

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.1
(661 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
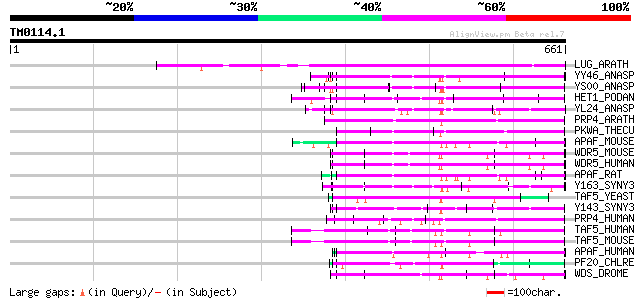
Score E
Sequences producing significant alignments: (bits) Value
LUG_ARATH (Q9FUY2) Transcriptional corepressor LEUNIG 327 6e-89
YY46_ANASP (Q8YRI1) Hypothetical WD-repeat protein alr3466 138 4e-32
YS00_ANASP (Q8YTC2) Hypothetical WD-repeat protein alr2800 135 3e-31
HET1_PODAN (Q00808) Vegetatible incompatibility protein HET-E-1 120 9e-27
YL24_ANASP (Q8YV57) Hypothetical WD-repeat protein all2124 105 3e-22
PRP4_ARATH (O22212) Hypothetical Trp-Asp repeats containing prot... 97 1e-19
PKWA_THECU (P49695) Probable serine/threonine-protein kinase pkw... 93 2e-18
APAF_MOUSE (O88879) Apoptotic protease activating factor 1 (Apaf-1) 90 2e-17
WDR5_MOUSE (P61965) WD-repeat protein 5 (WD-repeat protein BIG-3) 89 3e-17
WDR5_HUMAN (P61964) WD-repeat protein 5 89 3e-17
APAF_RAT (Q9EPV5) Apoptotic protease activating factor 1 (Apaf-1) 89 4e-17
Y163_SYNY3 (Q55563) Hypothetical WD-repeat protein sll0163 89 5e-17
TAF5_YEAST (P38129) Transcription initiation factor TFIID subuni... 89 5e-17
Y143_SYNY3 (P74442) Hypothetical WD-repeat protein slr0143 87 2e-16
PRP4_HUMAN (O43172) U4/U6 small nuclear ribonucleoprotein Prp4 (... 86 2e-16
TAF5_HUMAN (Q15542) Transcription initiation factor TFIID subuni... 86 4e-16
TAF5_MOUSE (Q8C092) Transcription initiation factor TFIID subuni... 84 9e-16
APAF_HUMAN (O14727) Apoptotic protease activating factor 1 (Apaf-1) 84 9e-16
PF20_CHLRE (P93107) Flagellar WD-repeat protein PF20 83 2e-15
WDS_DROME (Q9V3J8) Will die slowly protein 81 8e-15
>LUG_ARATH (Q9FUY2) Transcriptional corepressor LEUNIG
Length = 931
Score = 327 bits (838), Expect = 6e-89
Identities = 187/498 (37%), Positives = 265/498 (52%), Gaps = 37/498 (7%)
Query: 176 GRDVPRDPPDVVQKTMLPLDGPHQTNNNEALNLVPQPLNGGPVNTPNSHQQC-----QVL 230
G + P+ P L L P ++N +++ + GG + S QVL
Sbjct: 459 GGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSITMDGSISNSFRGNEQVL 518
Query: 231 KTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGE--H 288
K Q+ + P + T N P + S + +IS H
Sbjct: 519 KNQSGRKRKQPVSSSGPANSSGTA------NTAGPSPSSAPSTPSTHTPGDVISMPNLPH 572
Query: 289 QCNQDQQIQM---QSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPA 345
+ + M + S + + R VE D+NVESFLS ED
Sbjct: 573 SGGSSKSMMMFGTEGTGTLTSPSNQLADMDRFVEDGS-------LDDNVESFLSQEDGD- 624
Query: 346 DNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKK 405
+R + T + +KGF+ E+ + S KV CHFSSDGK++ASAGH+KK
Sbjct: 625 ----------QRDAVTRCMDVSKGFTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKK 674
Query: 406 VFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTG 465
+W + + + + H+ +ITD+RF P ATSSFD++VR+WDA N SL G
Sbjct: 675 AVLWYTDTMKPKTTLEEHTAMITDIRFSPSQLRLATSSFDKTVRVWDADNKGYSLRTFMG 734
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H V SLDFHP + DL+CS D+++ IR W++N C +GGS Q+RFQP+ G+ LA
Sbjct: 735 HSSMVTSLDFHPIKDDLICSCDNDNEIRYWSINNGSCTRVYKGGSTQIRFQPRVGKYLAA 794
Query: 526 AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWS--AASDGKC 583
+ N + ++DVET + H L+GH + S+CWD SG+FLASVSED ++W+ S+G+C
Sbjct: 795 SSANLVNVLDVETQAIRHSLQGHANPINSVCWDPSGDFLASVSEDMVKVWTLGTGSEGEC 854
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSP 643
+ EL GNKFQSC+FHP Y SLL+IG YQSLE W+ +E +KT ++ AH+GLI LA S
Sbjct: 855 VHELSCNGNKFQSCVFHPAYPSLLVIGCYQSLELWNMSE-NKTMTLPAHEGLITSLAVST 913
Query: 644 VGELIASASHDNSVKVWK 661
L+ASASHD VK+WK
Sbjct: 914 ATGLVASASHDKLVKLWK 931
Score = 43.1 bits (100), Expect = 0.002
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H E + M + PQ+
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH---SEVAASYIETQMIKAREQQLQQSQHPQVSQ 101
Query: 68 NEHRPQQFQ 76
+ + QQ Q
Sbjct: 102 QQQQQQQQQ 110
>YY46_ANASP (Q8YRI1) Hypothetical WD-repeat protein alr3466
Length = 1526
Score = 138 bits (348), Expect = 4e-32
Identities = 87/283 (30%), Positives = 145/283 (50%), Gaps = 9/283 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V FS DG +AS ++ V +W++ + + H+ + V F P + A+
Sbjct: 1159 NWVNAVAFSPDGATLASGSGDQTVRLWDISSSKCLYILQGHTSWVNSVVFNPDGSTLASG 1218
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S D++VRLW+ N + L GH V S+ F+P +L S S+ +RLW+++ S+C
Sbjct: 1219 SSDQTVRLWEI-NSSKCLCTFQGHTSWVNSVVFNPDG-SMLASGSSDKTVRLWDISSSKC 1276
Query: 503 MHTTRGGSK---QVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWD 558
+HT +G + V F P G +LA+ G+ ++ + ++ + LH +GH V S+ +
Sbjct: 1277 LHTFQGHTNWVNSVAFNPD-GSMLASGSGDQTVRLWEISSSKCLHTFQGHTSWVSSVTFS 1335
Query: 559 RSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA 617
G LAS S+D + R+WS +S G+C+ N S IF P L G Q++
Sbjct: 1336 PDGTMLASGSDDQTVRLWSISS-GECLYTFLGHTNWVGSVIFSPDGAILASGSGDQTVRL 1394
Query: 618 WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
WS + G +++ H + + SP G L+AS S D +V++W
Sbjct: 1395 WSISSGKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQTVRLW 1437
Score = 129 bits (325), Expect = 2e-29
Identities = 79/275 (28%), Positives = 136/275 (48%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F+ DG +AS ++ V +W + + + + H+ + V F P +M A+ S D++VR
Sbjct: 1208 FNPDGSTLASGSSDQTVRLWEINSSKCLCTFQGHTSWVNSVVFNPDGSMLASGSSDKTVR 1267
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD S+ + L GH V S+ F+P +L S + +RLW ++ S+C+HT +G
Sbjct: 1268 LWDISSS-KCLHTFQGHTNWVNSVAFNPDG-SMLASGSGDQTVRLWEISSSKCLHTFQGH 1325
Query: 510 SK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P L + + ++ + + + L+ GH V S+ + G LAS
Sbjct: 1326 TSWVSSVTFSPDGTMLASGSDDQTVRLWSISSGECLYTFLGHTNWVGSVIFSPDGAILAS 1385
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S D + R+WS +S GKC+ L N S +F P L Q++ W+ + G
Sbjct: 1386 GSGDQTVRLWSISS-GKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQTVRLWNISSGEC 1444
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++ H + +A S G ++AS S D ++K+W
Sbjct: 1445 LYTLHGHINSVRSVAFSSDGLILASGSDDETIKLW 1479
Score = 122 bits (305), Expect = 4e-27
Identities = 77/275 (28%), Positives = 136/275 (49%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DGK++AS ++ V +W++ + Q + H+ + V F P S M A+ S D++VR
Sbjct: 914 FSQDGKMLASGSDDQTVRLWDISSGQCLKTFKGHTSRVRSVVFSPNSLMLASGSSDQTVR 973
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD S+ L I GH V S+ F+ +L + + +RLW+++ S+C + +G
Sbjct: 974 LWDISSG-ECLYIFQGHTGWVYSVAFN-LDGSMLATGSGDQTVRLWDISSSQCFYIFQGH 1031
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ VR F L + + ++ + D+ + + L+ L+GH V S+ + G LAS
Sbjct: 1032 TSCVRSVVFSSDGAMLASGSDDQTVRLWDISSGNCLYTLQGHTSCVRSVVFSPDGAMLAS 1091
Query: 567 VSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
+D R+W +S G C+ L + + +F P +L Q + W +
Sbjct: 1092 GGDDQIVRLWDISS-GNCLYTLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLWDISSKKC 1150
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++ H + +A SP G +AS S D +V++W
Sbjct: 1151 LYTLQGHTNWVNAVAFSPDGATLASGSGDQTVRLW 1185
Score = 121 bits (303), Expect = 7e-27
Identities = 82/286 (28%), Positives = 138/286 (47%), Gaps = 9/286 (3%)
Query: 380 KSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMF 439
K + VLT FS DGK+ A+ V W + + H+ + V F M
Sbjct: 862 KILGSVLTVAFSPDGKLFATGDSGGIVRFWEAATGKELLTCKGHNSWVNSVGFSQDGKML 921
Query: 440 ATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
A+ S D++VRLWD S+ + L GH +V S+ F P + +L S S+ +RLW+++
Sbjct: 922 ASGSDDQTVRLWDISSG-QCLKTFKGHTSRVRSVVFSPNSL-MLASGSSDQTVRLWDISS 979
Query: 500 SECMHTTRGGS---KQVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSI 555
EC++ +G + V F G +LAT G+ ++ + D+ + + +GH V S+
Sbjct: 980 GECLYIFQGHTGWVYSVAFNLD-GSMLATGSGDQTVRLWDISSSQCFYIFQGHTSCVRSV 1038
Query: 556 CWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQS 614
+ G LAS S+D + R+W +S G C+ L + +S +F P L G Q
Sbjct: 1039 VFSSDGAMLASGSDDQTVRLWDISS-GNCLYTLQGHTSCVRSVVFSPDGAMLASGGDDQI 1097
Query: 615 LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ W + G+ +++ + + L SP G +A+ S D V++W
Sbjct: 1098 VRLWDISSGNCLYTLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLW 1143
Score = 118 bits (295), Expect = 6e-26
Identities = 74/276 (26%), Positives = 135/276 (48%), Gaps = 7/276 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FSSDG ++AS ++ V +W++ + + H+ + V F P M A+ D+ VR
Sbjct: 1040 FSSDGAMLASGSDDQTVRLWDISSGNCLYTLQGHTSCVRSVVFSPDGAMLASGGDDQIVR 1099
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD S+ L L G+ V L F P + L + S+ ++RLW+++ +C++T +G
Sbjct: 1100 LWDISSG-NCLYTLQGYTSWVRFLVFSPNGVTL-ANGSSDQIVRLWDISSKKCLYTLQGH 1157
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P L + + ++ + D+ + L+ L+GH V S+ ++ G+ LAS
Sbjct: 1158 TNWVNAVAFSPDGATLASGSGDQTVRLWDISSSKCLYILQGHTSWVNSVVFNPDGSTLAS 1217
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S D + R+W S KC+ + S +F+P L +++ W +
Sbjct: 1218 GSSDQTVRLWEINSS-KCLCTFQGHTSWVNSVVFNPDGSMLASGSSDKTVRLWDISSSKC 1276
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ H + +A +P G ++AS S D +V++W+
Sbjct: 1277 LHTFQGHTNWVNSVAFNPDGSMLASGSGDQTVRLWE 1312
Score = 118 bits (295), Expect = 6e-26
Identities = 78/280 (27%), Positives = 136/280 (47%), Gaps = 7/280 (2%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + F+ DG ++A+ ++ V +W++ + Q F H+ + V F M A+ S
Sbjct: 993 VYSVAFNLDGSMLATGSGDQTVRLWDISSSQCFYIFQGHTSCVRSVVFSSDGAMLASGSD 1052
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++VRLWD S+ L L GH V S+ F P +L S + ++RLW+++ C++
Sbjct: 1053 DQTVRLWDISSG-NCLYTLQGHTSCVRSVVFSPDGA-MLASGGDDQIVRLWDISSGNCLY 1110
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNSITII---DVETDSHLHDLKGHDKDVLSICWDRSG 561
T +G + VRF + A G+S I+ D+ + L+ L+GH V ++ + G
Sbjct: 1111 TLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLWDISSKKCLYTLQGHTNWVNAVAFSPDG 1170
Query: 562 NFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
LAS S D + R+W +S KC+ L + S +F+P +L Q++ W
Sbjct: 1171 ATLASGSGDQTVRLWDISS-SKCLYILQGHTSWVNSVVFNPDGSTLASGSSDQTVRLWEI 1229
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ H + + +P G ++AS S D +V++W
Sbjct: 1230 NSSKCLCTFQGHTSWVNSVVFNPDGSMLASGSSDKTVRLW 1269
Score = 96.3 bits (238), Expect = 2e-19
Identities = 68/241 (28%), Positives = 118/241 (48%), Gaps = 13/241 (5%)
