

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
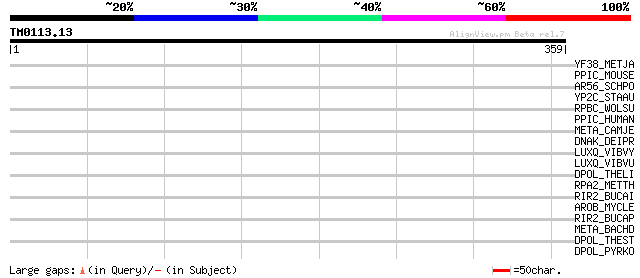
Query= TM0113.13
(359 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YF38_METJA (Q58933) Hypothetical protein MJ1538 34 0.50
PPIC_MOUSE (P30412) Peptidyl-prolyl cis-trans isomerase C (EC 5.... 32 1.9
AR56_SCHPO (P31318) Arg11 protein, mitochondrial precursor [Cont... 32 1.9
YP2C_STAAU (P14503) Hypothetical 27.7 kDa protein 32 3.2
RPBC_WOLSU (Q7MA56) Bifunctional DNA-directed RNA polymerase, be... 32 3.2
PPIC_HUMAN (P45877) Peptidyl-prolyl cis-trans isomerase C (EC 5.... 32 3.2
META_CAMJE (Q9PLV2) Homoserine O-succinyltransferase (EC 2.3.1.4... 32 3.2
DNAK_DEIPR (P94695) Chaperone protein dnaK (Heat shock protein 7... 31 4.2
LUXQ_VIBVY (Q7MD16) Autoinducer 2 sensor kinase/phosphatase luxQ... 31 5.5
LUXQ_VIBVU (Q8D5Z6) Autoinducer 2 sensor kinase/phosphatase luxQ... 31 5.5
DPOL_THELI (P30317) DNA polymerase (EC 2.7.7.7) (Vent DNA polyme... 31 5.5
RPA2_METTH (O27126) DNA-directed RNA polymerase subunit A'' (EC ... 30 7.2
RIR2_BUCAI (P57275) Ribonucleoside-diphosphate reductase beta ch... 30 7.2
AROB_MYCLE (Q9CCS4) 3-dehydroquinate synthase (EC 4.2.3.4) 30 7.2
RIR2_BUCAP (Q8K9W4) Ribonucleoside-diphosphate reductase beta ch... 30 9.4
META_BACHD (Q9KAK7) Homoserine O-succinyltransferase (EC 2.3.1.4... 30 9.4
DPOL_THEST (O33845) DNA polymerase (EC 2.7.7.7) 30 9.4
DPOL_PYRKO (P77933) DNA polymerase (EC 2.7.7.7) [Contains: Endon... 30 9.4
>YF38_METJA (Q58933) Hypothetical protein MJ1538
Length = 252
Score = 34.3 bits (77), Expect = 0.50
Identities = 24/74 (32%), Positives = 43/74 (57%), Gaps = 5/74 (6%)
Query: 271 EGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELF 330
E S +I + + + KN N+SD ID++TR + V++YI+ ++ + K + E + EL
Sbjct: 169 EKSKEIPKFYVLEENKNKNNNISDKIDKETR-KIVSEYIK--SKKLDKDKIKEVV--ELR 223
Query: 331 RRFKNKIVKLHGVE 344
+ F KI K+ V+
Sbjct: 224 KEFLKKIKKMEEVD 237
>PPIC_MOUSE (P30412) Peptidyl-prolyl cis-trans isomerase C (EC
5.2.1.8) (PPIase) (Rotamase) (Cyclophilin C)
Length = 212
Score = 32.3 bits (72), Expect = 1.9
Identities = 36/147 (24%), Positives = 55/147 (36%), Gaps = 25/147 (17%)
Query: 51 CLKVADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLP 110
CL + L SSG + V + D V ++ + + LFGN K++
Sbjct: 14 CLGLGALVSSSGSSG--VRKRGPSVTDKVFFDVRIGDKDVGRIVIGLFGN---VVPKTVE 68
Query: 111 DFYKRLQQEKGHKFGPCFF-----------------SGTPG-SFYGRLFPNDSVHFFHSS 152
+F EKG+ + F GT G S YG FP+++ H
Sbjct: 69 NFVALATGEKGYGYKGSIFHRVIKDFMIQGGDFTARDGTGGMSIYGETFPDENFKLKH-- 126
Query: 153 YSLHWLSKVNYTKPLNKGAIYLTKTSP 179
Y + W+S N N ++T T P
Sbjct: 127 YGIGWVSMANAGPDTNGSQFFITLTKP 153
>AR56_SCHPO (P31318) Arg11 protein, mitochondrial precursor
[Contains: N-acetyl-gamma-glutamyl-phosphate reductase
(EC 1.2.1.38) (N-acetyl-glutamate semialdehyde
dehydrogenase) (NAGSA dehydrogenase); Acetylglutamate
kinase (EC 2.7.2.8) (NAG kinase) (AGK)
