

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.4
(467 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
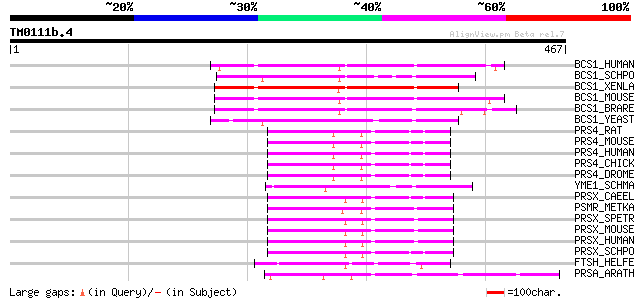
Score E
Sequences producing significant alignments: (bits) Value
BCS1_HUMAN (Q9Y276) Mitochondrial chaperone BCS1 (BCS1-like prot... 136 1e-31
BCS1_SCHPO (Q9P6Q3) Probable mitochondrial chaperone BCS1 (BCS1-... 136 1e-31
BCS1_XENLA (Q7ZTL7) Mitochondrial chaperone BCS1 (BCS1-like prot... 135 2e-31
BCS1_MOUSE (Q9CZP5) Mitochondrial chaperone BCS1 (BCS1-like prot... 132 2e-30
BCS1_BRARE (Q7ZV60) Mitochondrial chaperone BCS1 (BCS1-like prot... 130 9e-30
BCS1_YEAST (P32839) Mitochondrial chaperone BCS1 125 2e-28
PRS4_RAT (P62193) 26S protease regulatory subunit 4 (P26s4) (Pro... 71 7e-12
PRS4_MOUSE (P62192) 26S protease regulatory subunit 4 (P26s4) (P... 71 7e-12
PRS4_HUMAN (P62191) 26S protease regulatory subunit 4 (P26s4) (P... 71 7e-12
PRS4_CHICK (Q90732) 26S protease regulatory subunit 4 (P26s4) (P... 71 7e-12
PRS4_DROME (P48601) 26S protease regulatory subunit 4 (P26s4) 70 9e-12
YME1_SCHMA (P46508) YME1 protein homolog (EC 3.4.24.-) 69 2e-11
PRSX_CAEEL (O17071) Probable 26S protease regulatory subunit S10B 69 2e-11
PSMR_METKA (Q8TX03) Proteasome-activating nucleotidase (Proteaso... 69 3e-11
PRSX_SPETR (P62335) 26S protease regulatory subunit S10B (Protea... 69 3e-11
PRSX_MOUSE (P62334) 26S protease regulatory subunit S10B (Protea... 69 3e-11
PRSX_HUMAN (P62333) 26S protease regulatory subunit S10B (Protea... 69 3e-11
PRSX_SCHPO (O74445) Probable 26S protease subunit rpt4 69 3e-11
FTSH_HELFE (O32617) Cell division protein ftsH homolog (EC 3.4.2... 67 1e-10
PRSA_ARATH (O04019) 26S protease regulatory subunit 6A homolog (... 67 1e-10
>BCS1_HUMAN (Q9Y276) Mitochondrial chaperone BCS1 (BCS1-like
protein) (H-BCS1)
Length = 419
Score = 136 bits (343), Expect = 1e-31
Identities = 90/257 (35%), Positives = 138/257 (53%), Gaps = 18/257 (7%)
Query: 170 VLERAK--AIKQENMAVKIYTVEYEAWCRNG-TRFDHPMTFKTLAIDAGLKKELVSDLDR 226
+LE A+ A++QE +YT W G R P+ ++ + GL +V D+
Sbjct: 150 ILEEARELALQQEEGKTVMYTAVGSEWRPFGYPRRRRPLN--SVVLQQGLADRIVRDVQE 207
Query: 227 FLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLT--ALNDNKDL 284
F+ ++Y G ++RGYLLYGPPG GKSS I A+A L + I L LT +L+D++ L
Sbjct: 208 FIDNPKWYTDRGIPYRRGYLLYGPPGCGKSSFITALAGELEHSICLLSLTDSSLSDDR-L 266
Query: 285 KNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEE 344
+LL +S++++ED+D + + + G ++T SGLLN +DG+ S E
Sbjct: 267 NHLLSVAPQQSLVLLEDVDAAFLSRDLAVENPVKYQGLGRLTFSGLLNALDGVAST--EA 324
Query: 345 RIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLL 404
RI+ TTNH +RLDPAL+RPGR+D+ + YC+ Q+ + L E
Sbjct: 325 RIVFMTTNHVDRLDPALIRPGRVDLKEYVGYCSHWQLTQMFQRFYPGQAPSLAENFA--- 381