Query: 359 SATCSRNENKGFSLQEIG---CLHK---SINKVLTCHFSSDGKVMASAGHEKKVFIWNME 412
S S + +K L +I CLH N V + F+ DG ++AS ++ V +W +
Sbjct: 1255 SMLASGSSDKTVRLWDISSSKCLHTFQGHTNWVNSVAFNPDGSMLASGSGDQTVRLWEIS 1314
Query: 413 NFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMS 472
+ + + H+ ++ V F P TM A+ S D++VRLW S+ L GH V S
Sbjct: 1315 SSKCLHTFQGHTSWVSSVTFSPDGTMLASGSDDQTVRLWSISSG-ECLYTFLGHTNWVGS 1373
Query: 473 LDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGN 529
+ F P +L S + +RLW+++ +C++T +G + V F P L + +
Sbjct: 1374 VIFSPDGA-ILASGSGDQTVRLWSISSGKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQ 1432
Query: 530 SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELH 588
++ + ++ + L+ L GH V S+ + G LAS S+D + ++W + G+CI L
Sbjct: 1433 TVRLWNISSGECLYTLHGHINSVRSVAFSSDGLILASGSDDETIKLWDVKT-GECIKTLK 1491
Query: 589 S 589
S
Sbjct: 1492 S 1492
>YS00_ANASP (Q8YTC2) Hypothetical WD-repeat protein alr2800
Length = 1258
Score = 135 bits (340), Expect = 3e-31
Identities = 83/291 (28%), Positives = 142/291 (48%), Gaps = 9/291 (3%)
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
I LH N+V + FS DG+ +A ++ V +WN Q + ++ V F P
Sbjct: 887 IKTLHGHTNEVCSVAFSPDGQTLACVSLDQSVRLWNCRTGQCLKAWYGNTDWALPVAFSP 946
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
+ A+ S D++V+LWD + + L GH + + + F P L S+ ++ +RL
Sbjct: 947 DRQILASGSNDKTVKLWDWQTG-KYISSLEGHTDFIYGIAFSPDSQTL-ASASTDSSVRL 1004
Query: 495 WNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDK 550
WN++ +C + V F PQ G+++AT + ++ + ++ T L L H
Sbjct: 1005 WNISTGQCFQILLEHTDWVYAVVFHPQ-GKIIATGSADCTVKLWNISTGQCLKTLSEHSD 1063
Query: 551 DVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLII 609
+L + W G LAS S D S R+W + G+C+G L N+ S IF P +
Sbjct: 1064 KILGMAWSPDGQLLASASADQSVRLWDCCT-GRCVGILRGHSNRVYSAIFSPNGEIIATC 1122
Query: 610 GGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
Q+++ W +G ++ H + +A SP G+++ASASHD +V++W
Sbjct: 1123 STDQTVKIWDWQQGKCLKTLTGHTNWVFDIAFSPDGKILASASHDQTVRIW 1173
Score = 127 bits (320), Expect = 7e-29
Identities = 82/276 (29%), Positives = 141/276 (50%), Gaps = 9/276 (3%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS D +++AS ++K V +W+ + +Y +S + H+ I + F P S A++S D SVR
Sbjct: 944 FSPDRQILASGSNDKTVKLWDWQTGKYISSLEGHTDFIYGIAFSPDSQTLASASTDSSVR 1003
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ S + IL H + V ++ FHP+ ++ + ++ ++LWN++ +C+ T
Sbjct: 1004 LWNISTG-QCFQILLEHTDWVYAVVFHPQGK-IIATGSADCTVKLWNISTGQCLKTLSEH 1061
Query: 510 SKQV---RFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
S ++ + P GQLLA+A + S+ + D T + L+GH V S + +G +A
Sbjct: 1062 SDKILGMAWSPD-GQLLASASADQSVRLWDCCTGRCVGILRGHSNRVYSAIFSPNGEIIA 1120
Query: 566 SVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGS 624
+ S D + +IW GKC+ L N F P + L Q++ W G
Sbjct: 1121 TCSTDQTVKIWDW-QQGKCLKTLTGHTNWVFDIAFSPDGKILASASHDQTVRIWDVNTGK 1179
Query: 625 KTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
H L++ +A SP GE++AS S D +V++W
Sbjct: 1180 CHHICIGHTHLVSSVAFSPDGEVVASGSQDQTVRIW 1215
Score = 120 bits (302), Expect = 9e-27
Identities = 78/297 (26%), Positives = 145/297 (48%), Gaps = 9/297 (3%)
Query: 370 FSLQEIGC--LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLI 427
F+ ++ C +++ +L+ FS +G+++A+ + V +W +++ + HS+ +
Sbjct: 628 FANSDLSCCVFTETLGNILSAAFSPEGQLLATCDTDCHVRVWEVKSGKLLLICRGHSNWV 687
Query: 428 TDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
V F P + A+ D +V+LW + + + LTGH+ +V S+ FHP + L S+
Sbjct: 688 RFVVFSPDGEILASCGADENVKLWSVRDGV-CIKTLTGHEHEVFSVAFHPDG-ETLASAS 745
Query: 488 SNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHD 544
+ I+LW++ C+ T G + VR F P L ++A ++I + DV L
Sbjct: 746 GDKTIKLWDIQDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQGKCLRT 805
Query: 545 LKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
LK H V S+ + G LAS S D + +IW+ + G+C+ N S + P
Sbjct: 806 LKSHTGWVRSVAFSADGQTLASGSGDRTIKIWNYHT-GECLKTYIGHTNSVYSIAYSPDS 864
Query: 604 RSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ L+ G ++++ W ++ H + +A SP G+ +A S D SV++W
Sbjct: 865 KILVSGSGDRTIKLWDCQTHICIKTLHGHTNEVCSVAFSPDGQTLACVSLDQSVRLW 921
Score = 119 bits (298), Expect = 3e-26
Identities = 88/321 (27%), Positives = 153/321 (47%), Gaps = 11/321 (3%)
Query: 348 RIEPFSNLKRISATCSRNEN-KGFSLQEIGCLHKSI---NKVLTCHFSSDGKVMASAGHE 403
R FS I A+C +EN K +S+++ C+ ++V + F DG+ +ASA +
Sbjct: 688 RFVVFSPDGEILASCGADENVKLWSVRDGVCIKTLTGHEHEVFSVAFHPDGETLASASGD 747
Query: 404 KKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGIL 463
K + +W++++ + H+ + V F P A+S+ D +++LWD S + L L
Sbjct: 748 KTIKLWDIQDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQG-KCLRTL 806
Query: 464 TGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYG 520
H V S+ F L S + I++WN + EC+ T G + V + P
Sbjct: 807 KSHTGWVRSVAFSADG-QTLASGSGDRTIKIWNYHTGECLKTYIGHTNSVYSIAYSPDSK 865
Query: 521 QLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAAS 579
L++ + +I + D +T + L GH +V S+ + G LA VS D S R+W+ +
Sbjct: 866 ILVSGSGDRTIKLWDCQTHICIKTLHGHTNEVCSVAFSPDGQTLACVSLDQSVRLWNCRT 925
Query: 580 DGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGL 639
G+C+ + + F P + L ++++ W G S+ H I G+
Sbjct: 926 -GQCLKAWYGNTDWALPVAFSPDRQILASGSNDKTVKLWDWQTGKYISSLEGHTDFIYGI 984
Query: 640 ADSPVGELIASASHDNSVKVW 660
A SP + +ASAS D+SV++W
Sbjct: 985 AFSPDSQTLASASTDSSVRLW 1005
Score = 96.3 bits (238), Expect = 2e-19
Identities = 61/200 (30%), Positives = 105/200 (52%), Gaps = 9/200 (4%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F GK++A+ + V +WN+ Q + HS I + + P + A++S D+SVR
Sbjct: 1028 FHPQGKIIATGSADCTVKLWNISTGQCLKTLSEHSDKILGMAWSPDGQLLASASADQSVR 1087
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD R +GIL GH +V S F P +++ + ++ +++W+ Q +C+ T G
Sbjct: 1088 LWDCCTG-RCVGILRGHSNRVYSAIFSPNG-EIIATCSTDQTVKIWDWQQGKCLKTLTGH 1145
Query: 510 SK---QVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
+ + F P G++LA+A + ++ I DV T H GH V S+ + G +A
Sbjct: 1146 TNWVFDIAFSPD-GKILASASHDQTVRIWDVNTGKCHHICIGHTHLVSSVAFSPDGEVVA 1204
Query: 566 SVSED-SARIWSAASDGKCI 584
S S+D + RIW+ + G+C+
Sbjct: 1205 SGSQDQTVRIWNVKT-GECL 1223
Score = 77.0 bits (188), Expect = 1e-13
Identities = 54/213 (25%), Positives = 95/213 (44%), Gaps = 6/213 (2%)
Query: 452 DASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSK 511
D +N S + T ++S F P LL + D++ +R+W V + + RG S
Sbjct: 627 DFANSDLSCCVFTETLGNILSAAFSPEGQ-LLATCDTDCHVRVWEVKSGKLLLICRGHSN 685
Query: 512 QVRF---QPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVS 568
VRF P L + ++ + V + L GH+ +V S+ + G LAS S
Sbjct: 686 WVRFVVFSPDGEILASCGADENVKLWSVRDGVCIKTLTGHEHEVFSVAFHPDGETLASAS 745
Query: 569 EDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTW 627
D ++W DG C+ L + + F P +L +++ W ++G
Sbjct: 746 GDKTIKLWDI-QDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQGKCLR 804
Query: 628 SIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ +H G + +A S G+ +AS S D ++K+W
Sbjct: 805 TLKSHTGWVRSVAFSADGQTLASGSGDRTIKIW 837
Score = 75.5 bits (184), Expect = 4e-13
Identities = 38/133 (28%), Positives = 71/133 (52%), Gaps = 2/133 (1%)
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
+G L N+V + FS +G+++A+ ++ V IW+ + + + H++ + D+ F P
Sbjct: 1097 VGILRGHSNRVYSAIFSPNGEIIATCSTDQTVKIWDWQQGKCLKTLTGHTNWVFDIAFSP 1156
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
+ A++S D++VR+WD N + I GH V S+ F P +++ S + +R+
Sbjct: 1157 DGKILASASHDQTVRIWDV-NTGKCHHICIGHTHLVSSVAFSPDG-EVVASGSQDQTVRI 1214
Query: 495 WNVNQSECMHTTR 507
WNV EC+ R
Sbjct: 1215 WNVKTGECLQILR 1227
Score = 62.0 bits (149), Expect = 5e-09
Identities = 37/105 (35%), Positives = 57/105 (54%), Gaps = 4/105 (3%)
Query: 352 FSNLKRISATCSRNEN-KGFSLQEIGCLHK---SINKVLTCHFSSDGKVMASAGHEKKVF 407
FS I ATCS ++ K + Q+ CL N V FS DGK++ASA H++ V
Sbjct: 1112 FSPNGEIIATCSTDQTVKIWDWQQGKCLKTLTGHTNWVFDIAFSPDGKILASASHDQTVR 1171
Query: 408 IWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
IW++ + H+HL++ V F P + A+ S D++VR+W+
Sbjct: 1172 IWDVNTGKCHHICIGHTHLVSSVAFSPDGEVVASGSQDQTVRIWN 1216
>HET1_PODAN (Q00808) Vegetatible incompatibility protein HET-E-1
Length = 1356
Score = 120 bits (302), Expect = 9e-27
Identities = 83/283 (29%), Positives = 139/283 (48%), Gaps = 9/283 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
+ VL+ FS+DG+ +AS +K + IW+ + + + H + V F P A+
Sbjct: 842 SSVLSVAFSADGQRVASGSDDKTIKIWDTASGTGTQTLEGHGGSVWSVAFSPDRERVASG 901
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S D+++++WDA++ L GH +V S+ F P + SD + I++W+ C
Sbjct: 902 SDDKTIKIWDAASG-TCTQTLEGHGGRVQSVAFSPDGQRVASGSDDH-TIKIWDAASGTC 959
Query: 503 MHTTRG-GSK--QVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWD 558
T G GS V F P GQ +A+ G+ +I I D + + L+GH V S+ +
Sbjct: 960 TQTLEGHGSSVLSVAFSPD-GQRVASGSGDKTIKIWDTASGTCTQTLEGHGGSVWSVAFS 1018
Query: 559 RSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA 617
G +AS S+D + +IW AS G C L G QS +F P + + +++
Sbjct: 1019 PDGQRVASGSDDKTIKIWDTAS-GTCTQTLEGHGGWVQSVVFSPDGQRVASGSDDHTIKI 1077
Query: 618 WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
W G+ T ++ H + +A SP G+ +AS S D ++K+W
Sbjct: 1078 WDAVSGTCTQTLEGHGDSVWSVAFSPDGQRVASGSIDGTIKIW 1120
Score = 120 bits (302), Expect = 9e-27
Identities = 80/282 (28%), Positives = 134/282 (47%), Gaps = 7/282 (2%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
+ VL+ FS DG+ +AS +K + IW+ + + + H + V F P A+
Sbjct: 968 SSVLSVAFSPDGQRVASGSGDKTIKIWDTASGTCTQTLEGHGGSVWSVAFSPDGQRVASG 1027
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S D+++++WD ++ + L GH V S+ F P + SD + I++W+ C
Sbjct: 1028 SDDKTIKIWDTASGTCTQ-TLEGHGGWVQSVVFSPDGQRVASGSDDH-TIKIWDAVSGTC 1085
Query: 503 MHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
T G V F P ++ + +I +I I D + + L+GH V S+ +
Sbjct: 1086 TQTLEGHGDSVWSVAFSPDGQRVASGSIDGTIKIWDAASGTCTQTLEGHGGWVHSVAFSP 1145
Query: 560 SGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
G +AS S D +IW AAS G C L G QS F P + + ++++ W
Sbjct: 1146 DGQRVASGSIDGTIKIWDAAS-GTCTQTLEGHGGWVQSVAFSPDGQRVASGSSDKTIKIW 1204
Query: 619 SPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G+ T ++ H G + +A SP G+ +AS S DN++K+W
Sbjct: 1205 DTASGTCTQTLEGHGGWVQSVAFSPDGQRVASGSSDNTIKIW 1246
Score = 101 bits (251), Expect = 7e-21
