

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
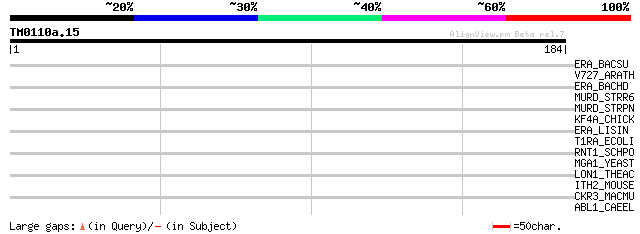
Query= TM0110a.15
(184 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ERA_BACSU (P42182) GTP-binding protein era homolog (Bex protein) 37 0.033
V727_ARATH (Q9M376) Vesicle-associated membrane protein 727 (AtV... 34 0.17
ERA_BACHD (Q9KD52) GTP-binding protein era homolog 32 0.82
MURD_STRR6 (Q8DQM2) UDP-N-acetylmuramoylalanine--D-glutamate lig... 30 2.4
MURD_STRPN (Q97RU8) UDP-N-acetylmuramoylalanine--D-glutamate lig... 30 2.4
KF4A_CHICK (Q90640) Chromosome-associated kinesin KIF4A (Chromok... 30 2.4
ERA_LISIN (Q92BP8) GTP-binding protein era homolog 30 4.1
T1RA_ECOLI (Q07736) Type I restriction enzyme EcoAI R protein (E... 29 5.3
RNT1_SCHPO (Q09820) Regulator of nonsense transcripts 1 homolog 29 5.3
MGA1_YEAST (P53050) MGA1 protein 29 5.3
LON1_THEAC (Q9HJ89) Putative protease La homolog type 1 (EC 3.4.... 29 5.3
ITH2_MOUSE (Q61703) Inter-alpha-trypsin inhibitor heavy chain H2... 29 5.3
CKR3_MACMU (P56483) C-C chemokine receptor type 3 (C-C CKR-3) (C... 29 7.0
ABL1_CAEEL (P03949) Tyrosine-protein kinase abl-1 (EC 2.7.1.112) 28 9.1
>ERA_BACSU (P42182) GTP-binding protein era homolog (Bex protein)
Length = 301
Score = 36.6 bits (83), Expect = 0.033
Identities = 31/106 (29%), Positives = 54/106 (50%), Gaps = 4/106 (3%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELV 141
+LI +++LHL V I +G + V +AT++ R S KG+ L+
Sbjct: 195 ELIREKVLHLTREEIPHSIAVAIESIKGQDNGSVHV-AATIVVERDSQKGIVIGKKGSLL 253
Query: 142 -ELEKDAVE--EALVGSAIQIEGFRRVWLDLRSRLSLLHESRSEKD 184
E+ K A EAL+GS + +E + +V D R+++S L + ++D
Sbjct: 254 KEVGKRARADIEALLGSRVYLELWVKVQKDWRNKMSQLRDFGFKED 299
>V727_ARATH (Q9M376) Vesicle-associated membrane protein 727
(AtVAMP727)
Length = 240
Score = 34.3 bits (77), Expect = 0.17
Identities = 15/61 (24%), Positives = 31/61 (50%), Gaps = 2/61 (3%)
Query: 14 SLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDS--VNDSTA 71
S ++++ + T+ FL+D F++ + D +S FL R++ K+ ++ ND
Sbjct: 44 SKYTYSCDGHTFNFLVDNGFVFLVVADESTGRSVPFVFLERVKEDFKKRYEASIKNDERH 103
Query: 72 P 72
P
Sbjct: 104 P 104
>ERA_BACHD (Q9KD52) GTP-binding protein era homolog
Length = 304
Score = 32.0 bits (71), Expect = 0.82
Identities = 27/106 (25%), Positives = 50/106 (46%), Gaps = 3/106 (2%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATEL- 140
+LI +++LHL V I + + + V AT++ R+S KG+ ++
Sbjct: 197 ELIREKVLHLTREEIPHSIAVVIEQIKRRQHQDTVYIGATIVVERSSQKGIIIGKQGKML 256
Query: 141 --VELEKDAVEEALVGSAIQIEGFRRVWLDLRSRLSLLHESRSEKD 184
V + A EAL+GS + +E + +V D R++ L + +D
Sbjct: 257 KEVGQQARADIEALLGSKVFLELWVKVQKDWRNKPQHLRDYGFRED 302
>MURD_STRR6 (Q8DQM2) UDP-N-acetylmuramoylalanine--D-glutamate ligase
(EC 6.3.2.9) (UDP-N-acetylmuramoyl-L-alanyl-D-glutamate
synthetase) (D-glutamic acid adding enzyme)
Length = 450
Score = 30.4 bits (67), Expect = 2.4