Query: 405 RHV-----QVTPAEVAG 416
HV Q++PA+V G
Sbjct: 382 EHVLRATNQISPAQVQG 398
>BCS1_SCHPO (Q9P6Q3) Probable mitochondrial chaperone BCS1
(BCS1-like protein)
Length = 449
Score = 136 bits (342), Expect = 1e-31
Identities = 83/223 (37%), Positives = 130/223 (58%), Gaps = 12/223 (5%)
Query: 175 KAIKQENMAVKIYTVEYEAWCRNGTRFDHPMTFKTLA---IDAGLKKELVSDLDRFLQGK 231
+A K A K T Y AW F HP + + L+ +++ +KK + D+ FL+
Sbjct: 172 EAQKFMQSAQKNKTTIYTAWATEWKPFGHPRSKRMLSSVVLESNVKKMITDDVHDFLRNS 231
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLT--ALNDNKDLKNLLL 289
++Y+ G ++RGYLLYGPPG+GK+S + A+A L YDI L+L L D++ L +LL
Sbjct: 232 QWYDTRGIPYRRGYLLYGPPGSGKTSFLYALAGELDYDICVLNLAEKGLTDDR-LNHLLS 290
Query: 290 AMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIF 349
+ ++++++ED+D + + RE E + + +T SGLLN +DG+ S +ERII
Sbjct: 291 NVPPKAVVLLEDVDSAFQ--GRERSGEVGFHAN--VTFSGLLNALDGVTS--SDERIIFM 344
Query: 350 TTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGIS 392
TTNH E+LDPAL+RPGR+D+ L T ++ + G S
Sbjct: 345 TTNHPEKLDPALVRPGRVDVKAYLGNATPEQVREMFTRFYGHS 387
>BCS1_XENLA (Q7ZTL7) Mitochondrial chaperone BCS1 (BCS1-like
protein)
Length = 419
Score = 135 bits (341), Expect = 2e-31
Identities = 77/208 (37%), Positives = 126/208 (60%), Gaps = 8/208 (3%)
Query: 173 RAKAIKQENMAVKIYTVEYEAWCRNG-TRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGK 231
R A+KQ+ +Y W + G R P++ ++ ++ G+ +++V D+ F++
Sbjct: 155 RELALKQQVGKTVMYNAVGAEWRQFGFPRRRRPLS--SVVLEQGISEKIVQDVKGFIENP 212
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDL--TALNDNKDLKNLLL 289
++Y+ G ++RGYLLYGPPG GKSS I A+A L Y I + L ++L+D++ L +LL
Sbjct: 213 KWYSDRGIPYRRGYLLYGPPGCGKSSFITALAGELEYSICLMSLSDSSLSDDR-LNHLLS 271
Query: 290 AMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIF 349
+SI+++ED+D + + + A G ++T SGLLN +DG+ S E RI+
Sbjct: 272 VAPQQSIILLEDVDAAFVSRDLNKQNPTAYQGMGRLTFSGLLNALDGVAST--EARIVFM 329
Query: 350 TTNHRERLDPALLRPGRMDMHVNLSYCT 377
TTNH +RLDPAL+RPGR+D+ + +CT
Sbjct: 330 TTNHIDRLDPALIRPGRVDVKQYVGHCT 357
>BCS1_MOUSE (Q9CZP5) Mitochondrial chaperone BCS1 (BCS1-like
protein)
Length = 418
Score = 132 bits (332), Expect = 2e-30
Identities = 86/249 (34%), Positives = 134/249 (53%), Gaps = 10/249 (4%)
Query: 173 RAKAIKQENMAVKIYTVEYEAWCRNG-TRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGK 231
RA A++QE +YT W G R P+ ++ + GL +V D+ F+
Sbjct: 155 RALALQQEEGKTVMYTAVGSEWRTFGYPRRRRPLD--SVVLQQGLADRIVKDIREFIDNP 212
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLT--ALNDNKDLKNLLL 289
++Y G ++RGYLLYGPPG GKSS I A+A L + I L LT +L+D++ L +LL
Sbjct: 213 KWYIDRGIPYRRGYLLYGPPGCGKSSFITALAGELEHSICLLSLTDSSLSDDR-LNHLLS 271
Query: 290 AMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIF 349
