

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.1
(863 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
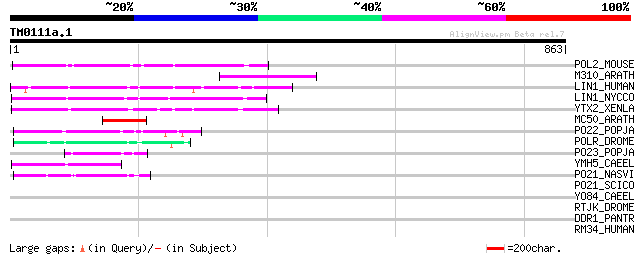
Score E
Sequences producing significant alignments: (bits) Value
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 132 4e-30
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 123 2e-27
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 115 4e-25
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 113 2e-24
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 108 6e-23
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 79 4e-14
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 63 4e-09
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 62 5e-09
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 62 5e-09
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 55 8e-07
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 52 7e-06
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 40 0.026
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 38 0.10
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 35 0.65
DDR1_PANTR (Q7YR43) Epithelial discoidin domain receptor 1 precu... 32 5.5
RM34_HUMAN (Q9BQ48) 39S ribosomal protein L34, mitochondrial pre... 32 9.4
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 132 bits (332), Expect = 4e-30
Identities = 105/406 (25%), Positives = 183/406 (44%), Gaps = 24/406 (5%)
Query: 5 EGYLPPFINETVVALVPKVDK-PENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQ 63
EG LP E + L+PK K P I RPIS N K++ KIL N+++ H+ +
Sbjct: 528 EGTLPNSFYEATITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRIQEHIKAIIHP 587
Query: 64 AQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTS 123
Q FI G Q NI + H++ N K K M I L+ KA+D+++ F+ L
Sbjct: 588 DQVGFIPGMQGWFNIRKSINVIHYI-NKLKDKNH-MIISLDAEKAFDKIQHPFMIKVLER 645
Query: 124 FGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLL 183
G +++ I + KVNGE + + + G RQG PLSPYLF + E ++ +
Sbjct: 646 SGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIVLEVLARAI 705
Query: 184 LKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMIS 243
+ +E I+G+++ E I+ L ADD + + + +L++++N + G I+
Sbjct: 706 RQQKE---IKGIQIGKEEVKIS--LLADDMIVYISDPKNSTRELLNLINSFGEVVGYKIN 760
Query: 244 IEKS-GIIFSRNTLSSTKHQISSILRMSAWDSPGKYLG--LPAEWGKSRNNTLNWIKLRV 300
KS ++++N + +I S + KYLG L E + +K +
Sbjct: 761 SNKSMAFLYTKN--KQAEKEIRETTPFSIVTNNIKYLGVTLTKEVKDLYDKNFKSLKKEI 818
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSI--IRFPQNFCQAICASIARFWW--R 356
+ + WK+ + G+ ++K + + Y + I+ P F + +I +F W +
Sbjct: 819 KEDLRRWKDLPCSWIGRINIVKMAILPKAIYRFNAIPIKIPTQFFNELEGAICKFVWNNK 878
Query: 357 SPGKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIY 402
P + + + + GG+ D A++ K AW Y
Sbjct: 879 KPRIAKSL-------LKDKRTSGGITMPDLKLYYRAIVIKTAWYWY 917
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 123 bits (309), Expect = 2e-27
Identities = 61/151 (40%), Positives = 85/151 (55%), Gaps = 1/151 (0%)
Query: 327 AMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWKDWHFISKAK-CKGGLGFKD 385
A+ Y MS R + C+ + +++ FWW S +R I W W + K+K GGLGF+D
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 386 FSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNSSFLQSGRTARNSWGWQSILQGKEF 445
N ALLAKQ++RI QP L R L+ YFP+SS ++ R S+ W+SI+ G+E
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGREL 121
Query: 446 LKEAGLWIIGDGSEVKVWKDKWVSDQVLQAP 476
