

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.13
(1389 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
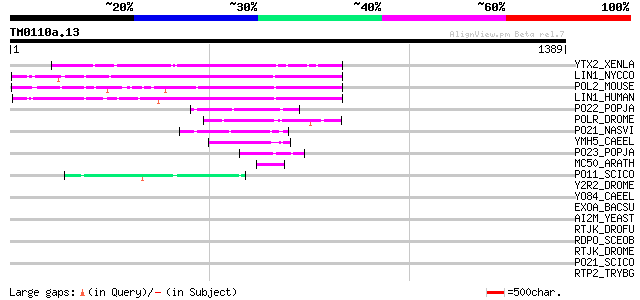
Score E
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 188 1e-46
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 173 3e-42
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 156 4e-37
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 153 4e-36
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 89 6e-17
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 85 1e-15
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 69 1e-10
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 64 2e-09
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 64 2e-09
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 62 1e-08
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 49 1e-04
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 45 0.001
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 44 0.004
EXOA_BACSU (P37454) Exodeoxyribonuclease (EC 3.1.11.2) 42 0.009
AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein 40 0.058
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 39 0.099
RDPO_SCEOB (P19593) Probable reverse transcriptase (EC 2.7.7.49) 37 0.38
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 37 0.49
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 37 0.49
RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa prot... 36 0.84
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 188 bits (477), Expect = 1e-46
Identities = 196/756 (25%), Positives = 342/756 (44%), Gaps = 47/756 (6%)
Query: 106 IMNVYGPNVDSERYDFFNLLSEAL-SLNNCFCKIMGGDFNAILNEGERIGIQM--GVESS 162
+MNVY P ER FF LS + ++++ I+GGDFN L+ +R + ES
Sbjct: 106 LMNVYAPTTGPERARFFESLSAYMETIDSDEALIIGGDFNYTLDARDRNVPKKRDSSESV 165
Query: 163 FSDFVADSNLIDLPLQSDK----FTWYSSRAGGL-WSRIDRWLVSEDAINKFDGISQSVE 217
+ +A +L+D+ + + FT+ R G + SRIDR +S +++ S ++
Sbjct: 166 LRELIAHFSLVDVWREQNPETVAFTYVRVRDGHVSQSRIDRIYISSHLMSRAQ--SSTIR 223
Query: 218 NWGLSDRRPIILSLGAINFGPKP--FLFYNHWLLDKEFEKLVSDWWCSATTVGWSAF--- 272
SD + L + PK + F N L D+ F K V D W GW AF
Sbjct: 224 LAPFSDHNCVSLRMSIAPSLPKAAYWHFNNSLLEDEGFAKSVRDTWR-----GWRAFQDE 278
Query: 273 --VLQQKLKGLKNRIKEW-QGYASSVSRVRINTIEADLQVTMD---RLASEEPGIELRKN 326
L Q K +K Q Y SVS R IEA +D RL+ E L+
Sbjct: 279 FATLNQWWDVGKVHLKLLCQEYTKSVSGQRNAEIEALNGEVLDLEQRLSGSEDQA-LQCE 337
Query: 327 RLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVV-GGV 385
L E L +++ ++R++ L + DR + FF+++ K + ++K I+ L G
Sbjct: 338 YLERKEALRNMEQRQARGAFVRSRMQLLCDMDRGSRFFYALEKKKGNRKQITCLFAEDGT 397
Query: 386 SIEDPNMIKDAIFSHFQSFFTSENRLRPKIRCANL----SRLSQANKMSLEKQFSEGEIW 441
+EDP I+D S +Q+ F S + + P C L +S+ K LE + E+
Sbjct: 398 PLEDPEAIRDRARSFYQNLF-SPDPISPDA-CEELWDGLPVVSERRKERLETPITLDELS 455
Query: 442 EVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPK 501
+ L+ N++PG DG + FF+ FW + D + + G+L +SL+PK
Sbjct: 456 QALRLMPHNKSPGLDGLTIEFFQFFWDTLGPDFHRVLTEAFKKGELPLSCRRAVLSLLPK 515
Query: 502 HNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIAN 561
+ ++RP+SL+ + YK+++K ++ RL+ V +I +Q GR I D + +
Sbjct: 516 KGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKSVLAEVIHPDQSYTVPGRTIFDNVFLIR 575
Query: 562 EVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSI 621
+++ F + L LD K +D +D +L L+ +FG +++ +++T ++A +
Sbjct: 576 DLLHFARRTGLSLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQFVGYLKTMYASAECLV 635
Query: 622 LVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNH 681
+N S T +G+RQG PLS L++L + CLL + L V P + +
Sbjct: 636 KINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCLLRKRLTGL----VLKEPDMRVVL 691
Query: 682 LQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKST-IIGCNVDVRETQRVAGM 740
+ADD +L + D ++ + S ++N KS+ ++ ++ V