Identities = 65/242 (26%), Positives = 112/242 (45%), Gaps = 7/242 (2%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H + V F A+ S D+++++WD ++ + L GH V S+ F P R +
Sbjct: 840 HGSSVLSVAFSADGQRVASGSDDKTIKIWDTASGTGTQ-TLEGHGGSVWSVAFSPDRERV 898
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETD 539
SD I++W+ C T G +V+ F P ++ + + ++I I D +
Sbjct: 899 ASGSDDK-TIKIWDAASGTCTQTLEGHGGRVQSVAFSPDGQRVASGSDDHTIKIWDAASG 957
Query: 540 SHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCI 598
+ L+GH VLS+ + G +AS S D +IW AS G C L G S
Sbjct: 958 TCTQTLEGHGSSVLSVAFSPDGQRVASGSGDKTIKIWDTAS-GTCTQTLEGHGGSVWSVA 1016
Query: 599 FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVK 658
F P + + ++++ W G+ T ++ H G + + SP G+ +AS S D+++K
Sbjct: 1017 FSPDGQRVASGSDDKTIKIWDTASGTCTQTLEGHGGWVQSVVFSPDGQRVASGSDDHTIK 1076
Query: 659 VW 660
+W
Sbjct: 1077 IW 1078
Score = 88.6 bits (218), Expect = 5e-17
Identities = 78/305 (25%), Positives = 127/305 (41%), Gaps = 27/305 (8%)
Query: 336 SFLSFEDEPADNRIEPFSNLKRI------SATCSRN-ENKGFSLQEIGCLHKSINKVLTC 388
S LS P R+ S K I S TC++ E G S+ +
Sbjct: 969 SVLSVAFSPDGQRVASGSGDKTIKIWDTASGTCTQTLEGHGGSVWSVA------------ 1016
Query: 389 HFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSV 448
FS DG+ +AS +K + IW+ + + + H + V F P A+ S D ++
Sbjct: 1017 -FSPDGQRVASGSDDKTIKIWDTASGTCTQTLEGHGGWVQSVVFSPDGQRVASGSDDHTI 1075
Query: 449 RLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG 508
++WDA + + L GH + V S+ F P + S + I++W+ C T G
Sbjct: 1076 KIWDAVSGTCTQ-TLEGHGDSVWSVAFSPDGQRV-ASGSIDGTIKIWDAASGTCTQTLEG 1133
Query: 509 GS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
V F P ++ + +I +I I D + + L+GH V S+ + G +A
Sbjct: 1134 HGGWVHSVAFSPDGQRVASGSIDGTIKIWDAASGTCTQTLEGHGGWVQSVAFSPDGQRVA 1193
Query: 566 SVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGS 624
S S D + +IW AS G C L G QS F P + + +++ W G+
Sbjct: 1194 SGSSDKTIKIWDTAS-GTCTQTLEGHGGWVQSVAFSPDGQRVASGSSDNTIKIWDTASGT 1252
Query: 625 KTWSI 629
T ++
Sbjct: 1253 CTQTL 1257
Score = 82.8 bits (203), Expect = 3e-15
Identities = 58/202 (28%), Positives = 95/202 (46%), Gaps = 6/202 (2%)
Query: 463 LTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG--GSK-QVRFQPQY 519
L GH V+S+ F + SD I++W+ T G GS V F P
Sbjct: 837 LEGHGSSVLSVAFSADGQRVASGSDDK-TIKIWDTASGTGTQTLEGHGGSVWSVAFSPDR 895
Query: 520 GQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAA 578
++ + + +I I D + + L+GH V S+ + G +AS S+D + +IW AA
Sbjct: 896 ERVASGSDDKTIKIWDAASGTCTQTLEGHGGRVQSVAFSPDGQRVASGSDDHTIKIWDAA 955
Query: 579 SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAG 638
S G C L G+ S F P + + G ++++ W G+ T ++ H G +
Sbjct: 956 S-GTCTQTLEGHGSSVLSVAFSPDGQRVASGSGDKTIKIWDTASGTCTQTLEGHGGSVWS 1014
Query: 639 LADSPVGELIASASHDNSVKVW 660
+A SP G+ +AS S D ++K+W
Sbjct: 1015 VAFSPDGQRVASGSDDKTIKIW 1036
Score = 73.6 bits (179), Expect = 2e-12
Identities = 54/203 (26%), Positives = 92/203 (44%), Gaps = 7/203 (3%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+ +AS + + IW+ + + + H + V F P A+ S D +++
Sbjct: 1059 FSPDGQRVASGSDDHTIKIWDAVSGTCTQTLEGHGDSVWSVAFSPDGQRVASGSIDGTIK 1118
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
+WDA++ + L GH V S+ F P + S + I++W+ C T G
Sbjct: 1119 IWDAASGTCTQ-TLEGHGGWVHSVAFSPDGQRV-ASGSIDGTIKIWDAASGTCTQTLEGH 1176
Query: 510 S---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P ++ + + +I I D + + L+GH V S+ + G +AS
Sbjct: 1177 GGWVQSVAFSPDGQRVASGSSDKTIKIWDTASGTCTQTLEGHGGWVQSVAFSPDGQRVAS 1236
Query: 567 VSEDSA-RIWSAASDGKCIGELH 588
S D+ +IW AS G C L+
Sbjct: 1237 GSSDNTIKIWDTAS-GTCTQTLN 1258
Score = 68.9 bits (167), Expect = 4e-11
Identities = 38/133 (28%), Positives = 68/133 (50%), Gaps = 2/133 (1%)
Query: 529 NSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGEL 587
++I++++ E ++ L+GH VLS+ + G +AS S+D +IW AS G L
Sbjct: 821 STISVVEAEWNACTQTLEGHGSSVLSVAFSADGQRVASGSDDKTIKIWDTAS-GTGTQTL 879
Query: 588 HSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGEL 647
G S F P + ++++ W G+ T ++ H G + +A SP G+
Sbjct: 880 EGHGGSVWSVAFSPDRERVASGSDDKTIKIWDAASGTCTQTLEGHGGRVQSVAFSPDGQR 939
Query: 648 IASASHDNSVKVW 660
+AS S D+++K+W
Sbjct: 940 VASGSDDHTIKIW 952
Score = 44.3 bits (103), Expect = 0.001
Identities = 21/86 (24%), Positives = 41/86 (47%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+ +AS +K + IW+ + + + H + V F P A+ S D +++
Sbjct: 1185 FSPDGQRVASGSSDKTIKIWDTASGTCTQTLEGHGGWVQSVAFSPDGQRVASGSSDNTIK 1244
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDF 475
+WD ++ + + G +S D+
Sbjct: 1245 IWDTASGTCTQTLNVGSTATCLSFDY 1270
>YL24_ANASP (Q8YV57) Hypothetical WD-repeat protein all2124
Length = 1683
Score = 105 bits (263), Expect = 3e-22
Identities = 80/323 (24%), Positives = 139/323 (42%), Gaps = 48/323 (14%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
+ V +FSSDGK +ASA + + +WN + + HS + V F P S + A++
Sbjct: 1282 DSVWDVNFSSDGKAIASASRDNTIKLWNRHGIE-LETFTGHSGGVYAVNFLPDSNIIASA 1340
Query: 443 SFDRSVRLWDASNPIRSLGILTG----------HDEQVMS-------------------- 472
S D ++RLW I L +L G HD +++
Sbjct: 1341 SLDNTIRLWQRPL-ISPLEVLAGNSGVYAVSFLHDGSIIATAGADGNIQLWHSQDGSLLK 1399
Query: 473 ----------LDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQY 519
+ F P+ DL+ S++++ +++W V + + T G +V F P
Sbjct: 1400 TLPGNKAIYGISFTPQG-DLIASANADKTVKIWRVRDGKALKTLIGHDNEVNKVNFSPDG 1458
Query: 520 GQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAA 578
L + + N++ + +V LKGH +V + + G +AS S D R+W +
Sbjct: 1459 KTLASASRDNTVKLWNVSDGKFKKTLKGHTDEVFWVSFSPDGKIIASASADKTIRLWDSF 1518
Query: 579 SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAG 638
S G I L + + S F+P L ++++ W +G + + H ++
Sbjct: 1519 S-GNLIKSLPAHNDLVYSVNFNPDGSMLASTSADKTVKLWRSHDGHLLHTFSGHSNVVYS 1577
Query: 639 LADSPVGELIASASHDNSVKVWK 661
+ SP G IASAS D +VK+W+
Sbjct: 1578 SSFSPDGRYIASASEDKTVKIWQ 1600
Score = 105 bits (261), Expect = 5e-22
Identities = 80/313 (25%), Positives = 145/313 (45%), Gaps = 11/313 (3%)
Query: 353 SNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNME 412
+NLK AT + + F +QE L + V++ S DG+ +AS +K + +W+ +
Sbjct: 1045 NNLKL--ATVTTLQQALFEMQERNRLEGHKDGVISISISRDGQTIASGSLDKTIKLWSRD 1102
Query: 413 NFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMS 472
+ F + + H + V F P A+ D++++LW S+ L +TGH++ V +
Sbjct: 1103 G-RLFRTLNGHEDAVYSVSFSPDGQTIASGGSDKTIKLWQTSDGTL-LKTITGHEQTVNN 1160
Query: 473 LDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSK---QVRFQPQYGQLLATAIGN 529
+ F P +L S+ S+ I+LW+ + + T G S VRF P + A +
Sbjct: 1161 VYFSPDGKNL-ASASSDHSIKLWDTTSGQLLMTLTGHSAGVITVRFSPDGQTIAAGSEDK 1219
Query: 530 SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELH 588
++ + + L L GH V S+ + G LAS S D ++W A DGK + L
Sbjct: 1220 TVKLWHRQDGKLLKTLNGHQDWVNSLSFSPDGKTLASASADKTIKLWRIA-DGKLVKTLK 1278
Query: 589 SVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELI 648
+ F +++ +++ W+ G + + H G + + P +I
Sbjct: 1279 GHNDSVWDVNFSSDGKAIASASRDNTIKLWN-RHGIELETFTGHSGGVYAVNFLPDSNII 1337
Query: 649 ASASHDNSVKVWK 661
ASAS DN++++W+
Sbjct: 1338 ASASLDNTIRLWQ 1350
Score = 97.4 bits (241), Expect = 1e-19
Identities = 71/282 (25%), Positives = 138/282 (48%), Gaps = 11/282 (3%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V F DG ++A+AG + + +W+ ++ + + I + F P + A+++
Sbjct: 1366 VYAVSFLHDGSIIATAGADGNIQLWHSQDGSLLKTLPGNK-AIYGISFTPQGDLIASANA 1424
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++V++W + ++L L GHD +V ++F P L +S N V +LWNV+ +
Sbjct: 1425 DKTVKIWRVRDG-KALKTLIGHDNEVNKVNFSPDGKTLASASRDNTV-KLWNVSDGKFKK 1482
Query: 505 TTRGGSKQV---RFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWDRS 560
T +G + +V F P G+++A+A + +I + D + + + L H+ V S+ ++
Sbjct: 1483 TLKGHTDEVFWVSFSPD-GKIIASASADKTIRLWDSFSGNLIKSLPAHNDLVYSVNFNPD 1541
Query: 561 GNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
G+ LAS S D + ++W + DG + N S F P R + ++++ W
Sbjct: 1542 GSMLASTSADKTVKLWRS-HDGHLLHTFSGHSNVVYSSSFSPDGRYIASASEDKTVKIWQ 1600
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+G ++ HQ + SP G+ + S S D + K+W+
Sbjct: 1601 -IDGHLLTTLPQHQAGVMSAIFSPDGKTLISGSLDTTTKIWR 1641
Score = 92.4 bits (228), Expect = 3e-18
Identities = 79/334 (23%), Positives = 143/334 (42%), Gaps = 52/334 (15%)
Query: 376 GCLHKSIN----KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVR 431
G L +++N V + FS DG+ +AS G +K + +W + + H + +V
Sbjct: 1103 GRLFRTLNGHEDAVYSVSFSPDGQTIASGGSDKTIKLWQTSDGTLLKTITGHEQTVNNVY 1162
Query: 432 FQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV 491
F P A++S D S++LWD ++ + L LTGH V+++ F P + S+ +
Sbjct: 1163 FSPDGKNLASASSDHSIKLWDTTSG-QLLMTLTGHSAGVITVRFSPDGQTIAAGSE-DKT 1220
Query: 492 IRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGH 548
++LW+ + + T G V F P L + + +I + + + LKGH
Sbjct: 1221 VKLWHRQDGKLLKTLNGHQDWVNSLSFSPDGKTLASASADKTIKLWRIADGKLVKTLKGH 1280
Query: 549 DKDVLSICWDRSGNFLASVSEDSA-RIW------------------------------SA 577
+ V + + G +AS S D+ ++W SA
Sbjct: 1281 NDSVWDVNFSSDGKAIASASRDNTIKLWNRHGIELETFTGHSGGVYAVNFLPDSNIIASA 1340
Query: 578 ASDG-------KCIGELHSV-GNK--FQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTW 627
+ D I L + GN + H G + G +++ W +GS
Sbjct: 1341 SLDNTIRLWQRPLISPLEVLAGNSGVYAVSFLHDG-SIIATAGADGNIQLWHSQDGSLLK 1399
Query: 628 SIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
++ ++ I G++ +P G+LIASA+ D +VK+W+
Sbjct: 1400 TLPGNKA-IYGISFTPQGDLIASANADKTVKIWR 1432
Score = 85.9 bits (211), Expect = 3e-16
Identities = 62/200 (31%), Positives = 104/200 (52%), Gaps = 13/200 (6%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N+V +FS DGK +ASA + V +WN+ + ++ + H+ + V F P + A++
Sbjct: 1447 NEVNKVNFSPDGKTLASASRDNTVKLWNVSDGKFKKTLKGHTDEVFWVSFSPDGKIIASA 1506
Query: 443 SFDRSVRLWD--ASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQS 500
S D+++RLWD + N I+S L H++ V S++F+P +L S+ ++ ++LW +
Sbjct: 1507 SADKTIRLWDSFSGNLIKS---LPAHNDLVYSVNFNPDG-SMLASTSADKTVKLWRSHDG 1562
Query: 501 ECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSH-LHDLKGHDKDVLSIC 556
+HT G S V F P G+ +A+A T+ + D H L L H V+S
Sbjct: 1563 HLLHTFSGHSNVVYSSSFSPD-GRYIASA-SEDKTVKIWQIDGHLLTTLPQHQAGVMSAI 1620
Query: 557 WDRSGNFLASVSED-SARIW 575
+ G L S S D + +IW
Sbjct: 1621 FSPDGKTLISGSLDTTTKIW 1640
