

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.7
(147 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
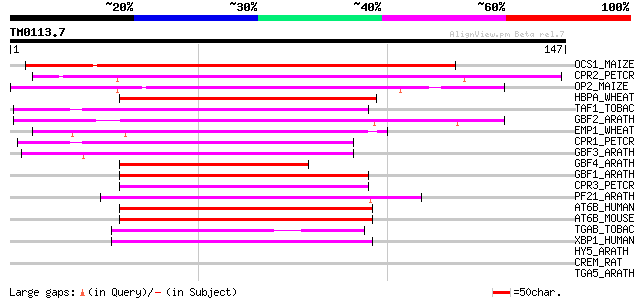
Score E
Sequences producing significant alignments: (bits) Value
OCS1_MAIZE (P24068) Ocs-element binding factor 1 (OCSBF-1) 85 6e-17
CPR2_PETCR (Q99090) Light-inducible protein CPRF-2 77 1e-14
OP2_MAIZE (P12959) Opaque-2 regulatory protein 63 3e-10
HBPA_WHEAT (P23922) Transcription factor HBP-1a (Histone-specifi... 57 1e-08
TAF1_TOBAC (Q99142) Transcriptional activator TAF-1 (Fragment) 54 1e-07
GBF2_ARATH (P42775) G-box binding factor 2 (AtbZIP54) 52 4e-07
EMP1_WHEAT (P25032) DNA-binding EMBP-1 protein 52 4e-07
CPR1_PETCR (Q99089) Common plant regulatory factor CPRF-1 51 1e-06
GBF3_ARATH (P42776) G-box binding factor 3 (AtbZIP55) 50 1e-06
GBF4_ARATH (P42777) G-box binding factor 4 (AtbZIP40) 48 7e-06
GBF1_ARATH (P42774) G-box binding factor 1 (AtbZIP41) 48 9e-06
CPR3_PETCR (Q99091) Light-inducible protein CPRF-3 45 4e-05
PF21_ARATH (Q04088) Possible transcription factor PosF21 (AtbZIP59) 45 6e-05
AT6B_HUMAN (Q99941) Cyclic-AMP-dependent transcription factor AT... 44 1e-04
AT6B_MOUSE (O35451) Cyclic-AMP-dependent transcription factor AT... 44 1e-04
TGAB_TOBAC (P14233) TGACG-sequence specific DNA-binding protein ... 42 5e-04
XBP1_HUMAN (P17861) X box binding protein-1 (XBP-1) (TREB5 protein) 42 6e-04
HY5_ARATH (O24646) Transcription factor HY5 (LONG HYPOCOL5 prote... 40 0.002
CREM_RAT (Q03061) cAMP responsive element modulator 40 0.002
TGA5_ARATH (Q39163) Transcription factor TGA5 (Ocs-element bindi... 40 0.002
>OCS1_MAIZE (P24068) Ocs-element binding factor 1 (OCSBF-1)
Length = 151
Score = 84.7 bits (208), Expect = 6e-17
Identities = 46/114 (40%), Positives = 72/114 (62%), Gaps = 1/114 (0%)
Query: 5 SGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLR 64
S +S + + SGS+ + A R+ KR +SNRESARRSR+RKQ+HLDEL +VA+L+
Sbjct: 2 SSSSLSPTAGRTSGSDGD-SAADTHRREKRRLSNRESARRSRLRKQQHLDELVQEVARLQ 60
Query: 65 SENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGV 118
++N ++ ++ ++ EN+VL A+ EL RL S+NE++ V +GV
Sbjct: 61 ADNARVAARARDIASQYTRVEQENTVLRARAAELGDRLRSVNEVLRLVEEFSGV 114
>CPR2_PETCR (Q99090) Light-inducible protein CPRF-2
Length = 401
Score = 77.4 bits (189), Expect = 1e-14
Identities = 53/150 (35%), Positives = 79/150 (52%), Gaps = 11/150 (7%)
Query: 7 TSSGSSMIQNSGSEEELQALM---------DQRKRKRMISNRESARRSRMRKQKHLDELA 57
T+SGSS +S ++EL+ D ++ +RM+SNRESARRSR RKQ H+ EL
Sbjct: 169 TTSGSSR-DHSDDDDELEGETETTRNGDPSDAKRVRRMLSNRESARRSRRRKQAHMTELE 227
Query: 58 AQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNG 117
QV+QLR EN +L L +QR+ +N VL A + + +++ E + V N
Sbjct: 228 TQVSQLRVENSSLLKRLTDISQRYNDAAVDNRVLKADIETMRAKVKMAEETVKRVTGLNP 287
Query: 118 VF-AANSFVDPIPSAYYLNHPIMASADILQ 146
+F + +S + I + P SAD Q
Sbjct: 288 MFQSMSSEISTIGMQSFSGSPSDTSADTTQ 317
>OP2_MAIZE (P12959) Opaque-2 regulatory protein
Length = 453
Score = 62.8 bits (151), Expect = 3e-10
Identities = 47/138 (34%), Positives = 74/138 (53%), Gaps = 11/138 (7%)
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALM---DQRKRKRMISNRESARRSRMRKQKHLDELA 57
+A+SS + S ++ E E+ ++R RK+ SNRESARRSR RK HL EL