Length = 885
Score = 32.3 bits (72), Expect = 1.9
Identities = 37/149 (24%), Positives = 58/149 (38%), Gaps = 25/149 (16%)
Query: 33 KHILEDSIIRLYCATFPSCLKVADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQF 92
K L+D +R+Y +F + + D SS P ++ A N
Sbjct: 383 KDTLKDRKLRIYSDSFNESVAIVDTTDSSLP--------VLLAFGAADNN---------- 424
Query: 93 FLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSS 152
+LN++ + T P RLQ K FFS + G+ LF N+ +F++
Sbjct: 425 WLNNVVDSILTTLKADFPSLLWRLQPSA--KNLEWFFSKSEGT----LFANNFYYFWYGV 478
Query: 153 YSLHWLSK-VNYTKPLNKGAIYLTKTSPP 180
L+ +SK + KP I T PP
Sbjct: 479 KDLNKISKFIQSDKPFADAIIQTQSTKPP 507
>YP2C_STAAU (P14503) Hypothetical 27.7 kDa protein
Length = 236
Score = 31.6 bits (70), Expect = 3.2
Identities = 27/92 (29%), Positives = 47/92 (50%), Gaps = 5/92 (5%)
Query: 241 ENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDT 300
+N E KL+ N+ + E+ Q E + ++RLE +++ ++V ++ +
Sbjct: 51 KNENESEKLE--NLKNRIEELEKSEQDRNERTNILMKRLEASTSNFNNKLDVFEEKTKHA 108
Query: 301 RA--EAVAK-YIRAVAESILKSEFGEAIMDEL 329
R E AK YI+ + E LK +F +AI DEL
Sbjct: 109 RLNFETTAKHYIKRLDEDNLKLDFQQAIQDEL 140
>RPBC_WOLSU (Q7MA56) Bifunctional DNA-directed RNA polymerase, beta
and beta' chain (EC 2.7.7.6) [Includes: DNA-directed RNA
polymerase beta chain (Transcriptase beta chain) (RNA
polymerase beta subunit); DNA-directed RNA polymerase
beta' chain (Transcrip
Length = 2883
Score = 31.6 bits (70), Expect = 3.2
Identities = 16/43 (37%), Positives = 27/43 (62%), Gaps = 2/43 (4%)
Query: 310 RAVAESILKSEFGEAIMDELFRRFKNKIVKLH--GVEKLELAN 350
R +AE I+ E GE I+D L + + K+ K+H G+++ +AN
Sbjct: 297 RHLAEPIVDKESGEVILDTLVQLDEGKLKKIHELGIKEFTIAN 339
>PPIC_HUMAN (P45877) Peptidyl-prolyl cis-trans isomerase C (EC
5.2.1.8) (PPIase) (Rotamase) (Cyclophilin C)
Length = 212
Score = 31.6 bits (70), Expect = 3.2
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 131 GTPG-SFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSP 179
GT G S YG FP+++ H Y + W+S N N ++T T P
Sbjct: 106 GTGGVSIYGETFPDENFKLKH--YGIGWVSMANAGPDTNGSQFFITLTKP 153
>META_CAMJE (Q9PLV2) Homoserine O-succinyltransferase (EC 2.3.1.46)
(Homoserine O-transsuccinylase) (HTS)
Length = 293
Score = 31.6 bits (70), Expect = 3.2
Identities = 14/31 (45%), Positives = 17/31 (54%)
Query: 103 NTTFKSLPDFYKRLQQEKGHKFGPCFFSGTP 133
NT F L FYK L++ K HKF +G P
Sbjct: 78 NTPFTHLEKFYKGLEEVKKHKFDGAIVTGAP 108
>DNAK_DEIPR (P94695) Chaperone protein dnaK (Heat shock protein 70)
(Heat shock 70 kDa protein) (HSP70) (Fragment)
Length = 618
Score = 31.2 bits (69), Expect = 4.2
Identities = 33/121 (27%), Positives = 50/121 (41%), Gaps = 17/121 (14%)
Query: 207 PGGAMVITLIGRDENNELINAWVVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQ 266
PGG++ I + G+D E ++A V+ +V N AS L +K +P+Y +
Sbjct: 83 PGGSVRIEVDGKDYAPEQVSAEVLRKLVAN--ASAKLGQKITDAVITVPAYFDNS----- 135