Sbjct: 692 SAYADDVILVAQ-DLVDLERAQECQEVYAAASSARINWSKSSGLLEGSLKVDFLPPAFRD 750
Query: 741 YGWSIGQLPILYLGAPLGGN--PRRASFWEPMLDKLRRKRRTWN--SKFISLSGRLTLMK 796
W I YLG L P +F E + + + + W +K +S+ GR ++
Sbjct: 751 ISWE--SKIIKYLGVYLSAEEYPVSQNFIE-LEECVLTRLGKWKGFAKVLSMRGRALVIN 807
Query: 797 ATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGGK 832
+ S + + + + ++++ L FL GK
Sbjct: 808 QLVASQIWYRLICLSPTQEFIAKIQRRLLDFLWIGK 843
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 173 bits (438), Expect = 3e-42
Identities = 197/864 (22%), Positives = 371/864 (42%), Gaps = 59/864 (6%)
Query: 5 SWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKL---DSSRVKTVEAFANAVKMDYVSV 61
S NV GL KR + + + LKP+I +QES L D R+K V+ +++ + +
Sbjct: 11 SINVNGLNCPLKRHRLADWIQKLKPDICCIQESHLTLKDKYRLK-VKGWSSIFQAN---- 65
Query: 62 DAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERY-- 119
AG I + + K+ + ++ + I+N+Y PN ++ ++
Sbjct: 66 GKQKKAGIAILFADAIGFKPTKIRKDKDGHFIFVKGNTQYDEISIINIYAPNHNAPQFIR 125
Query: 120 ----DFFNLLSEALSLNNCFCKIMGGDFN---AILNEGERIGIQMGVESSFSDFVADSNL 172
D NL+S I+ GDFN A+L+ + + + + + +L
Sbjct: 126 ETLTDMSNLISST--------SIVVGDFNTPLAVLDRSSKKKLSKEI-LDLNSTIQHLDL 176
Query: 173 IDLPL----QSDKFTWYSSRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPII 228
D+ ++T++SS A G +S+ID L + ++KF I + SD I
Sbjct: 177 TDIYRTFHPNKTEYTFFSS-AHGTYSKIDHILGHKSNLSKFKKIE--IIPCIFSDHHGIK 233
Query: 229 LSLGA---INFGPKPFLFYN-----HWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKG 280
+ L ++ K + N W++D E +K ++ + + L K
Sbjct: 234 VELNNNRNLHTHTKTWKLNNLMLKDTWVID-EIKKEITKFLEQNNNQDTNYQNLWDTAKA 292
Query: 281 -LKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYR 339
L+ + Q + R +N + L+ +++ P RK I L +
Sbjct: 293 VLRGKFIALQAFLKKTEREEVNNLMGHLK-QLEKEEHSNPKPSRRKEITKIRAELNEIEN 351
Query: 340 KEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIE-DPNMIKDAIF 398
K + K++ + + ++ ++++ ++ K LIS + G I DP+ I+ +
Sbjct: 352 KRIIQQINKSKSWFFEKINKIDKPLANLTRKKRVKSLISSIRNGNDEITTDPSEIQKILN 411
Query: 399 SHFQSFFTSE----NRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPG 454
+++ ++ + + + +L RLSQ L + S EI +++ ++PG
Sbjct: 412 EYYKKLYSHKYENLKEIDQYLEACHLPRLSQKEVEMLNRPISSSEIASTIQNLPKKKSPG 471
Query: 455 TDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNC-PSTVSDFRP 513
DGF F++ F + ++ F++ G L I+LIPK P+ ++RP
Sbjct: 472 PDGFTSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITLIPKPGKDPTRKENYRP 531
Query: 514 ISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNK-KQG 572
ISL+ K+++K+L +R+Q +I +Q F G Q I + V+ +NK K
Sbjct: 532 ISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINKLKNK 591
Query: 573 GGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFE 632
+L +D K +DNI F+ LK++ ++ I S T +I++NG F
Sbjct: 592 DHMILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFP 651
Query: 633 MEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFC 692
+ G RQG PLSP LFN+ + L+ + ++ G+ IG + FADD +++
Sbjct: 652 LRSGTRQGCPLSPLLFNIVMEVLAIAIR---EEKAIKGIHIG-SEEIKLSLFADDMIVYL 707
Query: 693 ENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILY 752
EN + L + + SG K+N HKS + + + V +++ + Y
Sbjct: 708 ENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNNNQAEKTVKDSIPFTVVPKKMKY 767
Query: 753 LGAPLGGNPR--RASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLC--SVPLFLMA 808
LG L + + +E + ++ W + S GR+ ++K ++ ++ F
Sbjct: 768 LGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVKMSILPKAIYNFNAI 827
Query: 809 IFKAPLKVVGEVEKILRSFLQGGK 832
KAPL ++EKI+ F+ K
Sbjct: 828 PIKAPLSYFKDLEKIILHFIWNQK 851
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 156 bits (394), Expect = 4e-37
Identities = 211/882 (23%), Positives = 370/882 (41%), Gaps = 93/882 (10%)
Query: 4 VSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSS-----RVKTVEAF--ANAVKM 56