Sbjct: 196 LATSSSSRDPSPSDEDMDGEVEILGFKMPTEERVRKKE-SNRESARRSRYRKAAHLKELE 254
Query: 58 AQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRL----ESLNEIIDFVN 113
QVAQL++EN +L + Q++ + +N VL A M L ++ +SL +I+
Sbjct: 255 DQVAQLKAENSCLLRRIAALNQKYNDANVDNRVLRADMETLRAKVKMGEDSLKRVIEM-- 312
Query: 114 ATNGVFAANSFVDPIPSA 131
++ V ++ P PS+
Sbjct: 313 -SSSVPSSMPISAPTPSS 329
>HBPA_WHEAT (P23922) Transcription factor HBP-1a (Histone-specific
transcription factor HBP1)
Length = 349
Score = 57.4 bits (137), Expect = 1e-08
Identities = 28/68 (41%), Positives = 45/68 (66%)
Query: 30 RKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENS 89
+K+KR +SNRESARRSR+RKQ +EL + L+SEN + L+ + + + ++N+
Sbjct: 254 KKQKRKLSNRESARRSRLRKQAECEELGQRAEALKSENSSLRIELDRIKKEYEELLSKNT 313
Query: 90 VLSAQMGE 97
L A++GE
Sbjct: 314 SLKAKLGE 321
>TAF1_TOBAC (Q99142) Transcriptional activator TAF-1 (Fragment)
Length = 265
Score = 53.9 bits (128), Expect = 1e-07
Identities = 34/94 (36%), Positives = 51/94 (54%), Gaps = 3/94 (3%)
Query: 2 ASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVA 61
AS + S S + N E LQ + ++ KR SNRESARRSR+RKQ +ELA +V
Sbjct: 171 ASPTNVSQLSPALPN---EAWLQNERELKREKRKQSNRESARRSRLRKQAEAEELAIRVQ 227
Query: 62 QLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQM 95
L +EN + + +N + + EN+ L ++
Sbjct: 228 SLTAENMTLKSEINKLMENSEKLKLENAALMERL 261
>GBF2_ARATH (P42775) G-box binding factor 2 (AtbZIP54)
Length = 360
Score = 52.4 bits (124), Expect = 4e-07
Identities = 38/138 (27%), Positives = 69/138 (49%), Gaps = 14/138 (10%)
Query: 2 ASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVA 61
A S S+G + + +E+E+ ++ KR SNRESARRSR+RKQ ++L+ +V
Sbjct: 229 AMSFQNSAGMNGVPQPWNEKEV------KREKRKQSNRESARRSRLRKQAETEQLSVKVD 282
Query: 62 QLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQM-GELSHRLESLNEIIDFVNATNG--- 117
L +EN + + L + EN + Q+ + + + E+L +D N+ +G
Sbjct: 283 ALVAENMSLRSKLGQLNNESEKLRLENEAILDQLKAQATGKTENLISRVDKNNSVSGSKT 342
Query: 118 ----VFAANSFVDPIPSA 131
+ A+ DP+ ++
Sbjct: 343 VQHQLLNASPITDPVAAS 360
>EMP1_WHEAT (P25032) DNA-binding EMBP-1 protein
Length = 354
Score = 52.4 bits (124), Expect = 4e-07
Identities = 34/103 (33%), Positives = 60/103 (58%), Gaps = 11/103 (10%)
Query: 7 TSSGSSMIQ---NSGSEEELQALMDQ------RKRKRMISNRESARRSRMRKQKHLDELA 57
++S SS++Q N+ + + A + Q ++ +R SNRESARRSR+RKQ+ +ELA
Sbjct: 220 SASPSSLVQGEVNAAASSQSNASLSQMDERELKRERRKQSNRESARRSRLRKQQECEELA 279
Query: 58 AQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSH 100
+V++L + N + + L+ + ++ EN L ++ LSH
Sbjct: 280 QKVSELTAANGTLRSELDQLKKDCKTMETENKKLMGKI--LSH 320
>CPR1_PETCR (Q99089) Common plant regulatory factor CPRF-1
Length = 411
Score = 50.8 bits (120), Expect = 1e-06
Identities = 33/89 (37%), Positives = 48/89 (53%), Gaps = 3/89 (3%)
Query: 3 SSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQ 62
S +G S+M+ N + L D ++ +R SNRESARRSR+RKQ +ELA +V
Sbjct: 247 SPAGGQQPSTMMPN---DSWLHNDRDLKRERRKQSNRESARRSRLRKQAEAEELAIKVDS 303
Query: 63 LRSENQQMLTSLNLTTQRFMIIDAENSVL 91
L +EN + +N T + +NS L
Sbjct: 304 LTAENMALKAEINRLTLTAEKLTNDNSRL 332
>GBF3_ARATH (P42776) G-box binding factor 3 (AtbZIP55)
Length = 382
Score = 50.4 bits (119), Expect = 1e-06