Query: 267 VIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIM 326
QR T + I +NV I+E T A R E+IL + G
Sbjct: 136 ----------QREATKQAGEIAGLNVLRVINEPTAAALAYGLERKGDETILVFDLGGGTF 185
Query: 327 D 327
D
Sbjct: 186 D 186
>LUXQ_VIBVY (Q7MD16) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 857
Score = 30.8 bits (68), Expect = 5.5
Identities = 19/73 (26%), Positives = 35/73 (47%), Gaps = 10/73 (13%)
Query: 52 LKVADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPD 111
LK D+ C+SG + L V ++I++ N+++ F F+ T +L +
Sbjct: 519 LKQVDVLCNSGEHLLAVLNDILDFSKIEQGKFNIKKRDFNFY----------DTLNTLEN 568
Query: 112 FYKRLQQEKGHKF 124
Y+ + +EKG F
Sbjct: 569 IYRPICREKGVSF 581
>LUXQ_VIBVU (Q8D5Z6) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 857
Score = 30.8 bits (68), Expect = 5.5
Identities = 19/73 (26%), Positives = 35/73 (47%), Gaps = 10/73 (13%)
Query: 52 LKVADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPD 111
LK D+ C+SG + L V ++I++ N+++ F F+ T +L +
Sbjct: 519 LKQVDVLCNSGEHLLAVLNDILDFSKIEQGKFNIKKRDFNFY----------DTLNTLEN 568
Query: 112 FYKRLQQEKGHKF 124
Y+ + +EKG F
Sbjct: 569 IYRPICREKGVSF 581
>DPOL_THELI (P30317) DNA polymerase (EC 2.7.7.7) (Vent DNA polymerase)
[Contains: Endonuclease PI-TliII (EC 3.1.-.-) (Tli pol-1
intein) (IVPS2); Endonuclease PI-TliI (EC 3.1.-.-) (Tli
pol-2 intein) (IVPS1)]
Length = 1702
Score = 30.8 bits (68), Expect = 5.5
Identities = 28/88 (31%), Positives = 42/88 (46%), Gaps = 20/88 (22%)
Query: 267 VIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIM 326
VI+EEG + LE +R DW +I ++T+A+ V E+ILK E +
Sbjct: 1527 VIDEEGRITTRGLEVVRRDW-------SEIAKETQAK--------VLEAILKEGSVEKAV 1571
Query: 327 DELFRRFKNKIVKLHGVEKLELANLVMH 354
E+ R KI K ++ L LV+H
Sbjct: 1572 -EVVRDVVEKIAKY----RVPLEKLVIH 1594
>RPA2_METTH (O27126) DNA-directed RNA polymerase subunit A'' (EC
2.7.7.6)
Length = 451
Score = 30.4 bits (67), Expect = 7.2
Identities = 25/104 (24%), Positives = 45/104 (43%), Gaps = 2/104 (1%)
Query: 232 GMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEE-EGSFDIQRLETIRTDWIKNI 290
G++L D E + A + P ++ E +G D + L+ + D+ + +
Sbjct: 16 GILLPDPVVEYVARIADEEKLKEPELQEMVRLFSRIAERNQGLDDDELLDAVEDDYQRIL 75
Query: 291 NVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFK 334
V + + RA K I +AE + K E + +DEL RR +
Sbjct: 76 KVQELVKRK-RARFPPKLIEDIAEVMKKHELSDDELDELIRRVR 118
>RIR2_BUCAI (P57275) Ribonucleoside-diphosphate reductase beta chain
(EC 1.17.4.1) (Ribonucleotide reductase small subunit)
Length = 376
Score = 30.4 bits (67), Expect = 7.2
Identities = 18/54 (33%), Positives = 29/54 (53%), Gaps = 3/54 (5%)
Query: 204 ELLPGGAMVITLIGRDENNELINAWVVIGMVLNDMASENL---IEKAKLDAFNI 254
EL+ G A +I LI RDE L ++ ++ N +EN+ + + K +A NI
Sbjct: 223 ELMEGNAKIIRLIARDEALHLTGTQHILNLLSNIKNNENMEDVVLECKEEAINI 276
>AROB_MYCLE (Q9CCS4) 3-dehydroquinate synthase (EC 4.2.3.4)
Length = 361
Score = 30.4 bits (67), Expect = 7.2