+S N+ GL KR + + L P +QE+ L RVK + AN +K
Sbjct: 37 ISLNINGLNSPIKRHRLTDWLHKQDPTFCCLQETHLREKDRHYLRVKGWKTIFQANGLKK 96
Query: 57 DYVSVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDS 116
AG I + + + + K+ +L++ + L I+N+Y PN +
Sbjct: 97 Q---------AGVAILISDKIDFQPKVIKKDKEGHFILIKGKILQEELSILNIYAPNARA 147
Query: 117 ERYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESS--FSDFVADSNLID 174
+ L+ L I+ GDFN L+ +R Q + ++ + +L D
Sbjct: 148 ATFIRDTLVK--LKAYIAPHTIIVGDFNTPLSSKDRSWKQKLNRDTVKLTEVMKQMDLTD 205
Query: 175 LPL----QSDKFTWYSSRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPI-IL 229
+ ++ +T++S+ G +S+ID + + +N++ I + LSD + ++
Sbjct: 206 IYRTFYPKTKGYTFFSA-PHGTFSKIDHIIGHKTGLNRYKNI--EIVPCILSDHHGLRLI 262
Query: 230 SLGAINFGPKPFLF------YNHWLLDKEFEKLVSDWW----CSATTVG--WSAF--VLQ 275
IN G F + N L+ + +K + D+ ATT W L+
Sbjct: 263 FNNNINNGKPTFTWKLNNTLLNDTLVKEGIKKEIKDFLEFNENEATTYPNLWDTMKAFLR 322
Query: 276 QKLKGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLW 335
KL L AS R +T + L + L +E R R I++ L
Sbjct: 323 GKLIALS---------ASKKKRETAHT--SSLTTHLKALEKKEANSPKRSRRQEIIK-LR 370
Query: 336 QEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKI-----RKSKKLISQLVVGGVSIE-- 388
E + E R R R+ R + FF I+KI R +K ++++ + E
Sbjct: 371 GEINQVETR-RTIQRINQTR-----SWFFEKINKIDKPLARLTKGHRDKILINKIRNEKG 424
Query: 389 ----DPNMIKDAIFSHFQSFFTSE----NRLRPKIRCANLSRLSQANKMSLEKQFSEGEI 440
DP I++ I S ++ ++++ + + + + +L+Q L S EI
Sbjct: 425 DITTDPEEIQNTIRSFYKRLYSTKLENLDEMDKFLDRYQVPKLNQDQVDHLNSPISPKEI 484
Query: 441 WEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVV----KFFEDFHSTGKLVKGLNATFI 496
V+ S ++PG DGF+ F++ F KED++ K F G L I
Sbjct: 485 EAVINSLPTKKSPGPDGFSAEFYQTF----KEDLIPILHKLFHKIEVEGTLPNSFYEATI 540
Query: 497 SLIPK-HNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISD 555
+LIPK P+ + +FRPISL+ K+++K+LA+R+Q +I +Q F G Q
Sbjct: 541 TLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRIQEHIKAIIHPDQVGFIPGMQGWF 600
Query: 556 CILIANEVVDFLNK-KQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCV 614
I + V+ ++NK K ++ LD K +D I F+ +L+ ++N I+
Sbjct: 601 NIRKSINVIHYINKLKDKNHMIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAIY 660
Query: 615 STATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIG 674
S +I VNG ++ G RQG PLSP+LFN+ + L+ + Q G++IG
Sbjct: 661 SKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIVLEVLARAIRQ---QKEIKGIQIG 717
Query: 675 PGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRET 734
L ADD +++ + +N L N + SF G K+N +KS + +
Sbjct: 718 KEEVKISL-LADDMIVYISDPKNSTRELLNLINSFGEVVGYKINSNKSMAFLYTKNKQAE 776
Query: 735 QRVAGMYGWSIGQLPILYLGAPLGGNPRRA--SFWEPMLDKLRRKRRTWNSKFISLSGRL 792
+ + +SI I YLG L + ++ + +++ R W S GR+
Sbjct: 777 KEIRETTPFSIVTNNIKYLGVTLTKEVKDLYDKNFKSLKKEIKEDLRRWKDLPCSWIGRI 836
Query: 793 TLMKATLC--SVPLFLMAIFKAPLKVVGEVEKILRSFLQGGK 832
++K + ++ F K P + E+E + F+ K
Sbjct: 837 NIVKMAILPKAIYRFNAIPIKIPTQFFNELEGAICKFVWNNK 878
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 153 bits (386), Expect = 4e-36
Identities = 198/865 (22%), Positives = 370/865 (41%), Gaps = 65/865 (7%)
Query: 7 NVRGLGRAEKRRAVVEALSGLKPEIVLVQESKL---DSSRVKTVEAFANAVKMDYVSVDA 63
NV GL KR + + P + +QE+ L D+ R+K ++ + N Y +
Sbjct: 13 NVNGLNAPIKRHRLANWIKSQDPSVCCIQETHLTCRDTHRLK-IKGWRNI----YQANGK 67
Query: 64 VGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFN 123
AG I + + + ++ ++++ + L I+N+Y PN + R F
Sbjct: 68 QKKAGVAILVSDKTDFKPTKIKRDKEGHYIMVKGSIQQEELTILNIYAPNTGAPR--FIK 125
Query: 124 LLSEALSLNNCFCKIMGGDFN---AILNEGERIGIQMGVESSFSDFVADSNLID----LP 176
+ L + I+ GDFN + L+ R I ++ + + ++LID L
Sbjct: 126 QVLSDLQRDLDSHTIIMGDFNTPLSTLDRSTRQKINKDIQE-LNSALHQADLIDIYRTLH 184
Query: 177 LQSDKFTWYSSRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINF 236
+S ++T++S+ +S+ D L S+ ++K ++ + N LSD I L L