+S++++ED+D + + + G ++T SGLLN +DG+ S E RI+
Sbjct: 272 VAPQQSLVLLEDVDAAFLSRDLAVENPIKYQGLGRLTFSGLLNALDGVAST--EARIVFM 329
Query: 350 TTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGG--LLRHV 407
TTN+ +RLDPAL+RPGR+D+ + YC+ Q+ + L E L
Sbjct: 330 TTNYIDRLDPALIRPGRVDLKEYVGYCSHWQLTQMFQRFYPGQAPSLAENFAEHVLKATS 389
Query: 408 QVTPAEVAG 416
+++PA+V G
Sbjct: 390 EISPAQVQG 398
>BCS1_BRARE (Q7ZV60) Mitochondrial chaperone BCS1 (BCS1-like
protein)
Length = 420
Score = 130 bits (326), Expect = 9e-30
Identities = 90/274 (32%), Positives = 146/274 (52%), Gaps = 28/274 (10%)
Query: 173 RAKAIKQENMAVKIYTVEYEAWCRNG-TRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGK 231
R A+KQE +YT W G R P++ ++ +++G+ + +V D+ F+
Sbjct: 155 RELALKQEEGRTVMYTAMGAEWRPFGFPRRRRPLS--SVVLESGVAERIVDDVKEFIGNP 212
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLT--ALNDNKDLKNLLL 289
++Y G ++RGYLLYGPPG GKSS I A+A L Y I + L+ +L+D++ L +LL
Sbjct: 213 KWYTDRGIPYRRGYLLYGPPGCGKSSFITALAGELGYSICLMSLSDRSLSDDR-LNHLLS 271
Query: 290 AMSNRSILVIEDIDCS-IKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIII 348
+SI+++ED+D + + + A G ++T SGLLN +DG+ S E RI+
Sbjct: 272 VAPQQSIILLEDVDAAFVSRELLPTENPLAYQGMGRLTFSGLLNALDGVAS--SEARIVF 329
Query: 349 FTTNHRERLDPALLRPGRMDMHVNLSYCTF--------------SAFEQLAFTYLGISQH 394
TTN ERLDPAL+RPGR+D+ + +C+ SA E F+ ++ H
Sbjct: 330 MTTNFIERLDPALVRPGRVDLKQYVGHCSHWQLTQMFRRFYPQESAAEADHFSEQALAAH 389
Query: 395 KLFE--QIGGLLRHVQVTPAEVAGQLTKIADVEE 426
Q+ G H + + AG + IA++++
Sbjct: 390 TDLSAAQVQG---HFMLYKTDPAGAIKNIAEIKD 420
>BCS1_YEAST (P32839) Mitochondrial chaperone BCS1
Length = 456
Score = 125 bits (315), Expect = 2e-28
Identities = 79/212 (37%), Positives = 122/212 (57%), Gaps = 12/212 (5%)
Query: 170 VLERAKAIKQENMAVKIYTVEYEAWCRNGTRFDHPMTFKTLA---IDAGLKKELVSDLDR 226
+L AK I + K TV Y ++ +F P + L +D+G+K+ ++ D+
Sbjct: 187 ILNEAKDIALKTTEGK--TVIYTSFGPEWRKFGQPKAKRMLPSVILDSGIKEGILDDVYD 244
Query: 227 FLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKD-LK 285
F++ ++Y+ G ++RGYLLYGPPG+GK+S I A+A L Y+I L+L+ N D L
Sbjct: 245 FMKNGKWYSDRGIPYRRGYLLYGPPGSGKTSFIQALAGELDYNICILNLSENNLTDDRLN 304
Query: 286 NLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEER 345
+L+ M RSIL++EDID + + + + + +T SGLLN +DG+ S EE
Sbjct: 305 HLMNNMPERSILLLEDIDAAF----NKRSQTGEQGFHSSVTFSGLLNALDGVTS--SEET 358
Query: 346 IIIFTTNHRERLDPALLRPGRMDMHVNLSYCT 377
I TTNH E+LD A++RPGR+D V + T
Sbjct: 359 ITFMTTNHPEKLDAAIMRPGRIDYKVFVGNAT 390
>PRS4_RAT (P62193) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 70.9 bits (172), Expect = 7e-12
Identities = 52/162 (32%), Positives = 82/162 (50%), Gaps = 13/162 (8%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E+Y G +G +LYGPPGTGK+ L A+AN S +