Identities = 32/93 (34%), Positives = 49/93 (52%), Gaps = 5/93 (5%)
Query: 4 SSGTSSGSSMIQNSG-----SEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAA 58
S G S+ S+ + E LQ + ++ +R SNRESARRSR+RKQ +ELA
Sbjct: 230 SPGVSANSNPFMSQSLAMVPPETWLQNERELKRERRKQSNRESARRSRLRKQAETEELAR 289
Query: 59 QVAQLRSENQQMLTSLNLTTQRFMIIDAENSVL 91
+V L +EN + + LN ++ + N+ L
Sbjct: 290 KVEALTAENMALRSELNQLNEKSDKLRGANATL 322
>GBF4_ARATH (P42777) G-box binding factor 4 (AtbZIP40)
Length = 270
Score = 48.1 bits (113), Expect = 7e-06
Identities = 24/50 (48%), Positives = 34/50 (68%)
Query: 30 RKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQ 79
+++KRMI NRESA RSR RKQ + EL A+L EN+Q+L + +T+
Sbjct: 189 QRQKRMIKNRESAARSRERKQAYQVELETLAAKLEEENEQLLKEIEESTK 238
>GBF1_ARATH (P42774) G-box binding factor 1 (AtbZIP41)
Length = 315
Score = 47.8 bits (112), Expect = 9e-06
Identities = 24/66 (36%), Positives = 40/66 (60%)
Query: 30 RKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENS 89
+++KR SNRESARRSR+RKQ ++L +V L +ENQ + L + + +EN+
Sbjct: 224 KRQKRKQSNRESARRSRLRKQAECEQLQQRVESLSNENQSLRDELQRLSSECDKLKSENN 283
Query: 90 VLSAQM 95
+ ++
Sbjct: 284 SIQDEL 289
>CPR3_PETCR (Q99091) Light-inducible protein CPRF-3
Length = 296
Score = 45.4 bits (106), Expect = 4e-05
Identities = 23/66 (34%), Positives = 40/66 (59%)
Query: 30 RKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENS 89
++++R SNRESARRSR+RKQ DEL ++ L EN+ + +L ++ + +EN
Sbjct: 198 KRQRRKQSNRESARRSRLRKQAKSDELQERLDNLSKENRILRKNLQRISEACAEVTSENH 257
Query: 90 VLSAQM 95
+ ++
Sbjct: 258 SIKEEL 263
>PF21_ARATH (Q04088) Possible transcription factor PosF21 (AtbZIP59)
Length = 398
Score = 45.1 bits (105), Expect = 6e-05
Identities = 28/88 (31%), Positives = 49/88 (54%), Gaps = 3/88 (3%)
Query: 25 ALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMII 84
AL+D ++ KR+ +NR+SA RS+ RK +++ EL +V L++E + L L + +
Sbjct: 198 ALIDPKRAKRIWANRQSAARSKERKTRYIFELERKVQTLQTEATTLSAQLTLLQRDTNGL 257
Query: 85 DAENSVLSAQ---MGELSHRLESLNEII 109
EN+ L + M + H + LNE +
Sbjct: 258 TVENNELKLRLQTMEQQVHLQDELNEAL 285
>AT6B_HUMAN (Q99941) Cyclic-AMP-dependent transcription factor ATF-6
beta (Activating transcription factor 6 beta)
(ATF6-beta) (cAMP responsive element binding
protein-like 1) (cAMP response element binding
protein-related protein) (Creb-rp) (G13 protein)
Length = 703
Score = 44.3 bits (103), Expect = 1e-04
Identities = 24/67 (35%), Positives = 41/67 (60%)
Query: 30 RKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENS 89
++++RMI NRESA +SR +K+++L L A++ + ++NQQ+ +R + AENS
Sbjct: 327 KRQQRMIKNRESACQSRRKKKEYLQGLEARLQAVLADNQQLRRENAALRRRLEALLAENS 386
Query: 90 VLSAQMG 96
L G
Sbjct: 387 ELKLGSG 393
>AT6B_MOUSE (O35451) Cyclic-AMP-dependent transcription factor ATF-6
beta (Activating transcription factor 6 beta)
(ATF6-beta) (cAMP responsive element binding
protein-like 1) (cAMP response element binding
protein-related protein) (Creb-rp)
Length = 699
Score = 43.9 bits (102), Expect = 1e-04
Identities = 24/67 (35%), Positives = 41/67 (60%)
Query: 30 RKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAENS 89
++++RMI NRESA +SR +K+++L L A++ + ++NQQ+ +R + AENS
Sbjct: 324 KRQQRMIKNRESACQSRRKKKEYLQGLEARLQAVLADNQQLRRENAALRRRLEALLAENS 383