Identities = 21/76 (27%), Positives = 32/76 (41%), Gaps = 3/76 (3%)
Query: 91 LDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEE 150
LD GN V I+ GLKK ++ SA + G+A AT++ + + A E
Sbjct: 356 LDRGNEFDELVPDIT---GLKKMVILGQSAERVKRAADKAGVAYVEATDIADATRKAYEL 412
Query: 151 ALVGSAIQIEGFRRVW 166
A G + + W
Sbjct: 413 ATQGDVVLLSPANASW 428
>MURD_STRPN (Q97RU8) UDP-N-acetylmuramoylalanine--D-glutamate ligase
(EC 6.3.2.9) (UDP-N-acetylmuramoyl-L-alanyl-D-glutamate
synthetase) (D-glutamic acid adding enzyme)
Length = 450
Score = 30.4 bits (67), Expect = 2.4
Identities = 21/76 (27%), Positives = 32/76 (41%), Gaps = 3/76 (3%)
Query: 91 LDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEE 150
LD GN V I+ GLKK ++ SA + G+A AT++ + + A E
Sbjct: 356 LDRGNEFDELVPDIT---GLKKMVILGQSAERVKRAADKAGVAYVEATDIADATRKAYEL 412
Query: 151 ALVGSAIQIEGFRRVW 166
A G + + W
Sbjct: 413 ATQGDVVLLSPANASW 428
>KF4A_CHICK (Q90640) Chromosome-associated kinesin KIF4A
(Chromokinesin)
Length = 1225
Score = 30.4 bits (67), Expect = 2.4
Identities = 20/49 (40%), Positives = 26/49 (52%), Gaps = 1/49 (2%)
Query: 107 DEGLKKKKMVVDSAT-VIANRTSAKGLAAAAATELVELEKDAVEEALVG 154
DE LK+ V+ + V+A S AA AATE+ E+DA EA G
Sbjct: 459 DEELKENVEVIRNLQQVLAQFQSESAAAAEAATEMANAEQDAAGEAETG 507
>ERA_LISIN (Q92BP8) GTP-binding protein era homolog
Length = 301
Score = 29.6 bits (65), Expect = 4.1
Identities = 23/100 (23%), Positives = 48/100 (48%), Gaps = 3/100 (3%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELV 141
+LI +++L L V I E K + + +AT+I R++ KG+ +++
Sbjct: 194 ELIREQVLQLTREEVPHSVAVVIEGIEKNPKTEKLTINATIIVERSTQKGIIIGKQGQML 253
Query: 142 E---LEKDAVEEALVGSAIQIEGFRRVWLDLRSRLSLLHE 178
+ + E+L+GS + +E + +V + R + LH+
Sbjct: 254 KQIGMRARKEIESLLGSKVFLEIWVKVQKNWRDKEHYLHD 293
>T1RA_ECOLI (Q07736) Type I restriction enzyme EcoAI R protein (EC
3.1.21.3) (R.EcoAI)
Length = 810
Score = 29.3 bits (64), Expect = 5.3
Identities = 40/161 (24%), Positives = 65/161 (39%), Gaps = 25/161 (15%)
Query: 34 IYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILHLDD 93
++F I D + K+ L R ++ +D+ + A P ++ F+ L+EI D+
Sbjct: 533 LWFTILDFK--KATELFADERFDGIPEKVMDTTPEDIADP----ESDFEEKLEEISEHDE 586
Query: 94 ----GNRSPPAV---------VGISHDEGLKK-KKMVVDSATV--IANRTSAKGLAAAAA 137
G PPA VG +E KK +K V+ V IA R
Sbjct: 587 EQVTGVDEPPAPPYQVTDTDDVGPLPEEDEKKIRKFHVNGVAVGVIAQRVQYYDADGKLV 646
Query: 138 TELVELEKDAVEEALVGSAIQIEGFRRVWLDLRSRLSLLHE 178
TE KD + L+ ++ F R W D + +++HE
Sbjct: 647 TESF---KDYTRKTLLKEYASLDDFTRKWQDADRKEAIIHE 684
>RNT1_SCHPO (Q09820) Regulator of nonsense transcripts 1 homolog
Length = 925
Score = 29.3 bits (64), Expect = 5.3
Identities = 14/55 (25%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Query: 30 DPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDS--VNDSTAPPPLSLQTPFD 82
D P +++A F + +FLNR +S E + + + + P + + TP+D
Sbjct: 671 DSPLMFYANFGQEELSASGTSFLNRTEASTCEKIVTTFLRSNVLPEQIGIVTPYD 725