Sbjct: 195 QEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQTSATFLRVVGSE 254
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L ++ SI+ I++ID + + D + + + T+
Sbjct: 255 LIQKYLGDGPKLVRELFRVAEEHAPSIVFIDEIDA---IGTKRYDSNSGGEREIQRTMLE 311
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + G+ ++I+ TN E LDPAL+RPGR+D +
Sbjct: 312 LLNQLDG-FDSRGDVKVIM-ATNRIETLDPALIRPGRIDRKI 351
>PRS4_MOUSE (P62192) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 70.9 bits (172), Expect = 7e-12
Identities = 52/162 (32%), Positives = 82/162 (50%), Gaps = 13/162 (8%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E+Y G +G +LYGPPGTGK+ L A+AN S +
Sbjct: 195 QEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQTSATFLRVVGSE 254
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L ++ SI+ I++ID + + D + + + T+
Sbjct: 255 LIQKYLGDGPKLVRELFRVAEEHAPSIVFIDEIDA---IGTKRYDSNSGGEREIQRTMLE 311
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + G+ ++I+ TN E LDPAL+RPGR+D +
Sbjct: 312 LLNQLDG-FDSRGDVKVIM-ATNRIETLDPALIRPGRIDRKI 351
>PRS4_HUMAN (P62191) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 70.9 bits (172), Expect = 7e-12
Identities = 52/162 (32%), Positives = 82/162 (50%), Gaps = 13/162 (8%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E+Y G +G +LYGPPGTGK+ L A+AN S +
Sbjct: 195 QEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQTSATFLRVVGSE 254
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L ++ SI+ I++ID + + D + + + T+
Sbjct: 255 LIQKYLGDGPKLVRELFRVAEEHAPSIVFIDEIDA---IGTKRYDSNSGGEREIQRTMLE 311
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + G+ ++I+ TN E LDPAL+RPGR+D +
Sbjct: 312 LLNQLDG-FDSRGDVKVIM-ATNRIETLDPALIRPGRIDRKI 351
>PRS4_CHICK (Q90732) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 70.9 bits (172), Expect = 7e-12
Identities = 52/162 (32%), Positives = 82/162 (50%), Gaps = 13/162 (8%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E+Y G +G +LYGPPGTGK+ L A+AN S +
Sbjct: 195 QEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQTSATFLRVVGSE 254
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L ++ SI+ I++ID + + D + + + T+
Sbjct: 255 LIQKYLGDGPKLVRELFRVAEEHGPSIVFIDEIDA---IGTKRYDSNSGGEREIQRTMLE 311
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + G+ ++I+ TN E LDPAL+RPGR+D +
Sbjct: 312 LLNQLDG-FDSRGDVKVIM-ATNRIETLDPALIRPGRIDRKI 351
>PRS4_DROME (P48601) 26S protease regulatory subunit 4 (P26s4)
Length = 439
Score = 70.5 bits (171), Expect = 9e-12
Identities = 52/162 (32%), Positives = 82/162 (50%), Gaps = 13/162 (8%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E+Y G +G +LYGPPGTGK+ L A+AN S +
Sbjct: 194 QEIKESVELPLTHPEYYEEMGIKPPKGVILYGPPGTGKTLLAKAVANQTSATFLRVVGSE 253
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L ++ SI+ I++ID + + D + + + T+
