/67 (44%), Gaps = 1/67 (1%)
Query: 473 PPSTYDCFSDPIHPSIIHQWTSP-DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPS 531
P ++C P PS +P P+P S PP+ P S S A P +P P+
Sbjct: 33 PGGIFNCPPSPAPPSPAPPSPAPPSPAPPSPAPPSPGPPSPAPPSPPSPAPPSPAPPSPA 92
Query: 532 PPQPVPP 538
PP P PP
Sbjct: 93 PPSPAPP 99
Score = 42.7 bits (99), Expect = 0.005
Identities = 28/76 (36%), Positives = 34/76 (43%), Gaps = 8/76 (10%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P P+P P + + P PS P P P PA TP S S +PP
Sbjct: 247 PSPAPPSPAPPSPKPPAPPPPPS------PPPPPPPRPPFPANTPMPPSPPSPPPSPAPP 300
Query: 525 SSPSP--PSPPQPVPP 538
+ P+P PSPP PVPP
Sbjct: 301 TPPTPPSPSPPSPVPP 316
Score = 42.7 bits (99), Expect = 0.005
Identities = 29/81 (35%), Positives = 36/81 (43%), Gaps = 13/81 (16%)
Query: 465 PFATMPTP-PPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASP 523
P P+P PPS P P+ P P+P S PP+ P P+ + S + P
Sbjct: 79 PSPAPPSPAPPSPAPPSPAPPSPA------PPSPAPPSPAPPSPAPPSPPSPAPPSPSPP 132
Query: 524 ----PSSPS--PPSPPQPVPP 538
PS PS PP PP P PP
Sbjct: 133 APPSPSPPSPAPPLPPSPAPP 153
Score = 42.0 bits (97), Expect = 0.009
Identities = 28/72 (38%), Positives = 35/72 (47%), Gaps = 16/72 (22%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P + P+PPPS P P+ P PSP+ P+ +PS P+ S S S SP
Sbjct: 327 PPSPAPSPPPS-------PAPPT-------PSPSPSP--SPSPSPSPSPSPSPSPSPSPI 370
Query: 525 SSPSPPSPPQPV 536
SPSP P PV
Sbjct: 371 PSPSPKPSPSPV 382
Score = 41.6 bits (96), Expect = 0.012
Identities = 28/78 (35%), Positives = 33/78 (41%), Gaps = 10/78 (12%)
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P P PPP S P P + + P P S PP+ PS P T + + P
Sbjct: 257 PSPKPPAPPPPP----SPPPPPPPRPPFPANTPMPPS--PPSPPPSPAPPTPPTPPSPSP 310
Query: 525 SSPSPPS----PPQPVPP 538
SP PPS PP P PP
Sbjct: 311 PSPVPPSPAPVPPSPAPP 328
Score = 41.2 bits (95), Expect = 0.015
Identities = 29/82 (35%), Positives = 35/82 (42%), Gaps = 13/82 (15%)
Query: 465 PFATMPTP-PPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPA----VTPSQLPTTSTSS 519
P P+P PPS P P+ P P+P S PP+ PS P S
Sbjct: 84 PSPAPPSPAPPSPAPPSPAPPSPA------PPSPAPPSPAPPSPPSPAPPSPSPPAPPSP 137
Query: 520 SASPPSSPSPPSP--PQPVPPI 539
S P+ P PPSP P P PP+
Sbjct: 138 SPPSPAPPLPPSPAPPSPSPPV 159
>WASL_BOVIN (Q95107) Neural Wiskott-Aldrich syndrome protein
(N-WASP)
Length = 505
Score = 49.7 bits (117), Expect = 4e-05
Identities = 29/80 (36%), Positives = 37/80 (46%), Gaps = 9/80 (11%)
Query: 462 TRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVP---PAVTPSQLPTTSTS 518
+R P A P PPPS + P P+ ++ P P +P P+ P P S S
Sbjct: 315 SRAPTAAPPPPPPSRPGVGAPPPPPNRMY------PPPLPALPSSAPSGPPPPPPPLSVS 368
Query: 519 SSASPPSSPSPPSPPQPVPP 538
S +PP P PP PP P PP
Sbjct: 369 GSVAPPPPPPPPPPPGPPPP 388
Score = 38.1 bits (87), Expect = 0.13
Identities = 32/111 (28%), Positives = 44/111 (38%), Gaps = 10/111 (9%)
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSP-- 527
P PPPS P P H P P P PS+ PT + PPS P
Sbjct: 278 PPPPPSRG---GPPPPPPPPHSSGPPPPPARGRGAPPPPPSRAPT--AAPPPPPPSRPGV 332