L L IGDG KVW D+W+ D+ P
Sbjct: 122 LSRGLLRTIGDGIHTKVWLDRWIMDETPLPP 152
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 115 bits (289), Expect = 4e-25
Identities = 115/454 (25%), Positives = 193/454 (42%), Gaps = 35/454 (7%)
Query: 1 SFFQEGYLPPFINETVVALVPK----VDKPENIGQLRPISCCNFLYKVITKILVNKLKPH 56
S +EG LP E + L+PK K EN RPIS N K++ KIL N+++ H
Sbjct: 497 SIEKEGILPNSFYEASIILIPKPGRDTTKKENF---RPISLMNIDAKILNKILANQIQQH 553
Query: 57 MNRLATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDF 116
+ +L Q FI Q NI + + T T M I ++ KA+D+++ F
Sbjct: 554 IKKLIHHDQVGFIPAMQGWFNIRKSINIIQHINRT--KDTNHMIISIDAEKAFDKIQQPF 611
Query: 117 LKACLTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAA 176
+ L G ++ I + +NG+ + + G RQG PLSP L +
Sbjct: 612 MLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAPPLKTGTRQGCPLSPLLPNIVL 671
Query: 177 ETMSHLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTL 236
E ++ + ++E I+G+++ E + LFADD + + + A L+ ++++++
Sbjct: 672 EVLARAI---RQEKEIKGIQLGKE--EVKLSLFADDMIVYLENPIVSAQNLLKLISNFSK 726
Query: 237 ASGQMISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAE------WGKSRN 290
SG I+++KS N T+ QI S L + KYLG+ + ++
Sbjct: 727 VSGYKINVQKSQAFLYTNN-RQTESQIMSELPFTIASKRIKYLGIQLTRDVKDLFKENYK 785
Query: 291 NTLNWIKLRVLDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSI--IRFPQNFCQAICA 348
LN IK + + WK + G+ ++K + Y + I+ P F +
Sbjct: 786 PLLNEIK----EDTNKWKNIPCSWVGRINIVKMAILPKVIYRFNAIPIKLPMTFFTELEK 841
Query: 349 SIARFWWRSPGKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDA- 407
+ +F W ++ H K K GG+ DF A + K AW Y D
Sbjct: 842 TTLKFIW----NQKRAHIAKSTLSQKNKA-GGITLPDFKLYYKATVTKTAWYWYQNRDID 896
Query: 408 LWVRTLKGLYFPN-SSFLQSGRTARN-SWGWQSI 439
W RT P+ ++L + +N WG S+
Sbjct: 897 QWNRTEPSEIMPHIYNYLIFDKPEKNKQWGKDSL 930
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 113 bits (282), Expect = 2e-24
Identities = 96/401 (23%), Positives = 171/401 (41%), Gaps = 18/401 (4%)
Query: 4 QEGYLPPFINETVVALVPKVDK-PENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLAT 62
+EG LP E + L+PK K P RPIS N K++ KIL N+++ H+ ++
Sbjct: 500 KEGILPNTFYEANITLIPKPGKDPTRKENYRPISLMNIDAKILNKILTNRIQQHIKKIIH 559
Query: 63 QAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLT 122
Q FI G Q NI + + N K K + M + ++ KA+D ++ F+ L
Sbjct: 560 HDQVGFIPGSQGWFNIRKSINVIQHI-NKLKNK-DHMILSIDAEKAFDNIQHPFMIRTLK 617
Query: 123 SFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHL 182
G ++ I + + +NG K + G RQG PLSP LF + E L
Sbjct: 618 KIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEV---L 674
Query: 183 LLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMI 242
+ +EE I+G+ + E I LFADD + + + R+ +L+ ++ +Y+ SG I
Sbjct: 675 AIAIREEKAIKGIHIGSE--EIKLSLFADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKI 732
Query: 243 SIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLG--LPAEWGKSRNNTLNWIKLRV 300
+ KS + F + + + + + KYLG L + ++ +
Sbjct: 733 NTHKS-VAFIYTNNNQAEKTVKDSIPFTVVPKKMKYLGVYLTKDVKDLYKENYETLRKEI 791
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSI--IRFPQNFCQAICASIARFWWRSP 358
+ ++ WK + G+ ++K + + Y + I+ P ++ + + I F W
Sbjct: 792 AEDVNKWKNIPCSWLGRINIVKMSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIWNQK 851
Query: 359 GKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAW 399
+ +S GG+ D +++ K AW
Sbjct: 852 KPQIA-----KTLLSNKNKAGGITLPDLRLYYKSIVIKTAW 887
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 108 bits (270), Expect = 6e-23