Sbjct: 254 LIQKYLGDGPKLVRELFRVAEEHAPSIVFIDEIDA---VGTKRYDSNSGGEREIQRTMLE 310
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + G+ ++I+ TN E LDPAL+RPGR+D +
Sbjct: 311 LLNQLDG-FDSRGDVKVIM-ATNRIETLDPALIRPGRIDRKI 350
>YME1_SCHMA (P46508) YME1 protein homolog (EC 3.4.24.-)
Length = 662
Score = 69.3 bits (168), Expect = 2e-11
Identities = 60/181 (33%), Positives = 84/181 (46%), Gaps = 14/181 (7%)
Query: 216 LKKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMAN-------YLSY 268
+KKELV D+ FL+ E +N+ G +G LL GPPG GK+ L A++ Y S
Sbjct: 174 VKKELV-DVVEFLRNPEKFNQIGAKLPKGVLLVGPPGVGKTLLAKAVSGEAQVPFLYASG 232
Query: 269 DIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLS 328
+D L L ++ + A N LV D S+ N T++
Sbjct: 233 SSFDEVLVGLGASRIRQLFTTAKQNSPCLVFIDEIDSVGGNRTFSPHHPFAN----QTIN 288
Query: 329 GLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTY 388
LL +DG S E I++ TN E LD ALLRPGR D+ +++S T+ L Y
Sbjct: 289 QLLAEMDGFQS--KEGIIVLGATNQAEVLDKALLRPGRFDVQIHVSPPTYEGRIALLNLY 346
Query: 389 L 389
L
Sbjct: 347 L 347
>PRSX_CAEEL (O17071) Probable 26S protease regulatory subunit S10B
Length = 406
Score = 69.3 bits (168), Expect = 2e-11
Identities = 51/164 (31%), Positives = 87/164 (52%), Gaps = 13/164 (7%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L E + R G +G LL+GPPGTGK+ L A+A+ L + + +A
Sbjct: 160 RELREVVELPLINPELFKRVGITPPKGCLLFGPPGTGKTLLARAVASQLDCNFLKVVSSA 219
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + ++ M N + I+ +++ID + R E + + + + TL
Sbjct: 220 IVDKYIGESARMIREMFNYARDHQPCIVFMDEIDA---IGGRRFSEGTSADREIQRTLME 276
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNL 373
LLN +DG + G+ ++I+ TN + LDPALLRPGR+D + +
Sbjct: 277 LLNQLDG-FDSLGKVKVIM-ATNRPDTLDPALLRPGRLDRKIEI 318
>PSMR_METKA (Q8TX03) Proteasome-activating nucleotidase (Proteasome
regulatory subunit)
Length = 436
Score = 68.9 bits (167), Expect = 3e-11
Identities = 50/164 (30%), Positives = 83/164 (50%), Gaps = 13/164 (7%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+E+ +++ L+ E + + G +G LLYGPPGTGK+ L A+AN+ L
Sbjct: 189 REIREVVEKPLKEPELFEKVGVEPPKGVLLYGPPGTGKTLLAKAVANHADATFIRLAAPE 248
Query: 278 L-----NDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L + L L ++ SI+ I++ID + R + + + + + TL+
Sbjct: 249 LVQKFIGEGARLVRELFELAREKAPSIIFIDEIDA---IGARRMRDATSGDREVQRTLTQ 305
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNL 373
LL +DG ++ +I TN ++ LDPALLRPGR D H+ +
Sbjct: 306 LLAEMDGFDPL--DDIKVIAATNRKDILDPALLRPGRFDRHIKI 347
>PRSX_SPETR (P62335) 26S protease regulatory subunit S10B
(Proteasome subunit p42) (Proteasome 26S subunit ATPase
6) (Conserved ATPase domain protein 44) (CADp44)
Length = 389
Score = 68.9 bits (167), Expect = 3e-11