Query: 528 -SPPSPPQPV--PPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNP 575
+PP PP + PP+ + + + G P ++S ++ P P P P
Sbjct: 333 GAPPPPPNRMYPPPLPALPSSAPSGPPPPPPPLSVSGSVAPPPPPPPPPPP 383
>Y091_NPVOP (O10341) Hypothetical 29.3 kDa protein (ORF92)
Length = 279
Score = 49.3 bits (116), Expect = 6e-05
Identities = 27/67 (40%), Positives = 35/67 (51%), Gaps = 1/67 (1%)
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
PTP PS S P+ T P P+P+ P+ TPS PT S + S +PP SP+P
Sbjct: 127 PTPTPSPTPTPSPTPSPTPSPTPT-PSPTPSPTPTPSPTPSPTPTPSPTPSPTPPPSPTP 185
Query: 530 PSPPQPV 536
P P P+
Sbjct: 186 PPSPSPL 192
Score = 48.9 bits (115), Expect = 7e-05
Identities = 26/69 (37%), Positives = 34/69 (48%), Gaps = 3/69 (4%)
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
P+P PS S PS SP PSPT P +P+ P+ + S + +P +PSP
Sbjct: 121 PSPTPSPTPTPSPTPTPS---PTPSPTPSPTPTPSPTPSPTPTPSPTPSPTPTPSPTPSP 177
Query: 530 PSPPQPVPP 538
PP P PP
Sbjct: 178 TPPPSPTPP 186
Score = 47.4 bits (111), Expect = 2e-04
Identities = 26/68 (38%), Positives = 32/68 (46%), Gaps = 3/68 (4%)
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
PTP P+ S PS SP PSPT P +P+ PT + S + SP +PSP
Sbjct: 67 PTPSPTPTPALSPTPTPS---PTLSPTPSPTPTPSPTPSPTPSPTPTPSPTPSPTPTPSP 123
Query: 530 PSPPQPVP 537
P P P
Sbjct: 124 TPSPTPTP 131
Score = 44.3 bits (103), Expect = 0.002
Identities = 37/113 (32%), Positives = 47/113 (40%), Gaps = 7/113 (6%)
Query: 461 ETRFPFAT-MPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSS 519
ET+ P + PTP PS + P+ +P P+PT P TP+ PT + S
Sbjct: 29 ETKPPLPSPTPTPTPSPTPTPTPSPTPT---PTPTPTPTPTPTPSPTPTPALSPTPTPSP 85
Query: 520 SASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLP 572
+ SP SP+P P P P T S P S T PT SP P
Sbjct: 86 TLSPTPSPTPTPSPTPSPTPSPTPTPSPTPSPTPTPSPTPSPT---PTPSPTP 135
Score = 42.4 bits (98), Expect = 0.007
Identities = 25/68 (36%), Positives = 31/68 (44%), Gaps = 3/68 (4%)
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
PTP P+ S PS SP P+P+ P TPS P+ + S +PP SPSP
Sbjct: 135 PTPSPTPSPTPSPTPTPS---PTPSPTPTPSPTPSPTPTPSPTPSPTPPPSPTPPPSPSP 191
Query: 530 PSPPQPVP 537
P P
Sbjct: 192 LGDPMYFP 199
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.329 0.140 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 128,544,701
Number of Sequences: 164201
Number of extensions: 5698802
Number of successful extensions: 45421
Number of sequences better than 10.0: 642
Number of HSP's better than 10.0 without gapping: 211
Number of HSP's successfully gapped in prelim test: 457
Number of HSP's that attempted gapping in prelim test: 33739
Number of HSP's gapped (non-prelim): 5271
length of query: 1082
length of database: 59,974,054
effective HSP length: 121
effective length of query: 961
effective length of database: 40,105,733
effective search space: 38541609413
effective search space used: 38541609413
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0114.10