>PRP4_ARATH (O22212) Hypothetical Trp-Asp repeats containing protein
At2g41500
Length = 554
Score = 97.4 bits (241), Expect = 1e-19
Identities = 78/293 (26%), Positives = 126/293 (42%), Gaps = 11/293 (3%)
Query: 375 IGCLHKSINKVLT-CHFSSDGKVMASAGHEKKVFIWNMENF-QYFASADTHSHLITDVRF 432
+ C + ++ LT C FS DGK++A+ +W M A H TDV F
Sbjct: 247 LDCSNFGDDRPLTGCSFSRDGKILATCSLSGVTKLWEMPQVTNTIAVLKDHKERATDVVF 306
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P AT+S DR+ +LW + L GH +++ + FHP L ++ +
Sbjct: 307 SPVDDCLATASADRTAKLWKTDGTL--LQTFEGHLDRLARVAFHPSGK-YLGTTSYDKTW 363
Query: 493 RLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
RLW++N + G S+ V FQ + + + + D+ T + +GH
Sbjct: 364 RLWDINTGAELLLQEGHSRSVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGHI 423
Query: 550 KDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLI 608
K V S+ + +G LAS ED+ RIW K + + + N + P L
Sbjct: 424 KPVFSVNFSPNGYHLASGGEDNQCRIWDLRMR-KSLYIIPAHANLVSQVKYEPQEGYFLA 482
Query: 609 IGGYQ-SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
Y + WS + S S+A H+ +A L + IA+ SHD ++K+W
Sbjct: 483 TASYDMKVNIWSGRDFSLVKSLAGHESKVASLDITADSSCIATVSHDRTIKLW 535
Score = 41.2 bits (95), Expect = 0.009
Identities = 19/62 (30%), Positives = 33/62 (52%)
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
+G +A+A ++ KV IW+ +F S H + + S+ AT S DR+++LW
Sbjct: 477 EGYFLATASYDMKVNIWSGRDFSLVKSLAGHESKVASLDITADSSCIATVSHDRTIKLWT 536
Query: 453 AS 454
+S
Sbjct: 537 SS 538
>PKWA_THECU (P49695) Probable serine/threonine-protein kinase pkwA
(EC 2.7.1.37)
Length = 742
Score = 93.2 bits (230), Expect = 2e-18
Identities = 61/274 (22%), Positives = 118/274 (42%), Gaps = 6/274 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS G ++A +K + +W++ + + + H+ + V F P + A+ S D +VR
Sbjct: 467 FSPGGSLLAGGSGDKLIHVWDVASGDELHTLEGHTDWVRAVAFSPDGALLASGSDDATVR 526
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD + + GH V+ + F P ++ S + RLWNV +G
Sbjct: 527 LWDVA-AAEERAVFEGHTHYVLDIAFSPDG-SMVASGSRDGTARLWNVATGTEHAVLKGH 584
Query: 510 SK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P + + + +I + DV T L+ ++V+S+ + G+ L
Sbjct: 585 TDYVYAVAFSPDGSMVASGSRDGTIRLWDVATGKERDVLQAPAENVVSLAFSPDGSMLVH 644
Query: 567 VSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKT 626
S+ + +W AS G+ + + ++ F P L +++ W +
Sbjct: 645 GSDSTVHLWDVAS-GEALHTFEGHTDWVRAVAFSPDGALLASGSDDRTIRLWDVAAQEEH 703
Query: 627 WSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ H + +A P G +ASAS D ++++W
Sbjct: 704 TTLEGHTEPVHSVAFHPEGTTLASASEDGTIRIW 737
Score = 89.0 bits (219), Expect = 4e-17
Identities = 64/235 (27%), Positives = 108/235 (45%), Gaps = 8/235 (3%)
Query: 430 VRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSN 489
V F PG ++ A S D+ + +WD ++ L L GH + V ++ F P LL S +
Sbjct: 465 VAFSPGGSLLAGGSGDKLIHVWDVASG-DELHTLEGHTDWVRAVAFSPDGA-LLASGSDD 522
Query: 490 DVIRLWNVNQSECMHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLK 546
+RLW+V +E G + + F P + + + + + +V T + LK
Sbjct: 523 ATVRLWDVAAAEERAVFEGHTHYVLDIAFSPDGSMVASGSRDGTARLWNVATGTEHAVLK 582
Query: 547 GHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRS 605
GH V ++ + G+ +AS S D + R+W A+ GK L + S F P S
Sbjct: 583 GHTDYVYAVAFSPDGSMVASGSRDGTIRLWDVAT-GKERDVLQAPAENVVSLAFSPD-GS 640
Query: 606 LLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+L+ G ++ W G + H + +A SP G L+AS S D ++++W
Sbjct: 641 MLVHGSDSTVHLWDVASGEALHTFEGHTDWVRAVAFSPDGALLASGSDDRTIRLW 695
Score = 58.9 bits (141), Expect = 4e-08
Identities = 45/159 (28%), Positives = 76/159 (47%), Gaps = 6/159 (3%)
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNS-ITIIDVETDSHLHDLKGHDKDVLSICWDRSGNF 563
TT + V F P G LLA G+ I + DV + LH L+GH V ++ + G
Sbjct: 457 TTDREAVAVAFSPG-GSLLAGGSGDKLIHVWDVASGDELHTLEGHTDWVRAVAFSPDGAL 515
Query: 564 LASVSED-SARIWS-AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPT 621
LAS S+D + R+W AA++ + + E H+ + F P + + W+
Sbjct: 516 LASGSDDATVRLWDVAAAEERAVFEGHT--HYVLDIAFSPDGSMVASGSRDGTARLWNVA 573
Query: 622 EGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G++ + H + +A SP G ++AS S D ++++W
Sbjct: 574 TGTEHAVLKGHTDYVYAVAFSPDGSMVASGSRDGTIRLW 612
>APAF_MOUSE (O88879) Apoptotic protease activating factor 1 (Apaf-1)
Length = 1249
Score = 89.7 bits (221), Expect = 2e-17
Identities = 69/290 (23%), Positives = 125/290 (42%), Gaps = 20/290 (6%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+ +AS G +K + ++ E + H + F + AT S D+ V+
Sbjct: 623 FSQDGQRIASCGADKTLQVFKAETGEKLLDIKAHEDEVLCCAFSSDDSYIATCSADKKVK 682
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV-IRLWNVNQSECMHTTRG 508
+WD++ + + H EQV F LL ++ SND ++LW++NQ EC +T G
Sbjct: 683 IWDSATG-KLVHTYDEHSEQVNCCHFTNSSNHLLLATGSNDFFLKLWDLNQKECRNTMFG 741
Query: 509 GSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLK-----------GHDKDVLS 554
+ V RF P L + + ++ + DV + + + D +V+
Sbjct: 742 HTNSVNHCRFSPDDELLASCSADGTLRLWDVRSANERKSINVKRFFLSSEDPPEDVEVIV 801
Query: 555 IC--WDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNK-FQSCIFHPGYRSLLIIGG 611
C W G+ + +++ ++ + G + E+H+ + Q C F P +I
Sbjct: 802 KCCSWSADGDKIIVAAKNKVLLFDIHTSG-LLAEIHTGHHSTIQYCDFSPYDHLAVIALS 860
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+E W+ K H + G+ SP G +AS D +++VW+
Sbjct: 861 QYCVELWNIDSRLKVADCRGHLSWVHGVMFSPDGSSFLTASDDQTIRVWE 910
Score = 56.6 bits (135), Expect = 2e-07
Identities = 75/346 (21%), Positives = 132/346 (37%), Gaps = 42/346 (12%)
Query: 337 FLSFEDEPADNRIEPFSNLKRISA------TCSRNENKGFSLQEIGCL---HKSINKVLT 387
FLS ED P D +E SA ++N+ F + G L H + +
Sbjct: 787 FLSSEDPPED--VEVIVKCCSWSADGDKIIVAAKNKVLLFDIHTSGLLAEIHTGHHSTIQ 844
Query: 388 -CHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDR 446
C FS + A + V +WN+++ A H + V F P + F T+S D+
Sbjct: 845 YCDFSPYDHLAVIALSQYCVELWNIDSRLKVADCRGHLSWVHGVMFSPDGSSFLTASDDQ 904
Query: 447 SVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTT 506
++R+W+ ++ I+ + +D++ + V+ + N+ +
Sbjct: 905 TIRVWETKKVCKNSAIVL------------KQEIDVVFQENETMVLAVDNIRG---LQLI 949
Query: 507 RGGSKQVRFQPQ--------YGQLLATAIGN---SITIIDVETDSHLHDLKGHDKDVLSI 555
G + Q+ + P+ L A G+ +I II++ + GH K V I
Sbjct: 950 AGKTGQIDYLPEAQVSCCCLSPHLEYVAFGDEDGAIKIIELPNNRVFSSGVGHKKAVRHI 1009
Query: 556 CWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQS 614
+ G L S SEDS ++W+ + + H K + S G +
Sbjct: 1010 QFTADGKTLISSSEDSVIQVWNWQTGDYVFLQAHQETVKDFRLLQDSRLLSWSFDG---T 1066
Query: 615 LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ W+ G HQG + A S +S S D + K+W
Sbjct: 1067 VKVWNVITGRIERDFTCHQGTVLSCAISSDATKFSSTSADKTAKIW 1112
Score = 49.3 bits (116), Expect = 3e-05
Identities = 54/247 (21%), Positives = 97/247 (38%), Gaps = 56/247 (22%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F++DGK + S+ + + +WN + Y H + D R S + + S FD +V+
Sbjct: 1011 FTADGKTLISSSEDSVIQVWNWQTGDY-VFLQAHQETVKDFRLLQDSRLLSWS-FDGTVK 1068
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
+W+ ++TG E+ DF + +L + S+D +
Sbjct: 1069 VWN---------VITGRIER----DFTCHQGTVLSCAISSDATKF--------------- 1100
Query: 510 SKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSE 569
+T+ + I + S LH+LKGH+ V + G LA+ +
Sbjct: 1101 -------------SSTSADKTAKIWSFDLLSPLHELKGHNGCVRCSAFSLDGILLATGDD 1147
Query: 570 D-SARIWSAASDGKCIGELHSV---------GNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
+ RIW+ SDG+ + + G F P ++L+ GGY L+ W+
Sbjct: 1148 NGEIRIWN-VSDGQLLHSCAPISVEEGTATHGGWVTDVCFSPDSKTLVSAGGY--LKWWN 1204
Query: 620 PTEGSKT 626
G +
Sbjct: 1205 VATGDSS 1211
Score = 38.5 bits (88), Expect = 0.057
Identities = 31/146 (21%), Positives = 55/146 (37%), Gaps = 14/146 (9%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
VL+C SSD +S +K IW+ + H+ + F + AT
Sbjct: 1088 VLSCAISSDATKFSSTSADKTAKIWSFDLLSPLHELKGHNGCVRCSAFSLDGILLATGDD 1147
Query: 445 DRSVRLWDASN--------PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWN 496
+ +R+W+ S+ PI H V + F P L+ + ++ WN
Sbjct: 1148 NGEIRIWNVSDGQLLHSCAPISVEEGTATHGGWVTDVCFSPDSKTLV---SAGGYLKWWN 1204
Query: 497 V---NQSECMHTTRGGSKQVRFQPQY 519
V + S+ +T K++ P +
Sbjct: 1205 VATGDSSQTFYTNGTNLKKIHVSPDF 1230
>WDR5_MOUSE (P61965) WD-repeat protein 5 (WD-repeat protein BIG-3)
Length = 334
Score = 89.4 bits (220), Expect = 3e-17
Identities = 65/246 (26%), Positives = 117/246 (47%), Gaps = 11/246 (4%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H+ ++ V+F P A+SS D+ +++W A + + ++GH + + + +L
Sbjct: 44 HTKAVSSVKFSPNGEWLASSSADKLIKIWGAYDG-KFEKTISGHKLGISDVAW-SSDSNL 101
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETD 539
L S+ + +++W+V+ +C+ T +G S V F PQ +++ + S+ I DV+T
Sbjct: 102 LVSASDDKTLKIWDVSSGKCLKTLKGHSNYVFCCNFNPQSNLIVSGSFDESVRIWDVKTG 161
Query: 540 SHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCI 598
L L H V ++ ++R G+ + S S D RIW AS G+C+ L N S +
Sbjct: 162 KCLKTLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTAS-GQCLKTLIDDDNPPVSFV 220
Query: 599 -FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQG---LIAGLADSPVGELIASASHD 654
F P + +L +L+ W ++G + H+ I G+ I S S D
Sbjct: 221 KFSPNGKYILAATLDNTLKLWDYSKGKCLKTYTGHKNEKYCIFANFSVTGGKWIVSGSED 280
Query: 655 NSVKVW 660
N V +W
Sbjct: 281 NLVYIW 286
Score = 82.8 bits (203), Expect = 3e-15
Identities = 68/292 (23%), Positives = 136/292 (46%), Gaps = 23/292 (7%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + FS +G+ +AS+ +K + IW + ++ + H I+DV + S + ++S
Sbjct: 48 VSSVKFSPNGEWLASSSADKLIKIWGAYDGKFEKTISGHKLGISDVAWSSDSNLLVSASD 107
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D+++++WD S+ + L L GH V +F+P + +L+ S ++ +R+W+V +C+
Sbjct: 108 DKTLKIWDVSSG-KCLKTLKGHSNYVFCCNFNP-QSNLIVSGSFDESVRIWDVKTGKCLK 165
Query: 505 TTRGGS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLS-ICWDRS 560
T S V F +++++ I D + L L D +S + + +
Sbjct: 166 TLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPN 225
Query: 561 GNF-LASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA-- 617
G + LA+ +++ ++W S GKC+ N+ + CIF + + GG +
Sbjct: 226 GKYILAATLDNTLKLWD-YSKGKCLKTYTGHKNE-KYCIF----ANFSVTGGKWIVSGSE 279
Query: 618 ------WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASAS--HDNSVKVWK 661
W+ + H ++ A P +IASA+ +D ++K+WK
Sbjct: 280 DNLVYIWNLQTKEIVQKLQGHTDVVISTACHPTENIIASAALENDKTIKLWK 331
Score = 66.2 bits (160), Expect = 3e-10