Identities = 107/423 (25%), Positives = 178/423 (41%), Gaps = 29/423 (6%)
Query: 3 FQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLAT 62
F++G LP V++L+PK I RP+S + YK++ K + +LK + +
Sbjct: 496 FKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKSVLAEVIH 555
Query: 63 QAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLT 122
QS + GR I DN+ + + HF + T + L+ KA+DR++ +L L
Sbjct: 556 PDQSYTVPGRTIFDNVFLIRDLLHFARRT---GLSLAFLSLDQEKAFDRVDHQYLIGTLQ 612
Query: 123 SFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHL 182
++ F ++V + A K+N ++ + RG+RQG PLS L+ LA E L
Sbjct: 613 AYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCL 672
Query: 183 LLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAK--ANREEAYQLISILNDYTLASGQ 240
L K R+ GL + + +ADD + A+ + E A + + Y AS
Sbjct: 673 LRK-----RLTGLVLKEPDMRVVLSAYADDVILVAQDLVDLERAQECQEV---YAAASSA 724
Query: 241 MISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSP-GKYLGLPAEWGKSRNNTLNWIKLR 299
I+ KS + S + R +W+S KYLG+ + + N+I+L
Sbjct: 725 RINWSKSSGLLEG---SLKVDFLPPAFRDISWESKIIKYLGVYLS-AEEYPVSQNFIELE 780
Query: 300 --VLDKIDGWK--EKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWW 355
VL ++ WK K+L+ G+ ++I ++ + Y + + Q F I + F W
Sbjct: 781 ECVLTRLGKWKGFAKVLSMRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRLLDFLW 840
Query: 356 RSPGKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWR-IYTQPDALWVRTLK 414
G HW S +GG G + +Q R +Y P W
Sbjct: 841 ------IGKHWVSAGVSSLPLKEGGQGVVCIRSQVHTFRLQQIQRYLYADPSPQWCTLAS 894
Query: 415 GLY 417
Y
Sbjct: 895 SFY 897
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 79.3 bits (194), Expect = 4e-14
Identities = 37/69 (53%), Positives = 48/69 (68%)
Query: 145 FKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPI 204
F +NG V P RG+RQG PLSPYLF+L E +S L +AQE+GR+ G+R++ P I
Sbjct: 12 FIINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSPRI 71
Query: 205 THLLFADDS 213
HLLFADD+
Sbjct: 72 NHLLFADDT 80
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 62.8 bits (151), Expect = 4e-09
Identities = 69/305 (22%), Positives = 127/305 (41%), Gaps = 26/305 (8%)
Query: 6 GYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQ 65
G++P L+PK EN RPI+ + L +++ +IL +L+ + Q
Sbjct: 50 GHVPTPWTAMRTTLIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVELHPAQKG 109
Query: 66 SAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFG 125
A I G + +L ++ + + + + + L++ KA+D + + L G
Sbjct: 110 YARIDGTLVNSLLLDT-----YISSRREQRKTYNVVSLDVRKAFDTVSHSSICRALQRLG 164
Query: 126 FSEKWVDRIMQCVTGASFRFKVN-GEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLL 184
E + I ++ ++ +V G ++++ +RG++QG PLSP+LF + LL
Sbjct: 165 IDEGTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLF---NAVLDELLC 221
Query: 185 KAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM--- 241
Q I G + E P+ L FADD L + L ++ N + L +
Sbjct: 222 SLQSTPGIGG-TIGEEKIPV--LAFADDLLLLEDNDVLLPTTLATVANFFRLRGMSLNAK 278
Query: 242 ----ISIEKSGIIFSRNTLSSTKHQISSIL-----RMSAWDSPGKYLGLPAEWGKSRNNT 292
IS+ SG + T + + ++L RM + G + GL + N
Sbjct: 279 KSVSISVAASGGVCIPRTKPFLR--VDNVLLPVTDRMQTFRYLGHFFGLSGAAKPTVYNL 336
Query: 293 LNWIK 297
W+K
Sbjct: 337 SRWLK 341
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 62.4 bits (150), Expect = 5e-09
Identities = 72/288 (25%), Positives = 112/288 (38%), Gaps = 30/288 (10%)
Query: 6 GYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQ 65
G LP I +PK + RPIS + L + + IL +L +N Q