Identities = 23/70 (32%), Positives = 35/70 (49%), Gaps = 1/70 (1%)
Query: 229 VVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIK 288
VVIG L+D E L + ++ + P TAE IR + +G D R+E + K
Sbjct: 20 VVIGTGLSDQLDELLANRHRVAILHQPVLTQTAEAIRSHLAGKG-VDAHRIEIPDAEAGK 78
Query: 289 NINVSDDIDE 298
+++V D I E
Sbjct: 79 DLSVMDFIWE 88
>RIR2_BUCAP (Q8K9W4) Ribonucleoside-diphosphate reductase beta chain
(EC 1.17.4.1) (Ribonucleotide reductase small subunit)
Length = 376
Score = 30.0 bits (66), Expect = 9.4
Identities = 23/84 (27%), Positives = 36/84 (42%), Gaps = 3/84 (3%)
Query: 160 KVNYTKPLNKGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRD 219
KV L K +YL S V+ +F F E++ G A +I LI RD
Sbjct: 182 KVEINLKLLKRRLYLCLIS---VNVLEAIRFYVSFACSFAFAEREIMEGNAKIIRLIARD 238
Query: 220 ENNELINAWVVIGMVLNDMASENL 243
E L ++ ++ N+ +EN+
Sbjct: 239 EALHLTGTQHILNILNNEKNNENM 262
>META_BACHD (Q9KAK7) Homoserine O-succinyltransferase (EC 2.3.1.46)
(Homoserine O-transsuccinylase) (HTS)
Length = 303
Score = 30.0 bits (66), Expect = 9.4
Identities = 18/66 (27%), Positives = 29/66 (43%), Gaps = 4/66 (6%)
Query: 103 NTTFKSLPDFYKRLQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVN 162
NT++ L FY+ + + K KF +G P P D V +++ + SK N
Sbjct: 79 NTSYDHLQTFYQTIDEVKQKKFDGMIITGAP----IETLPYDEVDYWNELKQIMEWSKTN 134
Query: 163 YTKPLN 168
T L+
Sbjct: 135 VTSTLH 140
>DPOL_THEST (O33845) DNA polymerase (EC 2.7.7.7)
Length = 1829
Score = 30.0 bits (66), Expect = 9.4
Identities = 29/93 (31%), Positives = 46/93 (49%), Gaps = 22/93 (23%)
Query: 267 VIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIM 326
VI+EEG + LE +R DW +I ++T+A+ V E+ILK + E +
Sbjct: 1654 VIDEEGRITTRGLEVVRRDW-------SEIAKETQAK--------VLEAILKEDSVEKAV 1698
Query: 327 DELFRRFKNKIVKLHGVEKLELANLVMH--ITK 357
E+ + +I K ++ L LV+H ITK
Sbjct: 1699 -EIVKDVVEEIAKY----QVPLEKLVIHEQITK 1726
>DPOL_PYRKO (P77933) DNA polymerase (EC 2.7.7.7) [Contains:
Endonuclease PI-PkoI (EC 3.1.-.-) (Pko pol-1 intein)
(IVS-A); Endonuclease PI-PkoII (EC 3.1.-.-) (Pko pol-2
intein) (IVS-B)]
Length = 1671
Score = 30.0 bits (66), Expect = 9.4
Identities = 17/59 (28%), Positives = 28/59 (46%), Gaps = 5/59 (8%)
Query: 267 VIEEEGSFDIQRLETIRTDWIK-----NINVSDDIDEDTRAEAVAKYIRAVAESILKSE 320
VI+EEG + LE +R DW + V + + +D E + ++ V E + K E
Sbjct: 1493 VIDEEGKITTRGLEIVRRDWSEIAKETQARVLEALLKDGDVEKAVRIVKEVTEKLSKYE 1551
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,436,478
Number of Sequences: 164201
Number of extensions: 1798971
Number of successful extensions: 4195
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 4192
Number of HSP's gapped (non-prelim): 19
length of query: 359
length of database: 59,974,054
effective HSP length: 111
effective length of query: 248
effective length of database: 41,747,743
effective search space: 10353440264
effective search space used: 10353440264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0113.13