Query: 90 VLSAQMG 96
L G
Sbjct: 384 GLKLGSG 390
>TGAB_TOBAC (P14233) TGACG-sequence specific DNA-binding protein
TGA-1B (TGA1b) (HSBF) (Fragment)
Length = 242
Score = 42.0 bits (97), Expect = 5e-04
Identities = 24/67 (35%), Positives = 38/67 (55%), Gaps = 7/67 (10%)
Query: 28 DQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAE 87
D++KR R++ NRESA+ SR RK+ +++EL +V + S Q + + I AE
Sbjct: 183 DEKKRARLVRNRESAQLSRQRKKHYVEELEDKVRIMHSTIQDL-------NAKVAYIIAE 235
Query: 88 NSVLSAQ 94
N+ L Q
Sbjct: 236 NATLKTQ 242
>XBP1_HUMAN (P17861) X box binding protein-1 (XBP-1) (TREB5 protein)
Length = 260
Score = 41.6 bits (96), Expect = 6e-04
Identities = 21/69 (30%), Positives = 38/69 (54%)
Query: 28 DQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAE 87
+++ +R + NR +A+ +R RK+ + EL QV L ENQ++L L ++ + E
Sbjct: 69 EEKALRRKLKNRVAAQTARDRKKARMSELEQQVVDLEEENQKLLLENQLLREKTHGLVVE 128
Query: 88 NSVLSAQMG 96
N L ++G
Sbjct: 129 NQELRQRLG 137
>HY5_ARATH (O24646) Transcription factor HY5 (LONG HYPOCOL5 protein)
(AtbZIP56)
Length = 168
Score = 40.0 bits (92), Expect = 0.002
Identities = 25/92 (27%), Positives = 49/92 (53%), Gaps = 12/92 (13%)
Query: 5 SGTSSGSSMIQNSGSEEELQ-----ALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
SG+++G Q + E + + A + ++ KR++ NR SA+++R RK+ +L EL +
Sbjct: 60 SGSATGQERTQATVGESQRKRGRTPAEKENKRLKRLLRNRVSAQQARERKKAYLSELENR 119
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVL 91
V L ++N ++ +R + EN +L
Sbjct: 120 VKDLENKNSEL-------EERLSTLQNENQML 144
>CREM_RAT (Q03061) cAMP responsive element modulator
Length = 341
Score = 40.0 bits (92), Expect = 0.002
Identities = 24/76 (31%), Positives = 42/76 (54%), Gaps = 3/76 (3%)
Query: 2 ASSSGTSSGSSMIQNSGSEEELQALMDQRKRKR---MISNRESARRSRMRKQKHLDELAA 58
A ++ G M + GS Q L ++ RKR ++ NRE+AR R +K++++ L
Sbjct: 254 APTTALPQGVVMAASPGSLHSPQQLAEEATRKRELRLMKNREAARECRRKKKEYVKCLEN 313
Query: 59 QVAQLRSENQQMLTSL 74
+VA L S+N+ ++ L
Sbjct: 314 RVAVLESQNKTLIEEL 329
>TGA5_ARATH (Q39163) Transcription factor TGA5 (Ocs-element
binding factor 5) (OBF5) (AtbZIP26)
Length = 330
Score = 39.7 bits (91), Expect = 0.002
Identities = 20/56 (35%), Positives = 36/56 (63%), Gaps = 3/56 (5%)
Query: 17 SGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDEL---AAQVAQLRSENQQ 69
S S + ++ MDQ+ +R+ NRE+AR+SR+RK+ ++ +L ++ QL E Q+
Sbjct: 33 SDSSDRSKSKMDQKTLRRLAQNREAARKSRLRKKAYVQQLENSRLKLTQLEQELQR 88
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.124 0.319
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,923,941
Number of Sequences: 164201
Number of extensions: 411860
Number of successful extensions: 2622
Number of sequences better than 10.0: 272
Number of HSP's better than 10.0 without gapping: 119
Number of HSP's successfully gapped in prelim test: 153
Number of HSP's that attempted gapping in prelim test: 2409
Number of HSP's gapped (non-prelim): 359
length of query: 147
length of database: 59,974,054
effective HSP length: 100
effective length of query: 47
effective length of database: 43,553,954
effective search space: 2047035838
effective search space used: 2047035838
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0113.7


