

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.10
(1408 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
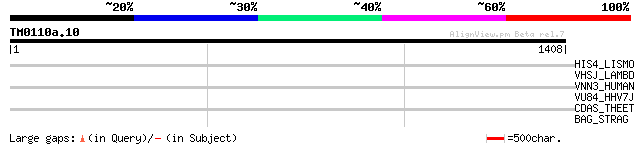
Score E
Sequences producing significant alignments: (bits) Value
HIS4_LISMO (Q8Y9G4) 1-(5-phosphoribosyl)-5-[(5-phosphoribosylami... 35 1.4
VHSJ_LAMBD (P03749) Host specificity protein J 33 4.2
VNN3_HUMAN (Q9NY84) Vascular non-inflammatory molecule 3 precurs... 33 7.2
VU84_HHV7J (P52534) Protein U84 32 9.4
CDAS_THEET (P29964) Cyclomaltodextrinase (EC 3.2.1.54) (CDase) (... 32 9.4
BAG_STRAG (P27951) IgA FC receptor precursor (Beta antigen) (B a... 32 9.4
>HIS4_LISMO (Q8Y9G4)
1-(5-phosphoribosyl)-5-[(5-
phosphoribosylamino)methylideneamino]
imidazole-4-carboxamide isomerase (EC 5.3.1.16)
(Phosphoribosylformimino-5-aminoimidazole carboxamide
ribotide isomerase)
Length = 240
Score = 35.0 bits (79), Expect = 1.4
Identities = 25/98 (25%), Positives = 45/98 (45%), Gaps = 3/98 (3%)
Query: 570 MKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV-VPYNPKLL-- 626
+KNG+C + F ++ T + D + T +T VDLD P N +++
Sbjct: 9 LKNGQCVRLFQGDFSKKTVVNEDPIAQAKAFATDGATYLHIVDLDGALEGRPINLEVIQK 68
Query: 627 MKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSS 664
MK A I ++ S+ + Y+ G+DRV + ++
Sbjct: 69 MKITAKIPVQVGGGIRSMAQVDYYLESGIDRVIIGSAA 106
>VHSJ_LAMBD (P03749) Host specificity protein J
Length = 1132
Score = 33.5 bits (75), Expect = 4.2
Identities = 30/90 (33%), Positives = 41/90 (45%), Gaps = 9/90 (10%)
Query: 765 PELFVYDP--KSKSWHPRKQGESIGRMSFVPPGTGE-LYYLRMLLNVQRGCTKYEDLRSV 821
P L VYDP + + W KQ I ++ G LY++ +N++ G Y +RSV
Sbjct: 733 PHLAVYDPTVQFEFWFSEKQIADIRQVETSTRYLGTALYWIAASINIKPGHDYYFYIRSV 792
Query: 822 NNNVHDTFRGACEALGLLKDDREFIDGILD 851
N F EA+G DD E G LD
Sbjct: 793 NTVGKSAF---VEAVGRASDDAE---GYLD 816
>VNN3_HUMAN (Q9NY84) Vascular non-inflammatory molecule 3 precursor
(Vanin 3)
Length = 501
Score = 32.7 bits (73), Expect = 7.2
Identities = 29/108 (26%), Positives = 44/108 (39%), Gaps = 20/108 (18%)
Query: 25 PYLADCPELLSNLL------RNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGP 78
PYL D P+ N + R +T + L + NS++ +IG K C +D + P
Sbjct: 95 PYLEDIPDPGVNWIPCRDPWRFGNTPVQQRLSCLAKDNSIYVVANIGDKKPCNASDSQCP 154
Query: 79 P--------QFVISGQ-----NYHRIGSLVP-TEGDNPKFAQLYIYDT 112
P V Q YH+ P + D PK ++L +DT
Sbjct: 155 PDGRYQYNTDVVFDSQGKLLARYHKYNLFAPEIQFDFPKDSELVTFDT 202
>VU84_HHV7J (P52534) Protein U84
Length = 310
Score = 32.3 bits (72), Expect = 9.4
Identities = 18/74 (24%), Positives = 34/74 (45%), Gaps = 9/74 (12%)
Query: 624 KLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSS-----GRTTQDKPVEDDEI 678
++L+K + N Y K + ++ +FK + + + + S KP+ EI
Sbjct: 64 EMLLKADKYTNAIY--KHSKLRKIFKAVKENLSTTNFATKSKGIVHSNFNDSKPIHSPEI 121
Query: 679 KQY--FDCRYLSPC 690
K Y F+C ++ PC
Sbjct: 122 KNYTDFNCDFMHPC 135
>CDAS_THEET (P29964) Cyclomaltodextrinase (EC 3.2.1.54) (CDase)
(Cyclomaltodextrin hydrolase, decycling)
Length = 574
Score = 32.3 bits (72), Expect = 9.4
Identities = 19/81 (23%), Positives = 39/81 (47%), Gaps = 4/81 (4%)
Query: 674 EDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVL 733
E++E+K C A+ R+F + +K + + + +++V+ E E ED+L
Sbjct: 496 ENEELKYGSFCTLY----AIGRVFAFKREYKGKSIIVVLNNSSKQEVIFLNEVEGKEDIL 551
Query: 734 RRNIVKNSMFLAWMDANCKYV 754
+ +K S L ++ N Y+
Sbjct: 552 KMKELKRSGNLLYLQPNSAYI 572
>BAG_STRAG (P27951) IgA FC receptor precursor (Beta antigen) (B
antigen)
Length = 1164
Score = 32.3 bits (72), Expect = 9.4
Identities = 19/72 (26%), Positives = 35/72 (48%), Gaps = 3/72 (4%)
Query: 627 MKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDK--PVEDDEIKQYFDC 684
M YQ H +E + ++Y+ KY + + RT +D +E+D +K YF+
Sbjct: 647 MNYQLHAQMEMLTRK-VVQYMNKYPDNAEIKKIFESDMKRTKEDNYGSLENDALKGYFEK 705
Query: 685 RYLSPCEAVWRI 696
+L+P + +I
Sbjct: 706 YFLTPFNKIKQI 717
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.353 0.158 0.566
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 144,190,271
Number of Sequences: 164201
Number of extensions: 5619933
Number of successful extensions: 25099
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 25099
Number of HSP's gapped (non-prelim): 6
length of query: 1408
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1285
effective length of database: 39,777,331
effective search space: 51113870335
effective search space used: 51113870335
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0110a.10