Sbjct: 185 PKSTEYTFFSA-PHHTYSKTDHILGSKTLLSKCKR-TEIITNC-LSDHSAIKLELRIKK- 240
Query: 237 GPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGY---AS 293
L NH K L++D+W +++ + +N+ +Q A
Sbjct: 241 -----LTQNHSTTWKLNNLLLNDYWVHNEMKA----EIKKFFETNENKDTTYQNLWDTAK 291
Query: 294 SVSRVRINTIEADLQVT----MDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKA 349
+V R + + A + +D L S+ +E ++ + QE K ++
Sbjct: 292 AVCRGKFIALNAHKRKQERSKIDTLISQLKELE-KQEQTNSKASRRQEIIKIRAELKEIE 350
Query: 350 RVRWLREGDRNTS-FFHSISKIR-------KSKKLISQLVV----GGVSIEDPNMIKDAI 397
+ L++ + + S FF I+KI K K+ +Q+ G DP I+ I
Sbjct: 351 TQKTLQKINESRSWFFEKINKIDRPLARLIKKKREKNQIDTIKNDRGDITTDPTEIQTTI 410
Query: 398 FSHFQSFFTSE----NRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAP 453
+++ + ++ + + L RL+Q SL + + EI ++ S ++P
Sbjct: 411 REYYKHLYANKLENLEEMDKFLDTYTLPRLNQEEVESLNRPITSSEIEAIINSLPNKKSP 470
Query: 454 GTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSD-FR 512
G +GF F++++ + ++K F+ G L I LIPK +T + FR
Sbjct: 471 GPEGFTAEFYQRYKEELVPFLLKLFQSIEKEGILPNSFYEASIILIPKPGRDTTKKENFR 530
Query: 513 PISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNK-KQ 571
PISL+ K+++K+LA+++Q LI +Q F Q I + ++ +N+ K
Sbjct: 531 PISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQGWFNIRKSINIIQHINRTKD 590
Query: 572 GGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFF 631
++ +D K +D I F+ L ++ ++ IR T +I++NG
Sbjct: 591 TNHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAP 650
Query: 632 EMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLF 691
++ G RQG PLSP L N+ + L+ + Q + G+++G L FADD +++
Sbjct: 651 PLKTGTRQGCPLSPLLPNIVLEVLARAIRQ---EKEIKGIQLGKEEVKLSL-FADDMIVY 706
Query: 692 CENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPIL 751
EN L + +F SG K+N+ KS + + ++ ++I I
Sbjct: 707 LENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMSELPFTIASKRIK 766
Query: 752 YLGAPLGGNPRR--ASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMK-ATLCSVPLFLMA 808
YLG L + + ++P+L++++ W + S GR+ ++K A L V A
Sbjct: 767 YLGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAILPKVIYRFNA 826
Query: 809 I-FKAPLKVVGEVEKILRSFLQGGK 832
I K P+ E+EK F+ K
Sbjct: 827 IPIKLPMTFFTELEKTTLKFIWNQK 851
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 89.4 bits (220), Expect = 6e-17
Identities = 75/276 (27%), Positives = 127/276 (45%), Gaps = 16/276 (5%)
Query: 452 APGTDGFNLNFFKQFWGIVKEDVVKFFEDFHST-GKLVKGLNATFISLIPKHNCPSTVSD 510
APG+DG + I + + + F H G + A +LIPK S+
Sbjct: 22 APGSDGLTVQ------AITRTRLPRNFVQLHLLRGHVPTPWTAMRTTLIPKDGDLENPSN 75
Query: 511 FRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKK 570
+RPI++ + +L+ ++LA RL+ L ++ +A G ++ +L + + ++
Sbjct: 76 WRPITIASALQRLLHRILAKRLEAAVELHPAQKGYARIDGTLVNSLLL--DTYISSRREQ 133
Query: 571 QGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVN-GSSTG 629
+ ++ LD K +D + + L+ + + N+I +S +T +I V GS T
Sbjct: 134 RKTYNVVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNYITGSLSDSTTTIRVGPGSQTR 193
Query: 630 FFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTL 689
+ +G++QGDPLSPFLFN ++ L C L G P LA FADD L
Sbjct: 194 KICIRRGVKQGDPLSPFLFNAVLDELLCSLQSTPGIGGTIGEEKIPVLA-----FADDLL 248
Query: 690 LFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTII 725
L +ND + T+ +F G+ +N KS I
Sbjct: 249 LLEDNDV-LLPTTLATVANFFRLRGMSLNAKKSVSI 283
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 85.1 bits (209), Expect = 1e-15
Identities = 85/355 (23%), Positives = 155/355 (42%), Gaps = 24/355 (6%)
Query: 485 GKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQ 544
G L + IPK DFRPIS+ + ++ +LA+RL + + Q
Sbjct: 389 GNLPHSIRLARTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLN--SSINWDPRQ 446
Query: 545 FAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGD 604
