

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.7
(402 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
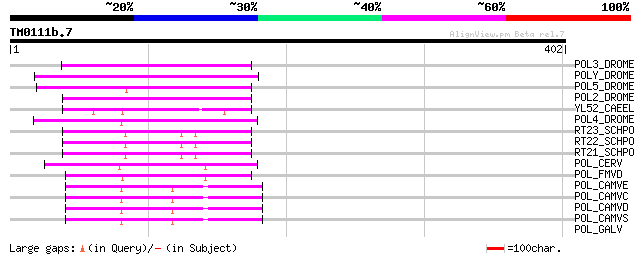
Score E
Sequences producing significant alignments: (bits) Value
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 92 2e-18
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 90 9e-18
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 89 2e-17
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 86 2e-16
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 84 6e-16
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 72 3e-12
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 64 5e-10
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 64 5e-10
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 64 5e-10
POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic prot... 54 5e-07
POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic prot... 48 5e-05
POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic pro... 45 3e-04
POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic pro... 45 3e-04
POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic pro... 45 4e-04
POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic pro... 44 6e-04
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 33 1.7
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 92.4 bits (228), Expect = 2e-18
Identities = 51/138 (36%), Positives = 79/138 (56%)
Query: 38 TSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERN 97
++F+++K ++ P+L +P + + + +AS LG VL Q ++Y S L HE N
Sbjct: 485 SAFKKLKYLISEDPILKVPDFTKKFTLTTDASDVALGAVLSQDGHPLSYISRTLNEHEIN 544
Query: 98 YLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYD 157
Y T + EL +V+A K +RHYL G F + S+H+ L +L+ KD N W + ++D
Sbjct: 545 YSTIEKELLAIVWATKTFRHYLLGRHFEISSDHQPLSWLYRMKDPNSKLTRWRVKLSEFD 604
Query: 158 FTLLYHPGKANVVADALS 175
F + Y GK N VADALS
Sbjct: 605 FDIKYIKGKENCVADALS 622
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 90.1 bits (222), Expect = 9e-18
Identities = 53/164 (32%), Positives = 92/164 (55%), Gaps = 2/164 (1%)
Query: 19 TVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPV-LTLPLEKEPYEVYCNAS*QGLGCVL 77
+V K S++ + R +FQ ++ L + V L P K+P+++ +AS G+G VL
Sbjct: 451 SVSKHMSKKIPVEFNETQRNAFQRLRNILASEDVILKYPDFKKPFDLTTDASASGIGAVL 510
Query: 78 MQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGS-TFTVFSNHKGLKYL 136
Q + S LK E+NY T++ EL +V+AL +++LYGS +F++H+ L +
Sbjct: 511 SQEGRPITMISRTLKQPEQNYATNERELLAIVWALGKLQNFLYGSREINIFTDHQPLTFA 570
Query: 137 FD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQAIH 180
++ N + W ++I+ ++ + Y PGK N VADALS Q ++
Sbjct: 571 VADRNTNAKIKRWKSYIDQHNAKVFYKPGKENFVADALSRQNLN 614
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 89.4 bits (220), Expect = 2e-17
Identities = 52/161 (32%), Positives = 87/161 (53%), Gaps = 5/161 (3%)
Query: 20 VDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQ 79
+ + S + +D SF ++K L ++ +L P +P+ + +AS +G VL Q
Sbjct: 393 IKSSQSSKVPITLDETALQSFNDLKSILCSSEILAFPCFTKPFHLTTDASNWAIGAVLSQ 452
Query: 80 HREA----MAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGS-TFTVFSNHKGLK 134
+ +AY S L E NY T + E+ ++++L R YLYG+ T V+++H+ L