Identities = 42/204 (20%), Positives = 97/204 (46%), Gaps = 10/204 (4%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V C+F+ ++ S ++ V IW+++ + + HS ++ V F ++ +S
Sbjct: 130 NYVFCCNFNPQSNLIVSGSFDESVRIWDVKTGKCLKTLPAHSDPVSAVHFNRDGSLIVSS 189
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S+D R+WD ++ ++ + V + F P +L ++ N ++LW+ ++ +C
Sbjct: 190 SYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPNGKYILAATLDN-TLKLWDYSKGKC 248
Query: 503 MHTTRGGSKQ-----VRFQPQYGQ-LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC 556
+ T G + F G+ +++ + N + I +++T + L+GH V+S
Sbjct: 249 LKTYTGHKNEKYCIFANFSVTGGKWIVSGSEDNLVYIWNLQTKEIVQKLQGHTDVVISTA 308
Query: 557 WDRSGNFLASV---SEDSARIWSA 577
+ N +AS ++ + ++W +
Sbjct: 309 CHPTENIIASAALENDKTIKLWKS 332
Score = 43.9 bits (102), Expect = 0.001
Identities = 32/117 (27%), Positives = 50/117 (42%), Gaps = 2/117 (1%)
Query: 545 LKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
L GH K V S+ + +G +LAS S D +IW A DGK + +
Sbjct: 41 LAGHTKAVSSVKFSPNGEWLASSSADKLIKIW-GAYDGKFEKTISGHKLGISDVAWSSDS 99
Query: 604 RSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
L+ ++L+ W + G ++ H + +P LI S S D SV++W
Sbjct: 100 NLLVSASDDKTLKIWDVSSGKCLKTLKGHSNYVFCCNFNPQSNLIVSGSFDESVRIW 156
Score = 31.2 bits (69), Expect = 9.1
Identities = 12/34 (35%), Positives = 22/34 (64%)
Query: 627 WSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++A H ++ + SP GE +AS+S D +K+W
Sbjct: 39 FTLAGHTKAVSSVKFSPNGEWLASSSADKLIKIW 72
>WDR5_HUMAN (P61964) WD-repeat protein 5
Length = 334
Score = 89.4 bits (220), Expect = 3e-17
Identities = 65/246 (26%), Positives = 117/246 (47%), Gaps = 11/246 (4%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H+ ++ V+F P A+SS D+ +++W A + + ++GH + + + +L
Sbjct: 44 HTKAVSSVKFSPNGEWLASSSADKLIKIWGAYDG-KFEKTISGHKLGISDVAW-SSDSNL 101
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETD 539
L S+ + +++W+V+ +C+ T +G S V F PQ +++ + S+ I DV+T
Sbjct: 102 LVSASDDKTLKIWDVSSGKCLKTLKGHSNYVFCCNFNPQSNLIVSGSFDESVRIWDVKTG 161
Query: 540 SHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCI 598
L L H V ++ ++R G+ + S S D RIW AS G+C+ L N S +
Sbjct: 162 KCLKTLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTAS-GQCLKTLIDDDNPPVSFV 220
Query: 599 -FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQG---LIAGLADSPVGELIASASHD 654
F P + +L +L+ W ++G + H+ I G+ I S S D
Sbjct: 221 KFSPNGKYILAATLDNTLKLWDYSKGKCLKTYTGHKNEKYCIFANFSVTGGKWIVSGSED 280
Query: 655 NSVKVW 660
N V +W
Sbjct: 281 NLVYIW 286
Score = 82.8 bits (203), Expect = 3e-15
Identities = 68/292 (23%), Positives = 136/292 (46%), Gaps = 23/292 (7%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + FS +G+ +AS+ +K + IW + ++ + H I+DV + S + ++S
Sbjct: 48 VSSVKFSPNGEWLASSSADKLIKIWGAYDGKFEKTISGHKLGISDVAWSSDSNLLVSASD 107
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D+++++WD S+ + L L GH V +F+P + +L+ S ++ +R+W+V +C+
Sbjct: 108 DKTLKIWDVSSG-KCLKTLKGHSNYVFCCNFNP-QSNLIVSGSFDESVRIWDVKTGKCLK 165
Query: 505 TTRGGS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLS-ICWDRS 560
T S V F +++++ I D + L L D +S + + +
Sbjct: 166 TLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPN 225
Query: 561 GNF-LASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA-- 617
G + LA+ +++ ++W S GKC+ N+ + CIF + + GG +
Sbjct: 226 GKYILAATLDNTLKLWD-YSKGKCLKTYTGHKNE-KYCIF----ANFSVTGGKWIVSGSE 279
Query: 618 ------WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASAS--HDNSVKVWK 661
W+ + H ++ A P +IASA+ +D ++K+WK
Sbjct: 280 DNLVYIWNLQTKEIVQKLQGHTDVVISTACHPTENIIASAALENDKTIKLWK 331
Score = 66.2 bits (160), Expect = 3e-10
Identities = 42/204 (20%), Positives = 97/204 (46%), Gaps = 10/204 (4%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V C+F+ ++ S ++ V IW+++ + + HS ++ V F ++ +S
Sbjct: 130 NYVFCCNFNPQSNLIVSGSFDESVRIWDVKTGKCLKTLPAHSDPVSAVHFNRDGSLIVSS 189
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S+D R+WD ++ ++ + V + F P +L ++ N ++LW+ ++ +C
Sbjct: 190 SYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPNGKYILAATLDN-TLKLWDYSKGKC 248
Query: 503 MHTTRGGSKQ-----VRFQPQYGQ-LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC 556
+ T G + F G+ +++ + N + I +++T + L+GH V+S
Sbjct: 249 LKTYTGHKNEKYCIFANFSVTGGKWIVSGSEDNLVYIWNLQTKEIVQKLQGHTDVVISTA 308
Query: 557 WDRSGNFLASV---SEDSARIWSA 577
+ N +AS ++ + ++W +
Sbjct: 309 CHPTENIIASAALENDKTIKLWKS 332
Score = 43.9 bits (102), Expect = 0.001
Identities = 32/117 (27%), Positives = 50/117 (42%), Gaps = 2/117 (1%)
Query: 545 LKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
L GH K V S+ + +G +LAS S D +IW A DGK + +
Sbjct: 41 LAGHTKAVSSVKFSPNGEWLASSSADKLIKIW-GAYDGKFEKTISGHKLGISDVAWSSDS 99
Query: 604 RSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
L+ ++L+ W + G ++ H + +P LI S S D SV++W
Sbjct: 100 NLLVSASDDKTLKIWDVSSGKCLKTLKGHSNYVFCCNFNPQSNLIVSGSFDESVRIW 156
Score = 31.2 bits (69), Expect = 9.1
Identities = 12/34 (35%), Positives = 22/34 (64%)
Query: 627 WSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++A H ++ + SP GE +AS+S D +K+W
Sbjct: 39 FTLAGHTKAVSSVKFSPNGEWLASSSADKLIKIW 72
>APAF_RAT (Q9EPV5) Apoptotic protease activating factor 1 (Apaf-1)
Length = 1249
Score = 89.0 bits (219), Expect = 4e-17
Identities = 69/290 (23%), Positives = 124/290 (41%), Gaps = 20/290 (6%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+ +AS G +K + ++ E + H + F + AT S D+ V+
Sbjct: 623 FSQDGQRIASCGADKTLQVFKAETGEKLLDIKAHEDEVLCCAFSSDDSYIATCSVDKKVK 682
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSND-VIRLWNVNQSECMHTTRG 508
+WD+ + + H EQV F + LL ++ SND ++LW++NQ EC +T G
Sbjct: 683 IWDSGTG-KLVHTYEEHSEQVNCCHFTNKSNHLLLATGSNDSFLKLWDLNQKECRNTMFG 741
Query: 509 GSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLK-----------GHDKDVLS 554
+ V RF P L + + ++ + DV + + + D +V+
Sbjct: 742 HTNSVTHCRFSPDDELLASCSADGTLKLWDVRSANEKKSINVKRFFLSSEDPPEDVEVIV 801
Query: 555 IC--WDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNK-FQSCIFHPGYRSLLIIGG 611
C W G+ + +++ + + G + E+H+ + Q C F P +I
Sbjct: 802 KCCSWSADGDRIIVAAKNKVLLLDIHTSG-LLTEIHTGHHSTIQYCDFSPYDHLAVIALS 860
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+E W+ K H + G+ SP G +AS D +++VW+
Sbjct: 861 QYCVELWNIDSRVKVADCRGHLSWVHGVMFSPDGSSFLTASDDQTIRVWE 910
Score = 56.2 bits (134), Expect = 3e-07
Identities = 67/301 (22%), Positives = 116/301 (38%), Gaps = 32/301 (10%)
Query: 372 LQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVR 431
L EI H S + C FS + A + V +WN+++ A H + V
Sbjct: 832 LTEIHTGHHST--IQYCDFSPYDHLAVIALSQYCVELWNIDSRVKVADCRGHLSWVHGVM 889
Query: 432 FQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV 491
F P + F T+S D+++R+W+ ++ I+ +Q + + F M +L
Sbjct: 890 FSPDGSSFLTASDDQTIRVWETRKVCKNSAIVL---KQEIDVVFQENEMMVLA------- 939
Query: 492 IRLWNVNQSECMHTTRGGSKQVRFQPQ--------YGQLLATAIGN---SITIIDVETDS 540
V+ + G + Q+ + P+ L A G+ +I II++ +
Sbjct: 940 -----VDNIRGLQLIAGKTGQIDYLPEAQVSCCCLSPHLEYVAFGDEEGAIKIIELPNNR 994
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIF 599
GH K V I + G L S SEDS ++W+ ++ + H K +
Sbjct: 995 VFSSGIGHKKAVRHIQFTADGKTLISSSEDSVIQVWNWQTEEYVFLQAHQETVKDFRLLR 1054
Query: 600 HPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKV 659
S G +++ W+ G HQG + A S +S S D + K+
Sbjct: 1055 DSRLLSWSFDG---TVKVWNVITGRIERDFTCHQGTVLSCAISSDATKFSSTSADKTAKI 1111
Query: 660 W 660
W
Sbjct: 1112 W 1112
Score = 53.9 bits (128), Expect = 1e-06
Identities = 63/265 (23%), Positives = 101/265 (37%), Gaps = 38/265 (14%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSS 443
+V C S + +A E + I + N + F+S H + ++F +SS
Sbjct: 963 QVSCCCLSPHLEYVAFGDEEGAIKIIELPNNRVFSSGIGHKKAVRHIQFTADGKTLISSS 1022
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
D +++W+ L H E V DF R L S + +++WNV
Sbjct: 1023 EDSVIQVWNWQT--EEYVFLQAHQETVK--DFRLLRDSRLLSWSFDGTVKVWNVIT---- 1074
Query: 504 HTTRGGSKQVRFQPQYGQLLATAIGNSIT------------IIDVETDSHLHDLKGHDKD 551
G + F G +L+ AI + T I E S LH+LKGH+
Sbjct: 1075 -----GRIERDFTCHQGTVLSCAISSDATKFSSTSADKTAKIWSFELPSPLHELKGHNSC 1129
Query: 552 VLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSV---------GNKFQSCIFHP 601
V + G LA+ ++ RIW+ SDG+ + + G F P
Sbjct: 1130 VRCSAFSLDGILLATGDDNGEIRIWN-VSDGQLLHLCAPISIEEGTATHGGWVTDVCFSP 1188
Query: 602 GYRSLLIIGGYQSLEAWSPTEGSKT 626
+ L+ GGY L+ W+ G +
Sbjct: 1189 DRKMLVSAGGY--LKWWNVVTGESS 1211
Score = 45.4 bits (106), Expect = 5e-04
Identities = 52/261 (19%), Positives = 104/261 (38%), Gaps = 48/261 (18%)
Query: 384 KVLTCHFS--SDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFAT 441
+V CHF+ S+ ++A+ ++ + +W++ + + H++ +T RF P + A+
Sbjct: 701 QVNCCHFTNKSNHLLLATGSNDSFLKLWDLNQKECRNTMFGHTNSVTHCRFSPDDELLAS 760
Query: 442 SSFDRSVRLWD--ASNPIRSLGILTGHDEQVMSLDFHPRRMDLL--CSSDSNDVIRLWNV 497
S D +++LWD ++N +S+ + +S + P ++++ C S S D
Sbjct: 761 CSADGTLKLWDVRSANEKKSINV----KRFFLSSEDPPEDVEVIVKCCSWSAD------- 809
Query: 498 NQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICW 557
G + A N + ++D+ T L ++ + C
Sbjct: 810 ----------------------GDRIIVAAKNKVLLLDIHTSGLLTEIHTGHHSTIQYCD 847
Query: 558 DRSGNFLA--SVSEDSARIWSAASDGK---CIGELHSVGNKFQSCIFHPGYRSLLIIGGY 612
+ LA ++S+ +W+ S K C G L V +F P S L
Sbjct: 848 FSPYDHLAVIALSQYCVELWNIDSRVKVADCRGHLSWV----HGVMFSPDGSSFLTASDD 903
Query: 613 QSLEAWSPTEGSKTWSIAAHQ 633
Q++ W + K +I Q
Sbjct: 904 QTIRVWETRKVCKNSAIVLKQ 924
>Y163_SYNY3 (Q55563) Hypothetical WD-repeat protein sll0163
Length = 1693
Score = 88.6 bits (218), Expect = 5e-17
Identities = 73/282 (25%), Positives = 126/282 (43%), Gaps = 17/282 (6%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V F+ G+++ +A + +W++E + H+ + + +F P T S
Sbjct: 1140 VRNAEFNCHGQILLTASRDGTARLWDLEGRE-IGLCQGHTSWVRNAQFSPDGQWIVTCSA 1198
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D + RLWD S+ + +L GH V + + P ++ SS S+ R+W+ +C+
Sbjct: 1199 DGTARLWDLSS--QCFAVLKGHQNWVNNALWSPDGQHIITSS-SDGTARVWS-RHGKCLG 1254
Query: 505 TTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
T RG + RF GQ + T ++ + + + L L+GH K+V + G
Sbjct: 1255 TLRGHDHNIHGARFSLD-GQKIVTYSTDNTARLWTKEGTLLTILRGHQKEVYDADFSADG 1313
Query: 562 NFLASVSED-SARIWSAASDG--KCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
F+ +VS D +AR W + G H V N F+P LL + ++ W