Sbjct: 389 GNLPHSIRLARTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSINW--DPRQ 446
Query: 66 SAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFG 125
F+ DN I L+++ K L+++KA+D L + L ++G
Sbjct: 447 RGFLPTDGCADNATIVDLV---LRHSHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYG 503
Query: 126 FSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLK 185
+ +VD + G +G S+ VP RG++QG PLSP LF L + + L
Sbjct: 504 APKGFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTL-- 561
Query: 186 AQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIE 245
E G G + FADD + FA+ R L+ D+ G ++ +
Sbjct: 562 PSEIGAKVGNAIT------NAAAFADDLVLFAE-TRMGLQVLLDKTLDFLSIVGLKLNAD 614
Query: 246 KS---GI---------IFSRNTLSSTKHQISSILRMSAWDSPGKYLGL 281
K GI + + +I S+ R W KYLG+
Sbjct: 615 KCFTVGIKGQPKQKCTVLEAQSFYVGSSEIPSLKRTDEW----KYLGI 658
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 62.4 bits (150), Expect = 5e-09
Identities = 41/130 (31%), Positives = 65/130 (49%), Gaps = 8/130 (6%)
Query: 86 HFLK-NTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFGFSEKWVDRIMQCVTGASFR 144
H++K KGKT + + L++ KA+D + + + +FG + D IM +T A
Sbjct: 11 HYIKLRRLKGKT-YNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTN 69
Query: 145 FKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPI 204
V G + ++ + G++QG PLSP LF + + L+ + +E G M P C I
Sbjct: 70 IVVGGRTTNKIYIRNGVKQGDPLSPVLFNI---VLDELVTRLNDEQ--PGASMTPAC-KI 123
Query: 205 THLLFADDSL 214
L FADD L
Sbjct: 124 ASLAFADDLL 133
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 55.1 bits (131), Expect = 8e-07
Identities = 37/173 (21%), Positives = 75/173 (42%), Gaps = 4/173 (2%)
Query: 3 FQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLAT 62
F + +P V+ +PK P + RPIS + +++ +I+ ++++ + L +
Sbjct: 656 FSDSAIPNRWKHAVIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLS 715
Query: 63 QAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLT 122
Q F++ R +++ + +H + K + + + KA+D++ L L
Sbjct: 716 PHQHGFLNFRSCPSSLVRSISLYH---SILKNEKSLDILFFDFAKAFDKVSHPILLKKLA 772
Query: 123 SFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVP-QRGIRQGYPLSPYLFVL 174
FG + + + +F K+N VS P G+ QG P LF+L
Sbjct: 773 LFGLDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFIL 825
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 52.0 bits (123), Expect = 7e-06
Identities = 54/213 (25%), Positives = 89/213 (41%), Gaps = 13/213 (6%)
Query: 6 GYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQ 65
G P ++ L+PK + RP+S + + +IL N++ H L Q
Sbjct: 363 GQCPRRYLDSRTVLIPKEPGTMDPACFRPLSIASVALRHFHRILANRIGEH--GLLDTRQ 420
Query: 66 SAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFG 125
AFI + +N + + KG ++AI L++ KA+D +E + L
Sbjct: 421 RAFIVADGVAENTSLLSAMIKEARMKIKGL--YIAI-LDVKKAFDSVEHRSILDALRRKK 477
Query: 126 FSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLK 185
+ + IM + R +V + + P RG+RQG PLSP LF + + L
Sbjct: 478 LPLEMRNYIMWVYRNSKTRLEVVKTKGRWIRPARGVRQGDPLSPLLFNCVMDAVLRRL-- 535
Query: 186 AQEEGRIQGLRMAPECPPITHLLFADDSLFFAK 218
+ G + G I L+FADD + A+
Sbjct: 536 PENTGFLMG------AEKIGALVFADDLVLLAE 562
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 40.0 bits (92), Expect = 0.026
Identities = 45/186 (24%), Positives = 70/186 (37%), Gaps = 9/186 (4%)
Query: 6 GYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQ 65
G +P I + PK++ + G RP+S C+ + + KIL + Q
Sbjct: 204 GRVPSPIKGSRTVFTPKIEGGPDPGVFRPLSICSVILREFNKILARRFVSCYTYDERQTA 263