F +D I + V+ +K ++ LD +K +D++ ++D L+
Sbjct: 447 RGFLPTDGCADNATIVDLVLRHSHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYGAPK 506
Query: 605 RWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLD 664
++++++ S+ +G S+ F +G++QGDPLSP LFNL ++ L L +
Sbjct: 507 GFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTLPSEI- 565
Query: 665 DSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTI 724
G ++G + N FADD +LF E + +L + L FL GLK+N K
Sbjct: 566 -----GAKVGNAIT-NAAAFADDLVLFAET-RMGLQVLLDKTLDFLSIVGLKLNADKCFT 618
Query: 725 IGCNVDVRETQRVAGMYGWSIG--QLPIL-------YLGAPLGGNPR-RASFWEPMLDKL 774
+G ++ V + +G ++P L YLG R R + E + KL
Sbjct: 619 VGIKGQPKQKCTVLEAQSFYVGSSEIPSLKRTDEWKYLGINFTATGRVRCNPAEDIGPKL 678
Query: 775 RRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQ 829
+R + + RL ++ L +A+ + V+ + +K++R +++
Sbjct: 679 QRLTKA----PLKPQQRLFALRTVLIPQLYHKLALGSVAIGVLRKTDKLIRYYVR 729
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 68.6 bits (166), Expect = 1e-10
Identities = 70/274 (25%), Positives = 118/274 (42%), Gaps = 19/274 (6%)
Query: 425 QANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHST 484
Q +L + S EI EV ++C A G DG W + E + F
Sbjct: 309 QGEIKNLWRPISNDEIKEV-EACK-RTAAGPDGMTTTA----WNSIDECIKSLFNMIMYH 362
Query: 485 GKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQ 544
G+ + + LIPK + FRP+S+ A + ++LA+R+ L +
Sbjct: 363 GQCPRRYLDSRTVLIPKEPGTMDPACFRPLSIASVALRHFHRILANRIGEHGLLDTRQRA 422
Query: 545 FAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGD 604
F A G +++ + + ++ K G ++ LD K +D+++ + D L+
Sbjct: 423 FIVADG--VAENTSLLSAMIKEARMKIKGLYIAILDVKKAFDSVEHRSILDALRRKKLPL 480
Query: 605 RWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLD 664
N+I + + V + + +G+RQGDPLSP LFN CV + +L +L +
Sbjct: 481 EMRNYIMWVYRNSKTRLEVVKTKGRWIRPARGVRQGDPLSPLLFN-CV--MDAVLRRLPE 537
Query: 665 DSNF--SGVRIGPGLALNHLQFADDTLLFCENDE 696
++ F +IG L FADD +L E E
Sbjct: 538 NTGFLMGAEKIGA------LVFADDLVLLAETRE 565
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 64.3 bits (155), Expect = 2e-09
Identities = 53/208 (25%), Positives = 93/208 (44%), Gaps = 20/208 (9%)
Query: 499 IPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCIL 558
IPK PS+ S++RPISL +++ +++ SR++ L+S +Q F R ++
Sbjct: 673 IPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLSPHQHGFLNFRSCPSSLV 732
Query: 559 IANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTAT 618
+ + + K + +L DFAK +D + L L +W + + T
Sbjct: 733 RSISLYHSILKNEKSLDILFFDFAKAFDKVSHPILLKKLALFGLDKLTCSWFKEFLHLRT 792
Query: 619 LSILVNG-SSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL 677
S+ +N S+ + + G+ QG P LF L +N L L+D + P +
Sbjct: 793 FSVKINKFVSSNAYPISSGVPQGSVSGPLLFILFINDL------LID--------LEPNI 838
Query: 678 ALNHLQFADDTLLFCEND---ENQIDLL 702
++ FADD +F N +N ID +
Sbjct: 839 HVS--CFADDIKIFHHNPSTLQNSIDTI 864
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 64.3 bits (155), Expect = 2e-09
Identities = 45/162 (27%), Positives = 77/162 (46%), Gaps = 7/162 (4%)
Query: 576 LLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEK 635
++ LD K +D + + ++ D ++I + ++ A +I+V G +T +
Sbjct: 25 VVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNIVVGGRTTNKIYIRN 84
Query: 636 GLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCEND 695
G++QGDPLSP LFN+ ++ L LN G + P + L FADD LL + D
Sbjct: 85 GVKQGDPLSPVLFNIVLDELVTRLN-----DEQPGASMTPACKIASLAFADDLLLLEDRD 139
Query: 696 ENQIDLLCNTLLSFLFTSGLKVNLHK-STIIGCNVDVRETQR 736
+ + L T ++ T G+ +N K ++I V R R
Sbjct: 140 IDVPNSLATT-CAYFRTRGMTLNPEKCASISAATVSGRSVPR 180
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 61.6 bits (148), Expect = 1e-08
Identities = 31/71 (43%), Positives = 41/71 (57%), Gaps = 1/71 (1%)
Query: 619 LSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL- 677
L ++NG+ G +GLRQGDPLSP+LF LC LS L + + G+R+