Sbjct: 453 DDQGRDRPIAYISRSLNKTEENYATIEKEMLAIIWSLDNLRAYLYGAGTIKVYTDHQPLT 512
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALS 175
+ ++ N + W IE+Y+ L+Y PGK+NVVADALS
Sbjct: 513 FALGNRNFNAKLKRWKARIEEYNCELIYKPGKSNVVADALS 553
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 85.9 bits (211), Expect = 2e-16
Identities = 48/137 (35%), Positives = 77/137 (56%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
+F+++K + P+L LP ++ + + +AS LG VL Q+ +++ S L HE NY
Sbjct: 485 AFEKLKALIIRDPILQLPDFEKKFVLTTDASNLALGAVLSQNGHPISFISRTLNDHELNY 544
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
+ EL +V+A K +RHYL G F + S+H+ L++L + K+ W + +Y F
Sbjct: 545 SAIEKELLAIVWATKTFRHYLLGRQFLIASDHQPLRWLHNLKEPGAKLERWRVRLSEYQF 604
Query: 159 TLLYHPGKANVVADALS 175
+ Y GK N VADALS
Sbjct: 605 KIDYIKGKENSVADALS 621
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 84.0 bits (206), Expect = 6e-16
Identities = 56/150 (37%), Positives = 82/150 (54%), Gaps = 15/150 (10%)
Query: 39 SFQEMKKRLTAAPVLTLPLEK------EPYEVYCNAS*QGLGCVLMQH-----REAMAYA 87
+FQE+KK + PVL P + P+ +Y +AS +G+G VL Q + +A+A
Sbjct: 1215 AFQELKKLVCQTPVLAQPDVEAALKGDRPFMIYTDASRKGIGAVLAQEGPDGQQHPIAFA 1274
Query: 88 SSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQR 147
S L E Y DLE ++FAL+ ++ +YG+ TVF++HK L L K + R
Sbjct: 1275 SKALSPAETRYHITDLEALAMMFALRRFKTIIYGTAITVFTDHKPLISLL--KGSPLADR 1332
Query: 148 CWMNFIE--DYDFTLLYHPGKANVVADALS 175
W IE ++D ++Y GKAN VADALS
Sbjct: 1333 LWRWSIEILEFDVKIVYLAGKANAVADALS 1362
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 72.0 bits (175), Expect = 3e-12
Identities = 53/166 (31%), Positives = 82/166 (48%), Gaps = 4/166 (2%)
Query: 18 RTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVL 77
R + + + F + +F +K +L +L P + + + +AS Q G VL
Sbjct: 572 RHITRLCKKNVPFEWTDECQKAFIHLKSQLINPTLLQYPDFSKEFCITTDASKQACGAVL 631
Query: 78 MQ----HREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGL 133
Q H+ +AYAS E N T + ELA + +A+ +R Y+YG FTV ++H+ L
Sbjct: 632 TQNHNGHQLPVAYASRAFTKGESNKSTTEQELAAIHWAIIHFRPYIYGKHFTVKTDHRPL 691
Query: 134 KYLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQAI 179
YLF + + +E+Y+FT+ Y GK N VADALS I
Sbjct: 692 TYLFSMVNPSSKLTRIRLELEEYNFTVEYLKGKDNHVADALSRITI 737
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 64.3 bits (155), Expect = 5e-10
Identities = 42/146 (28%), Positives = 75/146 (50%), Gaps = 9/146 (6%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHRE-----AMAYASSQLKL 93
+ + +K+ L + PVL + + +AS +G VL Q + + Y S+++
Sbjct: 685 AIENIKQCLVSPPVLRHFDFSKKILLETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSK 744
Query: 94 HERNYLTHDLELAVVVFALKIWRHYLYGST--FTVFSNHKGL--KYLFD*KDLNMGQRCW 149
+ NY D E+ ++ +LK WRHYL + F + ++H+ L + + + N W
Sbjct: 745 AQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKILTDHRNLIGRITNESEPENKRLARW 804
Query: 150 MNFIEDYDFTLLYHPGKANVVADALS 175
F++D++F + Y PG AN +ADALS
Sbjct: 805 QLFLQDFNFEINYRPGSANHIADALS 830
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 64.3 bits (155), Expect = 5e-10
Identities = 42/146 (28%), Positives = 75/146 (50%), Gaps = 9/146 (6%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHRE-----AMAYASSQLKL 93
+ + +K+ L + PVL + + +AS +G VL Q + + Y S+++
Sbjct: 685 AIENIKQCLVSPPVLRHFDFSKKILLETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSK 744