Query: 66 SAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFG 125
I G I ++L A + + + E L++ KA++ + L +T G
Sbjct: 264 YLPIDGVCINVSMLTA-----IIAEAKRLRKELHIAILDLVKAFNSVYHSALIDAITEAG 318
Query: 126 FSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLK 185
VD I + G+ + G+ QG PLS LF LA E L
Sbjct: 319 CPPGVVDYIADMYNNVITEMQFEGKCELASI-LAGVYQGDPLSGPLFTLAYEKALRAL-- 375
Query: 186 AQEEGR 191
EGR
Sbjct: 376 -NNEGR 380
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 38.1 bits (87), Expect = 0.10
Identities = 23/78 (29%), Positives = 40/78 (50%)
Query: 103 LEMNKAYDRLEWDFLKACLTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIR 162
L+ +KA+D++ D L LTS ++ + + +T SF+ KV +S+ G+
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 163 QGYPLSPYLFVLAAETMS 180
QG +SP LF + +S
Sbjct: 83 QGSVISPVLFGIFVNEIS 100
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 35.4 bits (80), Expect = 0.65
Identities = 40/181 (22%), Positives = 72/181 (39%), Gaps = 5/181 (2%)
Query: 1 SFFQEGYLPPFINETVVALVPKVDK-PENIGQLRPISCCNFLYKVITKILVNKLKPHMNR 59
S GY P + ++ K K P ++ RP S L K++ ++++N+L +
Sbjct: 483 SVLDVGYFPKAWKSASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTCKD- 541
Query: 60 LATQAQSAFISGRQIQDNI-LIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLK 118
T+A F G ++Q H +F + K + L++ +A+DR+ W
Sbjct: 542 -VTKAIPKFQFGFRLQHGTPEQLHRVVNFALEAMENKEYAVGAFLDIQQAFDRV-WHPGL 599
Query: 119 ACLTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAET 178
F + + + +F V+G S G+ QG L P L+ + A
Sbjct: 600 LYKAKRLFPPQLYLVVKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLYSVFASD 659
Query: 179 M 179
M
Sbjct: 660 M 660
>DDR1_PANTR (Q7YR43) Epithelial discoidin domain receptor 1
precursor (EC 2.7.1.112) (Tyrosine kinase DDR)
(Discoidin receptor tyrosine kinase)
Length = 909
Score = 32.3 bits (72), Expect = 5.5
Identities = 41/156 (26%), Positives = 62/156 (39%), Gaps = 33/156 (21%)
Query: 370 HFISKAKCKGGLGFKDFSTMNLALLAKQAW------RIYTQPDALWVRTLKGLYFPNSS- 422
HF AKC+ LG +D + + + A +W R + D W G FP
Sbjct: 25 HF-DPAKCRYALGMQDRTIPDSDISASSSWSDSTAARHSSDGDGAWCPA--GSVFPKEEE 81
Query: 423 FLQS-----------GRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQ 471
+LQ G R++ G GKEF + L DG WKD+W +
Sbjct: 82 YLQVDLQRLHLVALVGTQGRHAGGL-----GKEFSRSYRLRYSRDGRRWMGWKDRW-GQE 135
Query: 472 VLQAPEAGDAELTVNSLILPDVKQWNLPKLLSTFPR 507
V+ E D E ++L D+ + +L+ +PR
Sbjct: 136 VISGNE--DPE----GVVLKDLGPPMVARLVRFYPR 165
>RM34_HUMAN (Q9BQ48) 39S ribosomal protein L34, mitochondrial
precursor (L34mt) (MRP-L34)
Length = 92
Score = 31.6 bits (70), Expect = 9.4
Identities = 20/72 (27%), Positives = 33/72 (45%), Gaps = 10/72 (13%)
Query: 264 SSILRMSAWDSPGKYLGLPAEW---------GKSRNNTLNWIKLRVLDKIDGWKEKLLNQ 314
S+ L W P +LG P W GK+R N ++ +K GW +L
Sbjct: 15 SAALLGGRWLQPRAWLGFPDAWGLPTPQQARGKARGNEYQPSNIKRKNK-HGWVRRLSTP 73
Query: 315 AGKEVLIKAVLQ 326
AG +V+++ +L+
Sbjct: 74 AGVQVILRRMLK 85
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 102,524,009
Number of Sequences: 164201
Number of extensions: 4251052
Number of successful extensions: 8409
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 8379
Number of HSP's gapped (non-prelim): 21
length of query: 863
length of database: 59,974,054
effective HSP length: 119
effective length of query: 744
effective length of database: 40,434,135
effective search space: 30082996440
effective search space used: 30082996440
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0111a.1