Sbjct: 10 LLFIINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSP 69
Query: 678 ALNHLQFADDT 688
+NHL FADDT
Sbjct: 70 RINHLLFADDT 80
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 48.5 bits (114), Expect = 1e-04
Identities = 105/486 (21%), Positives = 193/486 (39%), Gaps = 51/486 (10%)
Query: 138 IMGGDFNAI----LNEGERIGIQMGVESS---FSDFVADSNLIDLPLQSDKFTWYSSRAG 190
IMG D NA+ ++GE +G E++ +++ + +I + S+ WY+
Sbjct: 94 IMGMDANAVSPLWFSKGENLGRGRLNEANGLLLEEWILEGRMIVINEPSE---WYTFSGP 150
Query: 191 GLWSRIDRWLVSEDAINKFDGISQSVE-NWGLSDRRPIIL--SLGAINFGPKPFLFYNHW 247
S ID LV+E A +F G SV+ WG+SD I + SL + P W
Sbjct: 151 NGSSDIDVTLVNE-AGGRF-GYEWSVQPEWGVSDHNLIRIRVSLDGLVADASPSPQSARW 208
Query: 248 LL-DKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEAD 306
D ++ + + D A G + + + + + EW Y ++ +R +T
Sbjct: 209 QTRDTDWGEYMGDVKAKADVFGLAQYE-NVSVDEKVDLLTEWI-YGANDWNMRRHTAVRT 266
Query: 307 LQVTMDRLASEEPGIELRKNRLTI----------LEGLWQEYR--KEEY-RWRQKARVRW 353
Q + E ELR+ R L Q +R K EY R +A++R
Sbjct: 267 FQNEWWSVELAEKRSELRRRRHAFQRIRNAGAASLADRLQAFRDCKIEYKRMLCEAKLRC 326
Query: 354 LRE---GDRNTSFFHSISKIRKSKKL---ISQLVVGGVSIEDPNMIKDAIFSHFQSFFTS 407
+E + N + + + K+ + ++ + + V GV + +A+ + F
Sbjct: 327 WQEFVASESNENPWGRVFKLCRGRRKPVDVCSVKVDGVYTDTWEGSVNAMMNVFFPASID 386
Query: 408 ENRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFW 467
+ + + RL +A L E+ + ++ C ++PG DG + W
Sbjct: 387 D--------ASEIDRL-KAIARPLPPDLEMDEVSDSVRRCKVRKSPGPDGIVGEMVRAVW 437
Query: 468 GIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPK--HNCPSTVSDFRPISLIGSAYKLIS 525
G + E + ++ + + ++ K S +RPI L+ + K++
Sbjct: 438 GAIPEYMFCLYKQCLLESYFPQKWKIASLVILLKLLDRIRSDPGSYRPICLLDNLGKVLE 497
Query: 526 KVLASRL-QVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKD 584
++ RL Q + + +S QFAF G+ D V+ K G L +DF
Sbjct: 498 GIMVKRLDQKLMDVEVSPYQFAFTYGKSTEDAWRCVQRHVECSEMKYVIG--LNIDFQGA 555
Query: 585 YDNIDW 590
+DN+ W
Sbjct: 556 FDNLGW 561
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 45.4 bits (106), Expect = 0.001
Identities = 61/268 (22%), Positives = 107/268 (39%), Gaps = 31/268 (11%)
Query: 439 EIWEV---LKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKL-VKGLNAT 494
E++EV + R+PG DG N K W + E + F G +
Sbjct: 435 EVFEVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLASLFSRCIRLGYFPAEWKCPR 494
Query: 495 FISLI---PKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGR 551
+SL+ K C S +R I L+ K++ ++ +R++ V P QF F QGR
Sbjct: 495 VVSLLKGPDKDKCEP--SSYRGICLLPVFGKVLEAIMVNRVREVLP-EGCRWQFGFRQGR 551
Query: 552 QISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIR 611
+ D V + G +DF +DN++W L ++ + + W
Sbjct: 552 CVEDAWRHVKSSVGASAAQYVLGTF--VDFKGAFDNVEWSAALSRLADLGCREMGL-W-- 606
Query: 612 TCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGV 671
+ +++ + S T + +G QG PF++++ L++ LL
Sbjct: 607 QSFFSGRRAVIRSSSGTVEVPVTRGCPQGSISGPFIWDI-------LMDVLLQ------- 652
Query: 672 RIGPGLALNHLQFADDTLLFCENDENQI 699
R+ P L+ +ADD LL E + +
Sbjct: 653 RLQPYCQLS--AYADDLLLLVEGNSRAV 678
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 43.5 bits (101), Expect = 0.004
Identities = 37/159 (23%), Positives = 66/159 (41%), Gaps = 23/159 (14%)
Query: 579 LDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLR 638
LDF+K +D + L D L + I W+ ++ + + V + + + G+
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 639 QGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQ 698
QG +SP LF + VN +S L + + QFADD L+ +NQ
Sbjct: 83 QGSVISPVLFGIFVNEISANL----------------PVGVYCKQFADDIKLYAAIPKNQ 126
Query: 699 ----IDLLCNTLLSFLFTSGLKVNLHKS---TIIGCNVD 730
+ L + + + + L +N K+ T+ CN +
Sbjct: 127 SQISLQLAIDVVTDWALNNKLLLNPSKTFHLTLGKCNTN 165
>EXOA_BACSU (P37454) Exodeoxyribonuclease (EC 3.1.11.2)
Length = 252
Score = 42.4 bits (98), Expect = 0.009