Sbjct: 1314 RFVFTVSADQTARQWDISQKDTITLTGHSHWVRNAH----FNPKGDRLLTVSRDKTARLW 1369
Query: 619 SPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ TEG +A HQG + SP G+ I + S D + ++W
Sbjct: 1370 T-TEGECVAVLADHQGWVREGQFSPDGQWIVTGSADKTAQLW 1410
Score = 75.1 bits (183), Expect = 5e-13
Identities = 63/280 (22%), Positives = 119/280 (42%), Gaps = 13/280 (4%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V HF+ G + + +K +W E AD H + + +F P T S
Sbjct: 1345 VRNAHFNPKGDRLLTVSRDKTARLWTTEGECVAVLAD-HQGWVREGQFSPDGQWIVTGSA 1403
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++ +LW+ + L +L GH + V+++ F P ++ +S + R+WN N +
Sbjct: 1404 DKTAQLWNVLG--KKLTVLRGHQDAVLNVRFSPDSQYIVTAS-KDGTARVWN-NTGRELA 1459
Query: 505 TTRGGSKQVRFQPQY---GQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
R K + F ++ GQ + TA ++ I + +GH+ V +
Sbjct: 1460 VLRHYEKNI-FAAEFSADGQFIVTASDDNTAGIWEIVGREVGICRGHEGPVYFAQFSADS 1518
Query: 562 NFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
++ + S D+ ARIW G+ + L + F P + + + W
Sbjct: 1519 RYILTASVDNTARIWDFL--GRPLLTLAGHQSIVYQARFSPEGNLIATVSADHTARLWDR 1576
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ G + HQGL+ + SP G+++ +AS+D + ++W
Sbjct: 1577 S-GKTVAVLYGHQGLVGTVDWSPDGQMLVTASNDGTARLW 1615
Score = 73.9 bits (180), Expect = 1e-12
Identities = 64/280 (22%), Positives = 114/280 (39%), Gaps = 11/280 (3%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSS 443
+V FS+DG+ + + ++ W++ HSH + + F P T S
Sbjct: 1303 EVYDADFSADGRFVFTVSADQTARQWDISQKDTITLTG-HSHWVRNAHFNPKGDRLLTVS 1361
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV--NQSE 501
D++ RLW + +L H V F P ++ S ++ +LWNV +
Sbjct: 1362 RDKTARLWTTEG--ECVAVLADHQGWVREGQFSPDGQWIVTGS-ADKTAQLWNVLGKKLT 1418
Query: 502 CMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
+ + VRF P Q + TA + + T L L+ ++K++ + + G
Sbjct: 1419 VLRGHQDAVLNVRFSPD-SQYIVTASKDGTARVWNNTGRELAVLRHYEKNIFAAEFSADG 1477
Query: 562 NFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
F+ + S+D +A IW G+ +G F R +L + W
Sbjct: 1478 QFIVTASDDNTAGIWEIV--GREVGICRGHEGPVYFAQFSADSRYILTASVDNTARIWD- 1534
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G ++A HQ ++ SP G LIA+ S D++ ++W
Sbjct: 1535 FLGRPLLTLAGHQSIVYQARFSPEGNLIATVSADHTARLW 1574
Score = 68.6 bits (166), Expect = 5e-11
Identities = 60/241 (24%), Positives = 103/241 (41%), Gaps = 10/241 (4%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H IT + + FAT+S D +V+LW + L GH++ V S+ F P L
Sbjct: 1054 HQEAITALDWSADGQYFATASADHTVKLWQRHG--EEVATLRGHEDWVRSVHFSPHHQFL 1111
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQP--QYGQLLATAIGNSITIIDVETDS 540
+ S N R+WN E + +G + VR +GQ+L TA + +
Sbjct: 1112 VTSGQDN-TARIWNF-AGEQLTLCQGHADWVRNAEFNCHGQILLTASRDGTARLWDLEGR 1169
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIF 599
+ +GH V + + G ++ + S D +AR+W +S +C L N + ++
Sbjct: 1170 EIGLCQGHTSWVRNAQFSPDGQWIVTCSADGTARLWDLSS--QCFAVLKGHQNWVNNALW 1227
Query: 600 HPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKV 659
P + ++ + WS G ++ H I G S G+ I + S DN+ ++
Sbjct: 1228 SPDGQHIITSSSDGTARVWS-RHGKCLGTLRGHDHNIHGARFSLDGQKIVTYSTDNTARL 1286
Query: 660 W 660
W
Sbjct: 1287 W 1287
Score = 60.8 bits (146), Expect = 1e-08
Identities = 70/339 (20%), Positives = 128/339 (37%), Gaps = 64/339 (18%)
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
+EIG + V FS DG+ + + + +W++ + Q FA H + + + +
Sbjct: 1169 REIGLCQGHTSWVRNAQFSPDGQWIVTCSADGTARLWDLSS-QCFAVLKGHQNWVNNALW 1227
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P TSS D + R+W + LG L GHD + F ++ S N
Sbjct: 1228 SPDGQHIITSSSDGTARVWSRHG--KCLGTLRGHDHNIHGARFSLDGQKIVTYSTDNTA- 1284
Query: 493 RLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHD-------- 544
RLW + + RG K+V + + +A G + + + + D
Sbjct: 1285 RLWT-KEGTLLTILRGHQKEV-YDADF-----SADGRFVFTVSADQTARQWDISQKDTIT 1337
Query: 545 LKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
L GH V + ++ G+ L +VS D +AR+W+ ++G+C+ L + F P
Sbjct: 1338 LTGHSHWVRNAHFNPKGDRLLTVSRDKTARLWT--TEGECVAVLADHQGWVREGQFSPDG 1395
Query: 604 RSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPV------------------- 644
+ ++ ++ + W+ G K + HQ + + SP
Sbjct: 1396 QWIVTGSADKTAQLWNVL-GKKLTVLRGHQDAVLNVRFSPDSQYIVTASKDGTARVWNNT 1454
Query: 645 ----------------------GELIASASHDNSVKVWK 661
G+ I +AS DN+ +W+
Sbjct: 1455 GRELAVLRHYEKNIFAAEFSADGQFIVTASDDNTAGIWE 1493
Score = 58.9 bits (141), Expect = 4e-08
Identities = 43/169 (25%), Positives = 77/169 (45%), Gaps = 8/169 (4%)
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
+E+G V FS+D + + +A + IW+ A H ++ RF
Sbjct: 1497 REVGICRGHEGPVYFAQFSADSRYILTASVDNTARIWDFLGRPLLTLAG-HQSIVYQARF 1555
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P + AT S D + RLWD S +++ +L GH V ++D+ P +L ++ ++
Sbjct: 1556 SPEGNLIATVSADHTARLWDRSG--KTVAVLYGHQGLVGTVDWSPDG-QMLVTASNDGTA 1612
Query: 493 RLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVET 538
RLW+++ E + T G VR F P +L ++ + + V+T
Sbjct: 1613 RLWDLSGRELL-TLEGHGNWVRSAEFSPDGRWVLTSSADGTAKLWPVKT 1660
Score = 53.9 bits (128), Expect = 1e-06
Identities = 53/225 (23%), Positives = 95/225 (41%), Gaps = 12/225 (5%)
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
+E+ L + FS+DG+ + +A + IW + + H + +F
Sbjct: 1456 RELAVLRHYEKNIFAAEFSADGQFIVTASDDNTAGIWEIVG-REVGICRGHEGPVYFAQF 1514
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
S T+S D + R+WD R L L GH V F P +L+ + ++
Sbjct: 1515 SADSRYILTASVDNTARIWDFLG--RPLLTLAGHQSIVYQARFSPEG-NLIATVSADHTA 1571
Query: 493 RLWNVNQS--ECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDK 550
RLW+ + ++ +G V + P GQ+L TA + + + L L+GH
Sbjct: 1572 RLWDRSGKTVAVLYGHQGLVGTVDWSPD-GQMLVTASNDGTARLWDLSGRELLTLEGHGN 1630
Query: 551 DVLSICWDRSGNF-LASVSEDSARIWSAASDGKCIGELHSVGNKF 594
V S + G + L S ++ +A++W K + +L S G ++
Sbjct: 1631 WVRSAEFSPDGRWVLTSSADGTAKLWPV----KTLPQLLSQGGQW 1671
>TAF5_YEAST (P38129) Transcription initiation factor TFIID subunit 5
(TBP-associated factor 5) (TBP-associated factor 90 kDa)
(TAFII-90)
Length = 798
Score = 88.6 bits (218), Expect = 5e-17
Identities = 68/238 (28%), Positives = 103/238 (42%), Gaps = 16/238 (6%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + FS D K + S +K V +W+M+ S H+H + DV F P FAT+S
Sbjct: 528 VYSTSFSPDNKYLLSGSEDKTVRLWSMDTHTALVSYKGHNHPVWDVSFSPLGHYFATASH 587
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++ RLW + I L I GH V + FHP + S S+ R+W+V+ + +
Sbjct: 588 DQTARLWSCDH-IYPLRIFAGHLNDVDCVSFHPNGCYVFTGS-SDKTCRMWDVSTGDSVR 645
Query: 505 TTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKD-VLSICWDRS 560
G + + P L + I + D+ T L ++GH K+ + S+ + +
Sbjct: 646 LFLGHTAPVISIAVCPDGRWLSTGSEDGIINVWDIGTGKRLKQMRGHGKNAIYSLSYSKE 705
Query: 561 GNFLASVSED-SARIW---------SAASDGKCIGELHSVGNKFQSCIFHPGYRSLLI 608
GN L S D + R+W SA D IG L V I G R +I
Sbjct: 706 GNVLISGGADHTVRVWDLKKATTEPSAEPDEPFIGYLGDVTASINQDIKEYGRRRTVI 763
Score = 52.4 bits (124), Expect = 4e-06
Identities = 60/285 (21%), Positives = 112/285 (39%), Gaps = 26/285 (9%)
Query: 380 KSINKVLTC-HFSSDGKVMASAGHEKKVFIWNME----NFQYFASADT------------ 422
++ NK ++C FS D ++ A+ + + IW+++ N A +
Sbjct: 463 QNTNKDMSCLDFSDDCRIAAAGFQDSYIKIWSLDGSSLNNPNIALNNNDKDEDPTCKTLV 522
Query: 423 -HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMD 481
HS + F P + + S D++VRLW + + +L GH+ V + F P
Sbjct: 523 GHSGTVYSTSFSPDNKYLLSGSEDKTVRLW-SMDTHTALVSYKGHNHPVWDVSFSPLG-H 580
Query: 482 LLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ---VRFQPQYGQLLATAIGNSITIIDVET 538
++ + RLW+ + + G V F P + + + + DV T
Sbjct: 581 YFATASHDQTARLWSCDHIYPLRIFAGHLNDVDCVSFHPNGCYVFTGSSDKTCRMWDVST 640
Query: 539 DSHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSC 597
+ GH V+SI G +L++ SED +W + GK + ++ G
Sbjct: 641 GDSVRLFLGHTAPVISIAVCPDGRWLSTGSEDGIINVWDIGT-GKRLKQMRGHGKNAIYS 699
Query: 598 IFHPGYRSLLIIGGY-QSLEAWSPTEGSKTWSIAAHQGLIAGLAD 641
+ + ++LI GG ++ W + + S + I L D
Sbjct: 700 LSYSKEGNVLISGGADHTVRVWDLKKATTEPSAEPDEPFIGYLGD 744
Score = 33.9 bits (76), Expect = 1.4
Identities = 18/65 (27%), Positives = 31/65 (47%)
Query: 596 SCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDN 655
S F P + LL +++ WS + S H + ++ SP+G A+ASHD
Sbjct: 530 STSFSPDNKYLLSGSEDKTVRLWSMDTHTALVSYKGHNHPVWDVSFSPLGHYFATASHDQ 589
Query: 656 SVKVW 660
+ ++W
Sbjct: 590 TARLW 594
>Y143_SYNY3 (P74442) Hypothetical WD-repeat protein slr0143
Length = 1191
Score = 86.7 bits (213), Expect = 2e-16
Identities = 71/280 (25%), Positives = 123/280 (43%), Gaps = 12/280 (4%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + SS ++ASA + V +W + ++ H+ I V F P +FAT+
Sbjct: 563 VTSVAISSHKNLIASASRDGTVHLWTPQG-EFLREFTGHTGSIYRVDFSPNGKIFATAGQ 621
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++V++WD + L L GH + V S+ F P ++L S+ + +RLW+ + +
Sbjct: 622 DQTVKIWDLDGNL--LQTLKGHQDSVYSVSFSPDG-EILASTSRDRTVRLWHWRSGKTLA 678
Query: 505 TTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
G +K V +F P L++ I + D++ + + + + W +G
Sbjct: 679 VLGGHTKSVDDAQFSPDGQTLVSVCRDGQIRLWDLD-GNLIRQFGLPEVAFFGVNWHPNG 737
Query: 562 NFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
N LA ++D + R+W+ G+ L +F P + L S+ WS
Sbjct: 738 NLLAVAADDGTVRLWT--PQGEIKATLSGHDEFVTRVVFTPDGKQLFSSSSNGSVIHWS- 794
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
T G + I GLA + G L+A + +N VKVW
Sbjct: 795 TSGKMLKKYQGYPEAIFGLALASNGALLAIGAENNLVKVW 834
Score = 60.5 bits (145), Expect = 1e-08
Identities = 48/198 (24%), Positives = 91/198 (45%), Gaps = 11/198 (5%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
+ V + FS DG+++AS ++ V +W+ + + A H+ + D +F P +
Sbjct: 643 DSVYSVSFSPDGEILASTSRDRTVRLWHWRSGKTLAVLGGHTKSVDDAQFSPDGQTLVSV 702
Query: 443 SFDRSVRLWDA-SNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSE 501
D +RLWD N IR G+ + +++HP +LL + + +RLW Q E
Sbjct: 703 CRDGQIRLWDLDGNLIRQFGL---PEVAFFGVNWHPNG-NLLAVAADDGTVRLW-TPQGE 757
Query: 502 CMHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWD 558
T G + +V F P QL +++ S+ I + L +G+ + + +
Sbjct: 758 IKATLSGHDEFVTRVVFTPDGKQLFSSSSNGSV-IHWSTSGKMLKKYQGYPEAIFGLALA 816
Query: 559 RSGNFLASVSEDS-ARIW 575
+G LA +E++ ++W
Sbjct: 817 SNGALLAIGAENNLVKVW 834
Score = 49.3 bits (116), Expect = 3e-05
Identities = 31/117 (26%), Positives = 54/117 (45%), Gaps = 4/117 (3%)