Query: 94 HERNYLTHDLELAVVVFALKIWRHYLYGST--FTVFSNHKGL--KYLFD*KDLNMGQRCW 149
+ NY D E+ ++ +LK WRHYL + F + ++H+ L + + + N W
Sbjct: 745 AQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKILTDHRNLIGRITNESEPENKRLARW 804
Query: 150 MNFIEDYDFTLLYHPGKANVVADALS 175
F++D++F + Y PG AN +ADALS
Sbjct: 805 QLFLQDFNFEINYRPGSANHIADALS 830
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 64.3 bits (155), Expect = 5e-10
Identities = 42/146 (28%), Positives = 75/146 (50%), Gaps = 9/146 (6%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHRE-----AMAYASSQLKL 93
+ + +K+ L + PVL + + +AS +G VL Q + + Y S+++
Sbjct: 685 AIENIKQCLVSPPVLRHFDFSKKILLETDASDVAVGAVLSQKHDDDKYYPVGYYSAKMSK 744
Query: 94 HERNYLTHDLELAVVVFALKIWRHYLYGST--FTVFSNHKGL--KYLFD*KDLNMGQRCW 149
+ NY D E+ ++ +LK WRHYL + F + ++H+ L + + + N W
Sbjct: 745 AQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKILTDHRNLIGRITNESEPENKRLARW 804
Query: 150 MNFIEDYDFTLLYHPGKANVVADALS 175
F++D++F + Y PG AN +ADALS
Sbjct: 805 QLFLQDFNFEINYRPGSANHIADALS 830
>POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 659
Score = 54.3 bits (129), Expect = 5e-07
Identities = 43/162 (26%), Positives = 72/162 (43%), Gaps = 8/162 (4%)
Query: 26 EESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLM----QHR 81
E+ST+ + ++KK L + P L P + + +AS + G +L H
Sbjct: 494 EDSTWTWNDTDSQYMAKIKKNLKSFPKLYHPEPNDKLVIETDASEEFWGGILKAIHNSHE 553
Query: 82 EAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*K- 140
YAS K ERNY +++ EL V+ +K + YL S F + +++K + +
Sbjct: 554 YICRYASGSFKAAERNYHSNEKELLAVIRVIKKFSIYLTPSRFLIRTDNKNFTHFVNINL 613
Query: 141 --DLNMGQRC-WMNFIEDYDFTLLYHPGKANVVADALSHQAI 179
D G+ W ++ YDF + + G NV AD L +
Sbjct: 614 KGDRKQGRLVRWQMWLSQYDFDVEHIAGTKNVFADFLQENTL 655
>POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 666
Score = 47.8 bits (112), Expect = 5e-05
Identities = 41/144 (28%), Positives = 64/144 (43%), Gaps = 9/144 (6%)
Query: 41 QEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQH-----REAMAYASSQLKLHE 95
+++KK L + P L LP ++ + +AS G VL Y+S K E
Sbjct: 518 KKIKKNLGSFPKLYLPKPEDHLIIETDASDSFWGGVLKARALDGVELICRYSSGSFKQAE 577
Query: 96 RNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*K---DLNMGQRC-WMN 151
+NY ++D EL V + + YL FTV +++K Y D G+ W N
Sbjct: 578 KNYHSNDKELLAVKQVITKFSAYLTPVRFTVRTDNKNFTYFLRINLKGDSKQGRLVRWQN 637
Query: 152 FIEDYDFTLLYHPGKANVVADALS 175
+ Y F + + G NV+AD L+
Sbjct: 638 WFSKYQFDVEHLEGVKNVLADCLT 661
>POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 45.4 bits (106), Expect = 3e-04
Identities = 46/158 (29%), Positives = 70/158 (44%), Gaps = 18/158 (11%)
Query: 41 QEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQ-------HREAMA-YASSQLK 92
Q++KK L P L PL +E + +AS G +L + E + YAS K
Sbjct: 525 QKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTELICRYASGSFK 584
Query: 93 LHERNYLTHDLELAVVVFALKIWR------HYLYGSTFTVFSNHKGLKYLFD*KDLNMGQ 146
ERNY ++D E V+ +K + H+L + T F + L Y D K +G+
Sbjct: 585 AAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNYKGDSK---LGR 641
Query: 147 RC-WMNFIEDYDFTLLYHPGKANVVADALSHQAIHVSS 183
W ++ Y F + + G N AD LS + V+S
Sbjct: 642 NIRWQAWLSHYSFDVEHIKGTDNHFADFLSREFNKVNS 679
>POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 45.4 bits (106), Expect = 3e-04