Identities = 65/255 (25%), Positives = 101/255 (39%), Gaps = 39/255 (15%)
Query: 4 VSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRV----KTVEAFAN-AVKMDY 58
+SWNV GL ++ + L +I+ +QE+K+ +V + + N AVK Y
Sbjct: 4 ISWNVNGLRAVMRKMDFLSYLKEEDADIICLQETKIQDGQVDLQPEDYHVYWNYAVKKGY 63
Query: 59 VSVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNV--DS 116
S + S P V+ G ++ ++ +F+M VY PN
Sbjct: 64 -------SGTAVFSKQEPLQVIYGIGVEEHDQEGRVI--TLEFENVFVMTVYTPNSRRGL 114
Query: 117 ERYDF-----FNLLSEALSLNNCFCKIMGGDFNAILNE---------GERIGIQMGVESS 162
ER D+ LLS L L+ I+ GD N E G +
Sbjct: 115 ERIDYRMQWEEALLSYILELDQKKPVILCGDLNVAHQEIDLKNPKANRNNAGFSDQEREA 174
Query: 163 FSDFV----ADSNLIDLPLQSDKFTWYSSRAG----GLWSRIDRWLVSEDAINKFDGISQ 214
F+ F+ DS P ++W+S RAG + RID ++VSE + + S
Sbjct: 175 FTRFLEAGFVDSFRHVYPDLEGAYSWWSYRAGARDRNIGWRIDYFVVSESLKEQIEDASI 234
Query: 215 SVENWGLSDRRPIIL 229
S + G SD P+ L
Sbjct: 235 SADVMG-SDHCPVEL 248
>AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein
Length = 789
Score = 39.7 bits (91), Expect = 0.058
Identities = 45/233 (19%), Positives = 94/233 (40%), Gaps = 37/233 (15%)
Query: 444 LKSCDGNRAPGT-----DGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISL 498
+KS GN + G+ DG N+++ + + ++ KF S
Sbjct: 237 IKSKKGNMSKGSNNITLDGINISYLNKLSKDINTNMFKF-------------------SP 277
Query: 499 IPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFA-FAQGRQISDCI 557
+ + P T FRP+S+ K++ + + L++ I N F+ ++ G + +
Sbjct: 278 VRRVEIPKTSGGFRPLSVGNPREKIVQESMRMMLEI-----IYNNSFSYYSHGFRPNLSC 332
Query: 558 LIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTA 617
L A ++ N Q + +K+D K +D I L ++L E +++ + +
Sbjct: 333 LTA--IIQCKNYMQYCNWFIKVDLNKCFDTIPHNMLINVLNERIKDKGFMDLLYKLLRAG 390
Query: 618 TLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSG 670
+ + + G+ QG +SP L N+ ++ L L ++ +G
Sbjct: 391 YVD-----KNNNYHNTTLGIPQGSVVSPILCNIFLDKLDKYLENKFENEFNTG 438
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 38.9 bits (89), Expect = 0.099
Identities = 47/218 (21%), Positives = 89/218 (40%), Gaps = 10/218 (4%)
Query: 439 EIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGL-NATFIS 497
E+ E++ +APG D + + +V F G K A+ I
Sbjct: 442 EVKELVSKLKPKKAPGEDLLDNRTIRLLPDQALLYLVLIFNSILRVGYFPKARPTASIIM 501
Query: 498 LIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPL--LISENQFAF-AQGRQIS 554
++ P V +RP SL+ S K++ +++ +R+ + I + QF F Q
Sbjct: 502 ILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRILTSEEVTRAIPKFQFGFRLQHGTPE 561
Query: 555 DCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDW--GFLFDILKEMNFGDRWINWIRT 612
+ N ++ L KK+ G LD + +D + W G L+ ++ + I++
Sbjct: 562 QLHRVVNFALEALEKKEYAGSCF-LDIQQAFDRV-WHPGLLYKAKSLLS--PQLFQLIKS 617
Query: 613 CVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNL 650
S+ +G + +E G+ QG L P L+++
Sbjct: 618 FWEGRKFSVTADGCRSSVKFIEAGVPQGSVLGPTLYSI 655
>RDPO_SCEOB (P19593) Probable reverse transcriptase (EC 2.7.7.49)
Length = 608
Score = 37.0 bits (84), Expect = 0.38
Identities = 62/322 (19%), Positives = 128/322 (39%), Gaps = 36/322 (11%)
Query: 408 ENRLRPKIRCA----NLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFF 463
EN+ + I CA N+ + + + + F + + S G+R+PG +
Sbjct: 36 ENK-QESIACAKREGNIVLVEKLAQEIVNSSFGRAVAVQTVASSKGSRSPGLSRESFKTN 94
Query: 464 KQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKL 523
K + ++ E S K + I IPK + RP+S+ +
Sbjct: 95 KNYVAMMAT-----LEQITSNPHKYKATPLSRI-YIPKRD-----GSARPLSIPSYTDRC 143
Query: 524 ISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGG-GFLLKLDFA 582
+ + ++ + + + + F R +S + V++ LN +++++D
Sbjct: 144 LQALYKLAIEPMAEEVADLSSYGFRPMRNVSWAV---GRVLNGLNNPLANYQYVVEIDIK 200
Query: 583 KDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDP 642
DNI+ F+ + + W W++ C S + ++TG + QG
Sbjct: 201 GCVDNINHQFISQVTPFIPKKILWA-WLK-CGYIERNSNTLQPTTTG-------VPQGGI 251