Query: 545 LKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
L GH V S+ N +AS S D + +W+ G+ + E F P
Sbjct: 556 LTGHRDGVTSVAISSHKNLIASASRDGTVHLWTP--QGEFLREFTGHTGSIYRVDFSPNG 613
Query: 604 RSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ G Q+++ W +G+ ++ HQ + ++ SP GE++AS S D +V++W
Sbjct: 614 KIFATAGQDQTVKIWD-LDGNLLQTLKGHQDSVYSVSFSPDGEILASTSRDRTVRLW 669
Score = 48.5 bits (114), Expect = 5e-05
Identities = 33/108 (30%), Positives = 52/108 (47%), Gaps = 4/108 (3%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
+S +GK + SAG E IW++E Q + + I + P S AT++ D VR
Sbjct: 1046 YSPNGKYLLSAGREGTAKIWSVEG-QLLHTLKSDPLPIDQIAISPDSQWIATAASDGMVR 1104
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV 497
LWD +R G T ++ LDF+ + LL + + D ++ W V
Sbjct: 1105 LWDQQGNLR--GEFTSTSGSLLGLDFNRQGQWLLAVAQNGD-LQSWPV 1149
Score = 33.5 bits (75), Expect = 1.8
Identities = 22/81 (27%), Positives = 39/81 (47%), Gaps = 1/81 (1%)
Query: 580 DGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGL 639
DG I + + G++ + P + LL G + + WS EG ++ + I +
Sbjct: 1027 DGTLIRSIVAHGDRLNQLQYSPNGKYLLSAGREGTAKIWS-VEGQLLHTLKSDPLPIDQI 1085
Query: 640 ADSPVGELIASASHDNSVKVW 660
A SP + IA+A+ D V++W
Sbjct: 1086 AISPDSQWIATAASDGMVRLW 1106
>PRP4_HUMAN (O43172) U4/U6 small nuclear ribonucleoprotein Prp4
(U4/U6 snRNP 60 kDa protein) (WD splicing factor Prp4)
(hPrp4)
Length = 522
Score = 86.3 bits (212), Expect = 2e-16
Identities = 76/291 (26%), Positives = 137/291 (46%), Gaps = 26/291 (8%)
Query: 388 CHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTM--------F 439
CHFS + K++A+A +W++ + + H+ + + F P ST+
Sbjct: 237 CHFSPNSKMLATACWSGLCKLWSVPDCNLLHTLRGHNTNVGAIVFHPKSTVSLDPKDVNL 296
Query: 440 ATSSFDRSVRLW--DASNPIRSLGILTGHDEQVMSLDFHP--RRMDLLCSSDSNDVIRLW 495
A+ + D SV+LW D+ P+ + GH +V + +HP R + C S RLW
Sbjct: 297 ASCAADGSVKLWSLDSDEPVADI---EGHTVRVARVMWHPSGRFLGTTCYDRS---WRLW 350
Query: 496 NVN-QSECMHTTRG--GSKQVRFQPQYGQLLATAIGNSI-TIIDVETDSHLHDLKGHDKD 551
++ Q E +H G + F Q G L T ++ + D+ T + L+GH K+
Sbjct: 351 DLEAQEEILHQEGHSMGVYDIAFH-QDGSLAGTGGLDAFGRVWDLRTGRCIMFLEGHLKE 409
Query: 552 VLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIG 610
+ I + +G +A+ S D+ ++W +C+ + + N F P + + L+ G
Sbjct: 410 IYGINFSPNGYHIATGSGDNTCKVWDLRQR-RCVYTIPAHQNLVTGVKFEPIHGNFLLTG 468
Query: 611 GYQSL-EAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
Y + + W+ S ++A H+G + GL S G+LIA+ S+D + K+W
Sbjct: 469 AYDNTAKIWTHPGWSPLKTLAGHEGKVMGLDISSDGQLIATCSYDRTFKLW 519
Score = 79.7 bits (195), Expect = 2e-14
Identities = 63/265 (23%), Positives = 111/265 (41%), Gaps = 18/265 (6%)
Query: 410 NMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQ 469
++ + F S I+ F P S M AT+ + +LW + L L GH+
Sbjct: 217 SLRSLNNFCSQIGDDRPISYCHFSPNSKMLATACWSGLCKLWSVPD-CNLLHTLRGHNTN 275
Query: 470 VMSLDFHPRRMDLLCSSDSN-------DVIRLWNVNQSECM-----HTTRGGSKQVRFQP 517
V ++ FHP+ L D N ++LW+++ E + HT R +V + P
Sbjct: 276 VGAIVFHPKSTVSLDPKDVNLASCAADGSVKLWSLDSDEPVADIEGHTVR--VARVMWHP 333
Query: 518 QYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWS 576
L T S + D+E + +GH V I + + G+ + D+ R+W
Sbjct: 334 SGRFLGTTCYDRSWRLWDLEAQEEILHQEGHSMGVYDIAFHQDGSLAGTGGLDAFGRVWD 393
Query: 577 AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLI 636
+ G+CI L + F P + G + + W + ++I AHQ L+
Sbjct: 394 LRT-GRCIMFLEGHLKEIYGINFSPNGYHIATGSGDNTCKVWDLRQRRCVYTIPAHQNLV 452
Query: 637 AGLADSPV-GELIASASHDNSVKVW 660
G+ P+ G + + ++DN+ K+W
Sbjct: 453 TGVKFEPIHGNFLLTGAYDNTAKIW 477
Score = 67.0 bits (162), Expect = 1e-10
Identities = 61/232 (26%), Positives = 98/232 (41%), Gaps = 21/232 (9%)
Query: 446 RSVRLWDASNPIRSLGILT---GHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
R+ ++ + +RSL G D + F P +L ++ + + +LW+V
Sbjct: 207 RTSQMQELHKSLRSLNNFCSQIGDDRPISYCHFSPNSK-MLATACWSGLCKLWSVPDCNL 265
Query: 503 MHTTRGGS-----------KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKD 551
+HT RG + V P+ L + A S+ + +++D + D++GH
Sbjct: 266 LHTLRGHNTNVGAIVFHPKSTVSLDPKDVNLASCAADGSVKLWSLDSDEPVADIEGHTVR 325
Query: 552 VLSICWDRSGNFLASVSED-SARIWSAASDGKCI-GELHSVGNKFQSCIFHPGYRSLLII 609
V + W SG FL + D S R+W + + + E HS+G FH SL
Sbjct: 326 VARVMWHPSGRFLGTTCYDRSWRLWDLEAQEEILHQEGHSMG--VYDIAFHQD-GSLAGT 382
Query: 610 GGYQSL-EAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
GG + W G + H I G+ SP G IA+ S DN+ KVW
Sbjct: 383 GGLDAFGRVWDLRTGRCIMFLEGHLKEIYGINFSPNGYHIATGSGDNTCKVW 434
Score = 44.7 bits (104), Expect = 8e-04
Identities = 28/120 (23%), Positives = 58/120 (48%), Gaps = 5/120 (4%)
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP-GS 436
L + ++ +FS +G +A+ + +W++ + + H +L+T V+F+P
Sbjct: 403 LEGHLKEIYGINFSPNGYHIATGSGDNTCKVWDLRQRRCVYTIPAHQNLVTGVKFEPIHG 462
Query: 437 TMFATSSFDRSVRLWDASNPIRS-LGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLW 495
T ++D + ++W ++P S L L GH+ +VM LD L+ + + +LW
Sbjct: 463 NFLLTGAYDNTAKIW--THPGWSPLKTLAGHEGKVMGLDISSDG-QLIATCSYDRTFKLW 519
>TAF5_HUMAN (Q15542) Transcription initiation factor TFIID subunit 5
(Transcription initiation factor TFIID 100 kDa subunit)
(TAF(II)100) (TAFII-100) (TAFII100)
Length = 800
Score = 85.5 bits (210), Expect = 4e-16
Identities = 67/247 (27%), Positives = 109/247 (44%), Gaps = 22/247 (8%)
Query: 336 SFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGK 395
S LS D+ +D+ +E + K S E+ L+ V FS D
Sbjct: 511 SDLSLIDKESDDVLERIMDEKTAS--------------ELKILYGHSGPVYGASFSPDRN 556
Query: 396 VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASN 455
+ S+ + V +W+++ F H++ + D +F P F + DR RLW A++
Sbjct: 557 YLLSSSEDGTVRLWSLQTFTCLVGYKGHNYPVWDTQFSPYGYYFVSGGHDRVARLW-ATD 615
Query: 456 PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHT---TRGGSKQ 512
+ L I GH V FHP + + + ++ +RLW+V C+ +G
Sbjct: 616 HYQPLRIFAGHLADVNCTRFHP-NSNYVATGSADRTVRLWDVLNGNCVRIFTGHKGPIHS 674
Query: 513 VRFQPQYGQLLAT-AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED- 570
+ F P G+ LAT A + + D+ + +LKGH V S+ + R G LAS S D
Sbjct: 675 LTFSPN-GRFLATGATDGRVLLWDIGHGLMVGELKGHTDTVCSLRFSRDGEILASGSMDN 733
Query: 571 SARIWSA 577
+ R+W A
Sbjct: 734 TVRLWDA 740
Score = 64.3 bits (155), Expect = 1e-09
Identities = 58/208 (27%), Positives = 86/208 (40%), Gaps = 12/208 (5%)
Query: 460 LGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQ 516
L IL GH V F P R LL SS+ V RLW++ C+ +G + V +F
Sbjct: 536 LKILYGHSGPVYGASFSPDRNYLLSSSEDGTV-RLWSLQTFTCLVGYKGHNYPVWDTQFS 594
Query: 517 PQYGQLLATAIGNSITIIDVETDSHLHDLK---GHDKDVLSICWDRSGNFLASVSED-SA 572
P YG + G + + H L+ GH DV + + N++A+ S D +
Sbjct: 595 P-YGYYFVS--GGHDRVARLWATDHYQPLRIFAGHLADVNCTRFHPNSNYVATGSADRTV 651
Query: 573 RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
R+W + G C+ S F P R L + W G + H
Sbjct: 652 RLWDVLN-GNCVRIFTGHKGPIHSLTFSPNGRFLATGATDGRVLLWDIGHGLMVGELKGH 710
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVW 660
+ L S GE++AS S DN+V++W
Sbjct: 711 TDTVCSLRFSRDGEILASGSMDNTVRLW 738
Score = 45.4 bits (106), Expect = 5e-04
Identities = 56/244 (22%), Positives = 97/244 (38%), Gaps = 30/244 (12%)
Query: 427 ITDVRFQPGSTMFATSSFDRSVRLWDAS-NPIRSLGILTGHDEQVMSLDFHPRRMDLLCS 485
+T V S++ A D +VR+W + +RS+ +Q L L
Sbjct: 473 LTAVDVTDDSSLIAGGFADSTVRVWSVTPKKLRSV-------KQASDLS--------LID 517
Query: 486 SDSNDVI-RLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSH 541
+S+DV+ R+ + + + G S V F P LL+++ ++ + ++T +
Sbjct: 518 KESDDVLERIMDEKTASELKILYGHSGPVYGASFSPDRNYLLSSSEDGTVRLWSLQTFTC 577
Query: 542 LHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKC---IGELHSVGNKFQSC 597
L KGH+ V + G + S D AR+W+ G L V +C
Sbjct: 578 LVGYKGHNYPVWDTQFSPYGYYFVSGGHDRVARLWATDHYQPLRIFAGHLADV-----NC 632
Query: 598 I-FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNS 656
FHP + +++ W G+ H+G I L SP G +A+ + D
Sbjct: 633 TRFHPNSNYVATGSADRTVRLWDVLNGNCVRIFTGHKGPIHSLTFSPNGRFLATGATDGR 692
Query: 657 VKVW 660
V +W
Sbjct: 693 VLLW 696
Score = 43.1 bits (100), Expect = 0.002
Identities = 19/71 (26%), Positives = 35/71 (48%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS +G+ +A+ + +V +W++ + H+ + +RF + A+ S D +VR
Sbjct: 677 FSPNGRFLATGATDGRVLLWDIGHGLMVGELKGHTDTVCSLRFSRDGEILASGSMDNTVR 736
Query: 450 LWDASNPIRSL 460
LWDA L
Sbjct: 737 LWDAIKAFEDL 747
>TAF5_MOUSE (Q8C092) Transcription initiation factor TFIID subunit 5
(Transcription initiation factor TFIID 100 kDa subunit)
(TAF(II)100) (TAFII-100) (TAFII100)
Length = 801
Score = 84.3 bits (207), Expect = 9e-16
Identities = 66/247 (26%), Positives = 109/247 (43%), Gaps = 22/247 (8%)
Query: 336 SFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGK 395
S LS D+ +D+ +E + K S E+ L+ V FS D
Sbjct: 512 SDLSLIDKESDDVLERIMDEKTAS--------------ELKILYGHSGPVYGASFSPDRN 557
Query: 396 VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASN 455
+ S+ + V +W+++ F H++ + D +F P F + DR RLW A++
Sbjct: 558 YLLSSSEDGTVRLWSLQTFTCLVGYKGHNYPVWDTQFSPYGYYFVSGGHDRVARLW-ATD 616
Query: 456 PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHT---TRGGSKQ 512
+ L I GH V +HP + + + ++ +RLW+V C+ +G
Sbjct: 617 HYQPLRIFAGHLADVNCTRYHP-NSNYVATGSADRTVRLWDVLNGNCVRIFTGHKGPIHS 675
Query: 513 VRFQPQYGQLLAT-AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED- 570
+ F P G+ LAT A + + D+ + +LKGH V S+ + R G LAS S D
Sbjct: 676 LTFSPN-GRFLATGATDGRVLLWDIGHGLMVGELKGHTDTVCSLRFSRDGEILASGSMDN 734
Query: 571 SARIWSA 577
+ R+W A
Sbjct: 735 TVRLWDA 741
Score = 64.7 bits (156), Expect = 7e-10
Identities = 58/208 (27%), Positives = 86/208 (40%), Gaps = 12/208 (5%)
Query: 460 LGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQ 516
L IL GH V F P R LL SS+ V RLW++ C+ +G + V +F
Sbjct: 537 LKILYGHSGPVYGASFSPDRNYLLSSSEDGTV-RLWSLQTFTCLVGYKGHNYPVWDTQFS 595
Query: 517 PQYGQLLATAIGNSITIIDVETDSHLHDLK---GHDKDVLSICWDRSGNFLASVSED-SA 572
P YG + G + + H L+ GH DV + + N++A+ S D +
Sbjct: 596 P-YGYYFVS--GGHDRVARLWATDHYQPLRIFAGHLADVNCTRYHPNSNYVATGSADRTV 652
Query: 573 RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
R+W + G C+ S F P R L + W G + H
Sbjct: 653 RLWDVLN-GNCVRIFTGHKGPIHSLTFSPNGRFLATGATDGRVLLWDIGHGLMVGELKGH 711
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVW 660
+ L S GE++AS S DN+V++W
Sbjct: 712 TDTVCSLRFSRDGEILASGSMDNTVRLW 739
Score = 43.1 bits (100), Expect = 0.002