Identities = 52/165 (31%), Positives = 86/165 (51%), Gaps = 15/165 (9%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L E + R G +G LLYGPPGTGK+ L A+A+ L + + ++
Sbjct: 143 RELREVIELPLTNPELFQRVGIIPPKGCLLYGPPGTGKTLLARAVASQLDCNFLKVVSSS 202
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + L+ M N + I+ +++ID + R E + + + + TL
Sbjct: 203 IVDKYIGESARLIREMFNYARDHQPCIIFMDEIDA---IGGRRFSEGTSADREIQRTLME 259
Query: 330 LLNVIDGLWSCCGEERI-IIFTTNHRERLDPALLRPGRMDMHVNL 373
LLN +DG + R+ +I TN + LDPALLRPGR+D +++
Sbjct: 260 LLNQMDGFDTL---HRVKMIMATNRPDTLDPALLRPGRLDRKIHI 301
>PRSX_MOUSE (P62334) 26S protease regulatory subunit S10B
(Proteasome subunit p42) (Proteasome 26S subunit ATPase
6)
Length = 389
Score = 68.9 bits (167), Expect = 3e-11
Identities = 52/165 (31%), Positives = 86/165 (51%), Gaps = 15/165 (9%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L E + R G +G LLYGPPGTGK+ L A+A+ L + + ++
Sbjct: 143 RELREVIELPLTNPELFQRVGIIPPKGCLLYGPPGTGKTLLARAVASQLDCNFLKVVSSS 202
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + L+ M N + I+ +++ID + R E + + + + TL
Sbjct: 203 IVDKYIGESARLIREMFNYARDHQPCIIFMDEIDA---IGGRRFSEGTSADREIQRTLME 259
Query: 330 LLNVIDGLWSCCGEERI-IIFTTNHRERLDPALLRPGRMDMHVNL 373
LLN +DG + R+ +I TN + LDPALLRPGR+D +++
Sbjct: 260 LLNQMDGFDTL---HRVKMIMATNRPDTLDPALLRPGRLDRKIHI 301
>PRSX_HUMAN (P62333) 26S protease regulatory subunit S10B
(Proteasome subunit p42) (Proteasome 26S subunit ATPase
6)
Length = 389
Score = 68.9 bits (167), Expect = 3e-11
Identities = 52/165 (31%), Positives = 86/165 (51%), Gaps = 15/165 (9%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L E + R G +G LLYGPPGTGK+ L A+A+ L + + ++
Sbjct: 143 RELREVIELPLTNPELFQRVGIIPPKGCLLYGPPGTGKTLLARAVASQLDCNFLKVVSSS 202
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + L+ M N + I+ +++ID + R E + + + + TL
Sbjct: 203 IVDKYIGESARLIREMFNYARDHQPCIIFMDEIDA---IGGRRFSEGTSADREIQRTLME 259
Query: 330 LLNVIDGLWSCCGEERI-IIFTTNHRERLDPALLRPGRMDMHVNL 373
LLN +DG + R+ +I TN + LDPALLRPGR+D +++
Sbjct: 260 LLNQMDGFDTL---HRVKMIMATNRPDTLDPALLRPGRLDRKIHI 301
>PRSX_SCHPO (O74445) Probable 26S protease subunit rpt4
Length = 388
Score = 68.6 bits (166), Expect = 3e-11
Identities = 51/164 (31%), Positives = 86/164 (52%), Gaps = 13/164 (7%)
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTA 277
+EL ++ L+ E + R G +G LLYGPPGTGK+ L A+A L + + +A
Sbjct: 142 RELREVIELPLKNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAVAASLGVNFLKVVSSA 201
Query: 278 LNDN--KDLKNLLLAMSNRS------ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
+ D + ++ M + ++ +++ID + R E + + + + TL
Sbjct: 202 IVDKYIGESARIIREMFGYAKEHEPCVIFMDEIDA---IGGRRFSEGTSADREIQRTLME 258
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNL 373