Query: 643 LSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLL 702
+SP + NL ++GL + + + S+ + ++ADD ++ +E + +
Sbjct: 252 ISPLIMNLTLDGLEFHIYKKIQKSS------SQSKGNTYCRYADDMVILTTTEETAL-IA 304
Query: 703 CNTLLSFLFTSGLKVNLHKSTI 724
+ FL GL+V L K+TI
Sbjct: 305 LPAVKEFLAVRGLEVKLAKTTI 326
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 36.6 bits (83), Expect = 0.49
Identities = 47/221 (21%), Positives = 91/221 (40%), Gaps = 16/221 (7%)
Query: 439 EIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISL 498
E+ ++ +APG D + + + + F G K + I +
Sbjct: 442 EVKNLIAKLPLKKAPGEDLLDNRTIRLLPDQALQFLALIFNSVLDVGYFPKAWKSASIIM 501
Query: 499 IPKHN-CPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPL--LISENQFAF----AQGR 551
I K P+ V +RP SL+ S K++ +++ +RL + I + QF F
Sbjct: 502 IHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTCKDVTKAIPKFQFGFRLQHGTPE 561
Query: 552 QISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDW--GFLFDILKEMNFGDRWINW 609
Q+ + A E ++ NK+ G LD + +D + W G L+ + F +
Sbjct: 562 QLHRVVNFALEAME--NKEYAVGAF--LDIQQAFDRV-WHPGLLYK--AKRLFPPQLYLV 614
Query: 610 IRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNL 650
+++ + T + V+G + + G+ QG L P L+++
Sbjct: 615 VKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLYSV 655
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 36.6 bits (83), Expect = 0.49
Identities = 56/250 (22%), Positives = 95/250 (37%), Gaps = 22/250 (8%)
Query: 488 VKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAF 547
+KG F I P FRP+S+ + +K+LA R V+ E Q A+
Sbjct: 210 IKGSRTVFTPKIEGGPDPGV---FRPLSICSVILREFNKILARRF--VSCYTYDERQTAY 264
Query: 548 AQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWI 607
+ + + ++ + + + LD K ++++ L D + E +
Sbjct: 265 LPIDGVCINVSMLTAIIAEAKRLRKELHIAILDLVKAFNSVYHSALIDAITEAGCPPGVV 324
Query: 608 NWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCV-NGLSCLLNQLLDDS 666
++I + + G + G+ QGDPLS LF L L L N+ D
Sbjct: 325 DYIADMYNNVITEMQFEGKCE-LASILAGVYQGDPLSGPLFTLAYEKALRALNNEGRFD- 382
Query: 667 NFSGVRIGPGLALNHLQFADDTLLFCEND---ENQIDLLCNTLLSFLFTSGLKVNLHKST 723
+ VR+ N ++DD LL ++ +D TL GL++N KS
Sbjct: 383 -IADVRV------NASAYSDDGLLLAMTVIGLQHNLDKFGETLAKI----GLRINSRKSK 431
Query: 724 IIGCNVDVRE 733
+ RE
Sbjct: 432 TVSLVPSGRE 441
>RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa protein
(ORF2)
Length = 1182
Score = 35.8 bits (81), Expect = 0.84
Identities = 51/224 (22%), Positives = 85/224 (37%), Gaps = 19/224 (8%)
Query: 452 APGTDGFNLNFFKQFW--GIVKEDVVKFFEDFHSTG---KLVKGLNATFISLIPKHNCPS 506
APG DG+ +K ++ +D + ++ + L AT ++++ K N
Sbjct: 527 APGLDGWTRELLYPLTLDPALKMEIAAVVKDIINADVSMEVGRRLQATSLTVLRKPN--- 583
Query: 507 TVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDF 566
+RPI KL S + SR+ S QF G I + I + DF
Sbjct: 584 --GKYRPIGAESVWAKLASHIAISRVMKTAEKKFSGIQFGV--GGHIEEAIAKIRK--DF 637
Query: 567 LNKKQGGGFLLKLDFAKDYDNIDWGFLFD-ILKEMNFGDRWINWIRTCVSTATLSILVNG 625
K G L LD Y+ I + + + + + W +T + NG
Sbjct: 638 ATK----GSLAMLDGRNAYNAISRRAILEAVYGDSTWSPLWRLVSLLLGTTGEVGFYENG 693
Query: 626 SSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFS 669
+E +G+RQG L P LF++ L Q ++ F+
Sbjct: 694 KLCHTWESTRGVRQGMVLGPLLFSIGTLATLRRLQQTFPEAQFT 737
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.147 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 154,860,597
Number of Sequences: 164201
Number of extensions: 6400050
Number of successful extensions: 21030
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 20978
Number of HSP's gapped (non-prelim): 33
length of query: 1389
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1266
effective length of database: 39,777,331
effective search space: 50358101046
effective search space used: 50358101046
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0110a.13