Identities = 19/71 (26%), Positives = 35/71 (48%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS +G+ +A+ + +V +W++ + H+ + +RF + A+ S D +VR
Sbjct: 678 FSPNGRFLATGATDGRVLLWDIGHGLMVGELKGHTDTVCSLRFSRDGEILASGSMDNTVR 737
Query: 450 LWDASNPIRSL 460
LWDA L
Sbjct: 738 LWDAVKAFEDL 748
>APAF_HUMAN (O14727) Apoptotic protease activating factor 1 (Apaf-1)
Length = 1248
Score = 84.3 bits (207), Expect = 9e-16
Identities = 69/297 (23%), Positives = 121/297 (40%), Gaps = 34/297 (11%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+ +AS G +K + ++ E + H + F AT S D+ V+
Sbjct: 623 FSEDGQRIASCGADKTLQVFKAETGEKLLEIKAHEDEVLCCAFSTDDRFIATCSVDKKVK 682
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV-IRLWNVNQSECMHTTRG 508
+W++ + H EQV F LL ++ S+D ++LW++NQ EC +T G
Sbjct: 683 IWNSMTG-ELVHTYDEHSEQVNCCHFTNSSHHLLLATGSSDCFLKLWDLNQKECRNTMFG 741
Query: 509 GSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLK-----------GHDKDVLS 554
+ V RF P L + + ++ + D + + + D +V+
Sbjct: 742 HTNSVNHCRFSPDDKLLASCSADGTLKLWDATSANERKSINVKQFFLNLEDPQEDMEVIV 801
Query: 555 ICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVG----------NKFQSCIFHPGYR 604
C S S D ARI AA + + ++H+ G + Q C F P
Sbjct: 802 KC--------CSWSADGARIMVAAKNKIFLFDIHTSGLLGEIHTGHHSTIQYCDFSPQNH 853
Query: 605 SLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
++ +E W+ SK H + G+ SP G ++S D ++++W+
Sbjct: 854 LAVVALSQYCVELWNTDSRSKVADCRGHLSWVHGVMFSPDGSSFLTSSDDQTIRLWE 910
Score = 56.6 bits (135), Expect = 2e-07
Identities = 59/278 (21%), Positives = 110/278 (39%), Gaps = 16/278 (5%)
Query: 388 CHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRS 447
C FS + A + V +WN ++ A H + V F P + F TSS D++
Sbjct: 846 CDFSPQNHLAVVALSQYCVELWNTDSRSKVADCRGHLSWVHGVMFSPDGSSFLTSSDDQT 905
Query: 448 VRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTR 507
+RLW+ ++ ++ +Q + + F + ++ + D ++L N + + T
Sbjct: 906 IRLWETKKVCKNSAVML---KQEVDVVFQENEV-MVLAVDHIRRLQLINGRTGQIDYLTE 961
Query: 508 GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASV 567
P + +I I+++ + H K V I + L S
Sbjct: 962 AQVSCCCLSPHLQYIAFGDENGAIEILELVNNRIFQSRFQHKKTVWHIQFTADEKTLISS 1021
Query: 568 SEDS-ARIWSAASDGKCI---GELHSVGNKFQSCIFHPGYRSLLIIGGYQ-SLEAWSPTE 622
S+D+ ++W+ D KCI G +V + F S L+ + +++ W+
Sbjct: 1022 SDDAEIQVWNWQLD-KCIFLRGHQETVKD------FRLLKNSRLLSWSFDGTVKVWNIIT 1074
Query: 623 GSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G+K HQG + S +S S D + K+W
Sbjct: 1075 GNKEKDFVCHQGTVLSCDISHDATKFSSTSADKTAKIW 1112
Score = 45.1 bits (105), Expect = 6e-04
Identities = 36/146 (24%), Positives = 57/146 (38%), Gaps = 15/146 (10%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
VL+C S D +S +K IW+ + H+ + F ST+ AT
Sbjct: 1088 VLSCDISHDATKFSSTSADKTAKIWSFDLLLPLHELRGHNGCVRCSAFSVDSTLLATGDD 1147
Query: 445 DRSVRLWDASN--------PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWN 496
+ +R+W+ SN P+ G T H V L F P L+ + I+ WN
Sbjct: 1148 NGEIRIWNVSNGELLHLCAPLSEEGAAT-HGGWVTDLCFSPDGKMLI---SAGGYIKWWN 1203
Query: 497 V---NQSECMHTTRGGSKQVRFQPQY 519
V S+ +T K++ P +
Sbjct: 1204 VVTGESSQTFYTNGTNLKKIHVSPDF 1229
>PF20_CHLRE (P93107) Flagellar WD-repeat protein PF20
Length = 606
Score = 83.2 bits (204), Expect = 2e-15
Identities = 59/292 (20%), Positives = 116/292 (39%), Gaps = 15/292 (5%)
Query: 381 SINKVLTCHFSSDGK--------VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
S+NK H S ++ +A +K +W+M + H + V F
Sbjct: 317 SLNKTFKGHLLSVANLALHPTKPILVTASDDKTWKMWHMPGGDLIMCGEGHKDWVAGVDF 376
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P T A+ D +V++WD R + T H + + S+ FH +++ S + +
Sbjct: 377 HPAGTCLASGGGDSAVKIWDFEKQ-RCVTTFTDHKQAIWSVRFH-HLGEVVASGSLDHTV 434
Query: 493 RLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
RLW++ +C RG V +QP L + ++++ D GH
Sbjct: 435 RLWDLPAGKCRMALRGHVDSVNDLAWQPFSSSLATASSDKTVSVWDARAGLCTQTYYGHQ 494
Query: 550 KDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLI 608
+ ++ G LAS D ++W + + +++ + F + L +
Sbjct: 495 NSCNGVSFNILGTQLASTDADGVVKLWDTRMTAE-VATINTGKHPANKSCFDRSGQVLAV 553
Query: 609 IGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++A+S T+G +A H+ + + P G+ + S DN+ ++W
Sbjct: 554 ACDDGKVKAYSTTDGVLQAELAGHEDAVQAVLFDPAGQYLVSCGSDNTFRLW 605
Score = 75.1 bits (183), Expect = 5e-13
Identities = 60/233 (25%), Positives = 92/233 (38%), Gaps = 5/233 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F G +AS G + V IW+ E + + H I VRF + A+ S D +VR
Sbjct: 376 FHPAGTCLASGGGDSAVKIWDFEKQRCVTTFTDHKQAIWSVRFHHLGEVVASGSLDHTVR 435
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD + L GH + V L + P L +S S+ + +W+ C T G
Sbjct: 436 LWDLPAG-KCRMALRGHVDSVNDLAWQPFSSSLATAS-SDKTVSVWDARAGLCTQTYYGH 493
Query: 510 SKQ---VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
V F QL +T + + D + + + C+DRSG LA
Sbjct: 494 QNSCNGVSFNILGTQLASTDADGVVKLWDTRMTAEVATINTGKHPANKSCFDRSGQVLAV 553
Query: 567 VSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
+D + +DG EL + Q+ +F P + L+ G + WS
Sbjct: 554 ACDDGKVKAYSTTDGVLQAELAGHEDAVQAVLFDPAGQYLVSCGSDNTFRLWS 606
Score = 68.6 bits (166), Expect = 5e-11
Identities = 53/200 (26%), Positives = 94/200 (46%), Gaps = 14/200 (7%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
+ + F G+V+AS + V +W++ + + H + D+ +QP S+ AT+S
Sbjct: 413 IWSVRFHHLGEVVASGSLDHTVRLWDLPAGKCRMALRGHVDSVNDLAWQPFSSSLATASS 472
Query: 445 DRSVRLWDASNPIRSLGILT----GHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQS 500
D++V +WDA G+ T GH + F+ L S+D++ V++LW+ +
Sbjct: 473 DKTVSVWDA-----RAGLCTQTYYGHQNSCNGVSFNILGTQ-LASTDADGVVKLWDTRMT 526
Query: 501 ECMHTTRGGSKQVRFQ--PQYGQLLATAIGNSITIIDVETDSHLH-DLKGHDKDVLSICW 557
+ T G + GQ+LA A + TD L +L GH+ V ++ +
Sbjct: 527 AEVATINTGKHPANKSCFDRSGQVLAVACDDGKVKAYSTTDGVLQAELAGHEDAVQAVLF 586
Query: 558 DRSGNFLASVSEDSA-RIWS 576
D +G +L S D+ R+WS
Sbjct: 587 DPAGQYLVSCGSDNTFRLWS 606
>WDS_DROME (Q9V3J8) Will die slowly protein
Length = 361
Score = 81.3 bits (199), Expect = 8e-15
Identities = 63/246 (25%), Positives = 112/246 (44%), Gaps = 11/246 (4%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H+ ++ V+F P A+SS D+ +++W A + + ++GH + + + L
Sbjct: 71 HTKAVSAVKFSPNGEWLASSSADKLIKIWGAYDG-KFEKTISGHKLGISDVAWSSDSR-L 128
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETD 539
L S + +++W ++ + + T +G S V F PQ +++ + S+ I DV T
Sbjct: 129 LVSGSDDKTLKVWELSTGKSLKTLKGHSNYVFCCNFNPQSNLIVSGSFDESVRIWDVRTG 188
Query: 540 SHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCI 598
L L H V ++ ++R G+ + S S D RIW AS G+C+ L N S +
Sbjct: 189 KCLKTLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTAS-GQCLKTLIDDDNPPVSFV 247
Query: 599 -FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQG---LIAGLADSPVGELIASASHD 654
F P + +L +L+ W ++G + H+ I G+ I S S D
Sbjct: 248 KFSPNGKYILAATLDNTLKLWDYSKGKCLKTYTGHKNEKYCIFANFSVTGGKWIVSGSED 307
Query: 655 NSVKVW 660
N V +W
Sbjct: 308 NMVYIW 313
Score = 79.7 bits (195), Expect = 2e-14
Identities = 66/283 (23%), Positives = 132/283 (46%), Gaps = 15/283 (5%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS +G+ +AS+ +K + IW + ++ + H I+DV + S + + S D++++
Sbjct: 80 FSPNGEWLASSSADKLIKIWGAYDGKFEKTISGHKLGISDVAWSSDSRLLVSGSDDKTLK 139
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
+W+ S +SL L GH V +F+P + +L+ S ++ +R+W+V +C+ T
Sbjct: 140 VWELSTG-KSLKTLKGHSNYVFCCNFNP-QSNLIVSGSFDESVRIWDVRTGKCLKTLPAH 197
Query: 510 S---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLS-ICWDRSGNF-L 564
S V F +++++ I D + L L D +S + + +G + L
Sbjct: 198 SDPVSAVHFNRDGSLIVSSSYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPNGKYIL 257
Query: 565 ASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFH----PGYRSLLIIGGYQSLEAWSP 620
A+ +++ ++W S GKC+ N+ + CIF G + ++ + W+
Sbjct: 258 AATLDNTLKLWD-YSKGKCLKTYTGHKNE-KYCIFANFSVTGGKWIVSGSEDNMVYIWNL 315
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASAS--HDNSVKVWK 661
+ H + A P +IASA+ +D ++K+WK
Sbjct: 316 QSKEVVQKLQGHTDTVLCTACHPTENIIASAALENDKTIKLWK 358
Score = 64.3 bits (155), Expect = 1e-09
Identities = 41/204 (20%), Positives = 95/204 (46%), Gaps = 10/204 (4%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V C+F+ ++ S ++ V IW++ + + HS ++ V F ++ +S
Sbjct: 157 NYVFCCNFNPQSNLIVSGSFDESVRIWDVRTGKCLKTLPAHSDPVSAVHFNRDGSLIVSS 216
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S+D R+WD ++ ++ + V + F P +L ++ N ++LW+ ++ +C
Sbjct: 217 SYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPNGKYILAATLDN-TLKLWDYSKGKC 275
Query: 503 MHTTRGGSKQ-----VRFQPQYGQ-LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC 556
+ T G + F G+ +++ + N + I ++++ + L+GH VL
Sbjct: 276 LKTYTGHKNEKYCIFANFSVTGGKWIVSGSEDNMVYIWNLQSKEVVQKLQGHTDTVLCTA 335
Query: 557 WDRSGNFLASV---SEDSARIWSA 577
+ N +AS ++ + ++W +
Sbjct: 336 CHPTENIIASAALENDKTIKLWKS 359
Score = 45.1 bits (105), Expect = 6e-04
Identities = 36/119 (30%), Positives = 55/119 (45%), Gaps = 6/119 (5%)
Query: 545 LKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
L GH K V ++ + +G +LAS S D +IW A DGK E G+K
Sbjct: 68 LAGHTKAVSAVKFSPNGEWLASSSADKLIKIW-GAYDGKF--EKTISGHKLGISDVAWSS 124
Query: 604 RSLLIIGGY--QSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
S L++ G ++L+ W + G ++ H + +P LI S S D SV++W
Sbjct: 125 DSRLLVSGSDDKTLKVWELSTGKSLKTLKGHSNYVFCCNFNPQSNLIVSGSFDESVRIW 183
Score = 31.2 bits (69), Expect = 9.1
Identities = 12/34 (35%), Positives = 22/34 (64%)
Query: 627 WSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++A H ++ + SP GE +AS+S D +K+W
Sbjct: 66 FTLAGHTKAVSAVKFSPNGEWLASSSADKLIKIW 99
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 82,476,591
Number of Sequences: 164201
Number of extensions: 3649187
Number of successful extensions: 12619
Number of sequences better than 10.0: 501
Number of HSP's better than 10.0 without gapping: 248
Number of HSP's successfully gapped in prelim test: 254
Number of HSP's that attempted gapping in prelim test: 9252
Number of HSP's gapped (non-prelim): 1935
length of query: 661
length of database: 59,974,054
effective HSP length: 117
effective length of query: 544
effective length of database: 40,762,537
effective search space: 22174820128
effective search space used: 22174820128
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0114.1