Identities = 46/158 (29%), Positives = 70/158 (44%), Gaps = 18/158 (11%)
Query: 41 QEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQ-------HREAMA-YASSQLK 92
Q++KK L P L PL +E + +AS G +L + E + YAS K
Sbjct: 525 QKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTELICRYASGSFK 584
Query: 93 LHERNYLTHDLELAVVVFALKIWR------HYLYGSTFTVFSNHKGLKYLFD*KDLNMGQ 146
ERNY ++D E V+ +K + H+L + T F + L Y D K +G+
Sbjct: 585 AAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNYKGDSK---LGR 641
Query: 147 RC-WMNFIEDYDFTLLYHPGKANVVADALSHQAIHVSS 183
W ++ Y F + + G N AD LS + V+S
Sbjct: 642 NIRWQAWLSHYSFDVEHIKGTDNHFADFLSREFNKVNS 679
>POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 674
Score = 44.7 bits (104), Expect = 4e-04
Identities = 45/158 (28%), Positives = 70/158 (43%), Gaps = 18/158 (11%)
Query: 41 QEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQ-------HREAMA-YASSQLK 92
Q++KK L P L PL +E + +AS G +L + E + YAS K
Sbjct: 520 QKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTELICRYASGSFK 579
Query: 93 LHERNYLTHDLELAVVVFALKIWR------HYLYGSTFTVFSNHKGLKYLFD*KDLNMGQ 146
E+NY ++D E V+ +K + H+L + T F + L Y D K +G+
Sbjct: 580 AAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNYKGDSK---LGR 636
Query: 147 RC-WMNFIEDYDFTLLYHPGKANVVADALSHQAIHVSS 183
W ++ Y F + + G N AD LS + V+S
Sbjct: 637 NIRWQAWLSHYSFDVEHIKGTDNHFADFLSREFNRVNS 674
>POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 44.3 bits (103), Expect = 6e-04
Identities = 45/158 (28%), Positives = 70/158 (43%), Gaps = 18/158 (11%)
Query: 41 QEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQ-------HREAMA-YASSQLK 92
Q++KK L P L PL +E + +AS G +L + E + YAS K
Sbjct: 525 QKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTELICRYASGSFK 584
Query: 93 LHERNYLTHDLELAVVVFALKIWR------HYLYGSTFTVFSNHKGLKYLFD*KDLNMGQ 146
E+NY ++D E V+ +K + H+L + T F + L Y D K +G+
Sbjct: 585 AAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNYKGDSK---LGR 641
Query: 147 RC-WMNFIEDYDFTLLYHPGKANVVADALSHQAIHVSS 183
W ++ Y F + + G N AD LS + V+S
Sbjct: 642 NIRWQAWLSHYSFDVEHIKGTDNHFADFLSREFNKVNS 679
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 32.7 bits (73), Expect = 1.7
Identities = 28/96 (29%), Positives = 41/96 (42%), Gaps = 4/96 (4%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQ----HREAMAYASSQLKLH 94
+F +KK L +AP L LP +P+ +Y + VL Q R +AY S +L
Sbjct: 460 AFDHIKKALLSAPALALPDLTKPFTLYIDERAGVARGVLTQTLGPWRRPVAYLSKKLDPV 519
Query: 95 ERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNH 130
+ T +A V LK G TV ++H
Sbjct: 520 ASGWPTCLKAVAAVALLLKDADKLTLGQNVTVIASH 555
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.353 0.155 0.544
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,050,666
Number of Sequences: 164201
Number of extensions: 1416064
Number of successful extensions: 6305
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 6285
Number of HSP's gapped (non-prelim): 16
length of query: 402
length of database: 59,974,054
effective HSP length: 112
effective length of query: 290
effective length of database: 41,583,542
effective search space: 12059227180
effective search space used: 12059227180
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0111b.7