LLN +DG + G+ +II+ TN + LDPALLRPGR+D + +
Sbjct: 259 LLNQMDG-FDYLGQTKIIM-ATNRPDTLDPALLRPGRLDRKIEI 300
>FTSH_HELFE (O32617) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 638
Score = 67.0 bits (162), Expect = 1e-10
Identities = 58/177 (32%), Positives = 87/177 (48%), Gaps = 20/177 (11%)
Query: 207 FKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYL 266
F +A + K+E+V +D FL+ + Y G +G LL GPPGTGK+ L A+A
Sbjct: 175 FNDMAGNEEAKEEVVEIVD-FLKYPDRYASLGAKIPKGVLLVGPPGTGKTLLAKAVAGEA 233
Query: 267 SYDIYDLDLTALNDN---------KDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEA 317
S + + ++ + +DL + + SI+ I++ID K +R
Sbjct: 234 SVPFFSMGGSSFIEMFVGLGASRVRDLFD-IAKKEAPSIIFIDEIDAIGK--SRAAGGMI 290
Query: 318 AKNGDNKMTLSGLLNVIDGLWSCCGEER---IIIFTTNHRERLDPALLRPGRMDMHV 371
+ N + + TL+ LL +DG G E I++ TN E LDPALLRPGR D V
Sbjct: 291 SGNDEREQTLNQLLAEMDGF----GSENAPVIVLAATNRPEILDPALLRPGRFDRQV 343
>PRSA_ARATH (O04019) 26S protease regulatory subunit 6A homolog
(TAT-binding protein homolog 1) (TBP-1)
Length = 419
Score = 66.6 bits (161), Expect = 1e-10
Identities = 72/261 (27%), Positives = 115/261 (43%), Gaps = 26/261 (9%)
Query: 215 GLKK---ELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAM-----ANYL 266
GL+K ELV + + KE + + G +G LLYGPPGTGK+ + A A +L
Sbjct: 170 GLEKQIQELVEAIVLPMTHKEQFEKLGIRPPKGVLLYGPPGTGKTLMARACAAQTNATFL 229
Query: 267 SYDIYDLDLTALNDNKDLKN---LLLAMSNRSILVIEDIDCSIKLPNREEDEEAAKNGDN 323
L + D L LL + I+ I++ID + + D E + + +
Sbjct: 230 KLAGPQLVQMFIGDGAKLVRDAFLLAKEKSPCIIFIDEIDA---IGTKRFDSEVSGDREV 286
Query: 324 KMTLSGLLNVIDGLWSCCGEERI-IIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFE 382
+ T+ LLN +DG S ++RI +I TN + LDPAL+R GR+D + + T E
Sbjct: 287 QRTMLELLNQLDGFSS---DDRIKVIAATNRADILDPALMRSGRLDRKIEFPHPT----E 339
Query: 383 QLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQ-N 441
+ L I K+ + + + G K VE G++ +D T+ N
Sbjct: 340 EARGRILQIHSRKMNVNADVNFEELARSTDDFNGAQLKAVCVEA---GMLALRRDATEVN 396
Query: 442 DQNINQDNASQKEKKNQDHEY 462
++ N+ + KK Y
Sbjct: 397 HEDFNEGIIQVQAKKKASLNY 417
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,471,902
Number of Sequences: 164201
Number of extensions: 2178990
Number of successful extensions: 8844
Number of sequences better than 10.0: 535
Number of HSP's better than 10.0 without gapping: 366
Number of HSP's successfully gapped in prelim test: 171
Number of HSP's that attempted gapping in prelim test: 8146
Number of HSP's gapped (non-prelim): 655
length of query: 467
length of database: 59,974,054
effective HSP length: 114
effective length of query: 353
effective length of database: 41,255,140
effective search space: 14563064420
effective search space used: 14563064420
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0111b.4


