

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
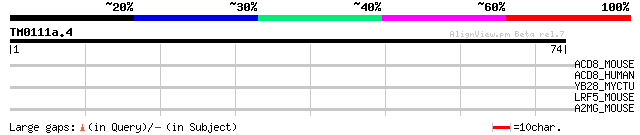
Query= TM0111a.4
(74 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ACD8_MOUSE (Q9D7B6) Acyl-CoA dehydrogenase family member 8, mito... 28 4.3
ACD8_HUMAN (Q9UKU7) Acyl-CoA dehydrogenase family member 8, mito... 28 4.3
YB28_MYCTU (O06580) Hypothetical protein Rv1128c/MT1160 27 9.6
LRF5_MOUSE (Q8BXA0) Leucine rich repeat and fibronectin type III... 27 9.6
A2MG_MOUSE (Q61838) Alpha-2-macroglobulin precursor (Alpha-2-M) 27 9.6
>ACD8_MOUSE (Q9D7B6) Acyl-CoA dehydrogenase family member 8,
mitochondrial precursor (EC 1.3.99.-) (ACAD-8)
(Isobutyryl-CoA dehydrogenase)
Length = 413
Score = 28.1 bits (61), Expect = 4.3
Identities = 11/27 (40%), Positives = 13/27 (47%)
Query: 13 AWMLAMNVNHENNHELCAPECTSTSIA 39
AWM+ N E H+ C P CT A
Sbjct: 128 AWMIDSFGNEEQRHKFCPPLCTMEKFA 154
>ACD8_HUMAN (Q9UKU7) Acyl-CoA dehydrogenase family member 8,
mitochondrial precursor (EC 1.3.99.-) (ACAD-8)
(Isobutyryl-CoA dehydrogenase) (Activator-recruited
cofactor 42 kDa component) (ARC42)
Length = 415
Score = 28.1 bits (61), Expect = 4.3
Identities = 11/27 (40%), Positives = 13/27 (47%)
Query: 13 AWMLAMNVNHENNHELCAPECTSTSIA 39
AWM+ N E H+ C P CT A
Sbjct: 130 AWMIDSFGNEEQRHKFCPPLCTMEKFA 156
>YB28_MYCTU (O06580) Hypothetical protein Rv1128c/MT1160
Length = 451
Score = 26.9 bits (58), Expect = 9.6
Identities = 11/25 (44%), Positives = 15/25 (60%)
Query: 31 PECTSTSIAVWDIPKCSGLCNPEKR 55
PE +T AVW G+CNPE++
Sbjct: 215 PELRATIEAVWAKLAAPGMCNPEQK 239
>LRF5_MOUSE (Q8BXA0) Leucine rich repeat and fibronectin type III
domain containing protein 5 precursor
Length = 719
Score = 26.9 bits (58), Expect = 9.6
Identities = 13/43 (30%), Positives = 19/43 (43%)
Query: 20 VNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQM 62
++ E++ E CA T W IP+ LC P H+M
Sbjct: 254 LSREDDLETCASPALLTGRYFWSIPEEEFLCEPPLITRHTHEM 296
>A2MG_MOUSE (Q61838) Alpha-2-macroglobulin precursor (Alpha-2-M)
Length = 1495
Score = 26.9 bits (58), Expect = 9.6
Identities = 21/59 (35%), Positives = 27/59 (45%), Gaps = 3/59 (5%)
Query: 8 DVTIGAWMLAMNVN-HENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQMESC 65
D+ I LA+ V HEN+H +C E + S AV PK G N + L E C
Sbjct: 839 DLEISPDFLAVPVGGHENSHCICGNERKTVSWAV--TPKSLGEVNFTRTAEALESQELC 895
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.127 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,898,333
Number of Sequences: 164201
Number of extensions: 281197
Number of successful extensions: 537
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 533
Number of HSP's gapped (non-prelim): 5
length of query: 74
length of database: 59,974,054
effective HSP length: 50
effective length of query: 24
effective length of database: 51,764,004
effective search space: 1242336096
effective search space used: 1242336096
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0111a.4