>MGA1_YEAST (P53050) MGA1 protein
Length = 456
Score = 29.3 bits (64), Expect = 5.3
Identities = 27/117 (23%), Positives = 48/117 (40%), Gaps = 16/117 (13%)
Query: 70 TAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSA 129
T PP S TPF ++ + + + SP +G+ ++ KK + + N
Sbjct: 294 TVPPYSSYSTPFPSMMNSLSN--SASNSP--ALGVCNNNVTLPKKSNISERQALDNHIQT 349
Query: 130 KGLAAAAATELVELEKDAVEE----ALVGSAIQIEGFRRVWLDLRSRLSLLHESRSE 182
+ + T+L+E ++ + L A+ DLR+ LSLL S+ E
Sbjct: 350 LKNSLSTITDLIEKHINSASQDENKTLTNDAMN--------KDLRTSLSLLQNSKEE 398
>LON1_THEAC (Q9HJ89) Putative protease La homolog type 1 (EC
3.4.21.-)
Length = 657
Score = 29.3 bits (64), Expect = 5.3
Identities = 16/45 (35%), Positives = 26/45 (57%)
Query: 101 VVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEK 145
+V ++ D + +KK VV +A VIA + AK L A +E++K
Sbjct: 380 LVRVAGDIAVSQKKTVVTAADVIAAKNLAKPLEQQIADRSIEIKK 424
>ITH2_MOUSE (Q61703) Inter-alpha-trypsin inhibitor heavy chain H2
precursor (ITI heavy chain H2) (Inter-alpha-inhibitor
heavy chain 2)
Length = 946
Score = 29.3 bits (64), Expect = 5.3
Identities = 25/83 (30%), Positives = 38/83 (45%), Gaps = 11/83 (13%)
Query: 9 KGSTHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVND 68
KG+ S F+ TVN T+T I + A++ K +T ++ TLD N
Sbjct: 112 KGAFISNFTMTVNGMTFTSSIKEKTVGRALYSQARAKGKTAGWVR------SRTLDMENF 165
Query: 69 ST---APPPLSLQTPFDLILQEI 88
+T PP +Q F+L QE+
Sbjct: 166 NTEVNIPPGAKVQ--FELHYQEV 186
>CKR3_MACMU (P56483) C-C chemokine receptor type 3 (C-C CKR-3)
(CC-CKR-3) (CCR-3) (CCR3) (CKR3)
Length = 355
Score = 28.9 bits (63), Expect = 7.0
Identities = 21/63 (33%), Positives = 29/63 (45%), Gaps = 1/63 (1%)
Query: 16 FSHT-VNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAPPP 74
+SH VN Y F+ + Y F HRH ++ + S E SV+ STA P
Sbjct: 291 YSHCCVNPVIYAFVGERFRKYLRHFFHRHVLMHLGKYIPFLPSEKLERTSSVSPSTAEPE 350
Query: 75 LSL 77
LS+
Sbjct: 351 LSI 353
>ABL1_CAEEL (P03949) Tyrosine-protein kinase abl-1 (EC 2.7.1.112)
Length = 1224
Score = 28.5 bits (62), Expect = 9.1
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 5/69 (7%)
Query: 8 SKGSTHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKE-----T 62
++G+ S S TV+N + L P + + + H S FL +IRS LK+
Sbjct: 743 NEGAGSSSLSRTVSNDSLDTLPLPDSMNSSTYVKMHPASGENVFLRQIRSKLKKRSETPE 802
Query: 63 LDSVNDSTA 71
LD ++ TA
Sbjct: 803 LDHIDSDTA 811
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,455,044
Number of Sequences: 164201
Number of extensions: 762912
Number of successful extensions: 2323
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2321
Number of HSP's gapped (non-prelim): 14
length of query: 184
length of database: 59,974,054
effective HSP length: 104
effective length of query: 80
effective length of database: 42,897,150
effective search space: 3431772000
effective search space used: 3431772000
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0110a.15


