

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
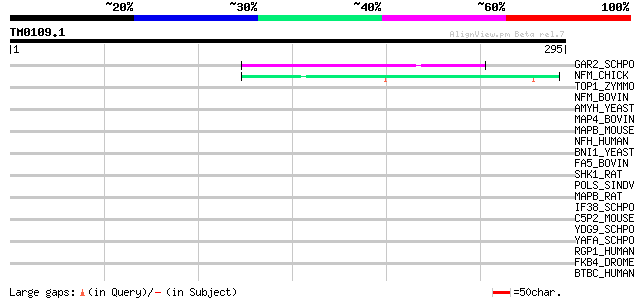
Query= TM0109.1
(295 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
GAR2_SCHPO (P41891) Protein gar2 46 1e-04
NFM_CHICK (P16053) Neurofilament triplet M protein (160 kDa neur... 44 5e-04
TOP1_ZYMMO (Q9X3X7) DNA topoisomerase I (EC 5.99.1.2) (Omega-pro... 41 0.003
NFM_BOVIN (O77788) Neurofilament triplet M protein (160 kDa neur... 41 0.003
AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3) (G... 41 0.003
MAP4_BOVIN (P36225) Microtubule-associated protein 4 (MAP 4) (Mi... 40 0.009
MAPB_MOUSE (P14873) Microtubule-associated protein 1B (MAP 1B) (... 39 0.015
NFH_HUMAN (P12036) Neurofilament triplet H protein (200 kDa neur... 39 0.020
BNI1_YEAST (P41832) BNI1 protein (Synthetic lethal 39) 39 0.020
FA5_BOVIN (Q28107) Coagulation factor V precursor (Activated pro... 38 0.026
SHK1_RAT (Q9WV48) SH3 and multiple ankyrin repeat domains protei... 38 0.034
POLS_SINDV (P03316) Structural polyprotein (P130) [Contains: Coa... 38 0.034
MAPB_RAT (P15205) Microtubule-associated protein 1B (MAP 1B) (Ne... 38 0.034
IF38_SCHPO (O14164) Probable eukaryotic translation initiation f... 38 0.034
C5P2_MOUSE (Q8K389) CDK5 regulatory subunit associated protein 2... 38 0.034
YDG9_SCHPO (Q10496) Hypothetical protein C26F1.09 in chromosome I 37 0.045
YAFA_SCHPO (Q09863) Hypothetical protein C29E6.10c in chromosome I 37 0.045
RGP1_HUMAN (P46060) Ran GTPase-activating protein 1 37 0.045
FKB4_DROME (P54397) 39 kDa FK506-binding nuclear protein (EC 5.2... 37 0.045
BTBC_HUMAN (Q8IY92) BTB/POZ domain containing protein 12 37 0.045
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 46.2 bits (108), Expect = 1e-04
Identities = 33/130 (25%), Positives = 58/130 (44%), Gaps = 2/130 (1%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
EEE +K E+ + S E ++E+E+ES + + EE E TE+ E +E +
Sbjct: 148 EEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEGSSESSSD 207
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
E + +++ D+ + + + +RK A + R A+I + NE+ T
Sbjct: 208 SESSSDSSSESGDSDSSSDSESESSSEDEKKRK--AEPASEERPAKITKPSQDSNETCTV 265
Query: 244 DVASLSLGDD 253
V LS D
Sbjct: 266 FVGRLSWNVD 275
Score = 29.6 bits (65), Expect = 9.3
Identities = 32/176 (18%), Positives = 70/176 (39%), Gaps = 17/176 (9%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKD-----------VPEETENESLTED 172
E ++S E+ +SE+E+ S E ++ +E+ES + + V E + ES +E
Sbjct: 82 ESSSESESESSSSESESSS-SESESSSSESESSSSESSSSESEEEVIVKTEEKKESSSES 140
Query: 173 VNESLTEDVGEM-----EKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRS 227
+ S +E+ E EK ES + + S + S + + ++V +
Sbjct: 141 SSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEG 200
Query: 228 ARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRS 283
+ + + E + +++ D+ + + + +RK P + S
Sbjct: 201 SSESSSDSESSSDSSSESGDSDSSSDSESESSSEDEKKRKAEPASEERPAKITKPS 256
>NFM_CHICK (P16053) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M)
Length = 857
Score = 43.9 bits (102), Expect = 5e-04
Identities = 41/184 (22%), Positives = 70/184 (37%), Gaps = 17/184 (9%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
+EE K+ +E V E ++ E A E E E ++ EE +S + S E++ E
Sbjct: 485 QEEEKAEEEAVEEEAVSEKAAEQAAEEEEKEE--EEAEEEEAAKSDAAEEGGSKKEEIEE 542
Query: 184 MEKNESLTTDVASLS-----LGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKN 238
E+ E + A G P K+ K P ++ ++ E
Sbjct: 543 KEEGEEAEEEEAEAKGKAEEAGAKVEKVKSPPAKSPPKSPPKSPVTEQAKAVQKAAAEVG 602
Query: 239 ESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQ----------PAVRRSARIKT 288
+ + A+ + A+ P +P T + + +PA P+ P RS T
Sbjct: 603 KDQKAEKAAEKAAKEEKAASPEKPATPKVTSPEKPATPEKPPTPEKAITPEKVRSPEKPT 662
Query: 289 TPKK 292
TP+K
Sbjct: 663 TPEK 666
>TOP1_ZYMMO (Q9X3X7) DNA topoisomerase I (EC 5.99.1.2) (Omega-protein)
(Relaxing enzyme) (Untwisting enzyme) (Swivelase)
Length = 1217
Score = 41.2 bits (95), Expect = 0.003
Identities = 39/133 (29%), Positives = 57/133 (42%), Gaps = 19/133 (14%)
Query: 160 VPEETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVA 219
+P++T ESLT + +L E +M K + A A P KT K A
Sbjct: 1091 LPKDTTPESLTFEEALALLEARAKMPKKKKTKK----------AAAKKPAAKKTTTKKAA 1140
Query: 220 PQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPA 279
P+ A ++A K S TTD A+ GD A P + K A +PA+ + A
Sbjct: 1141 PKKATTKTATPK--------SATTDNATSEDGDATPAKSPAKKAVAIKTTAKKPAS-KSA 1191
Query: 280 VRRSARIKTTPKK 292
+++ KTT K
Sbjct: 1192 TKKAPSSKTTAAK 1204
Score = 30.8 bits (68), Expect = 4.2
Identities = 23/85 (27%), Positives = 36/85 (42%), Gaps = 8/85 (9%)
Query: 216 KLVAPQPAVRRSARIKTGEM-------EKNESLTTDVASLSLG-DDALASPPPQPKTRRK 267
K+ P + ++ G E N SL D SL ++ALA + K +K
Sbjct: 1059 KVFGDSPVTEKPVQLMNGRYGPYVTDGETNASLPKDTTPESLTFEEALALLEARAKMPKK 1118
Query: 268 LAAPQPAAPQPAVRRSARIKTTPKK 292
+ AA +PA +++ K PKK
Sbjct: 1119 KKTKKAAAKKPAAKKTTTKKAAPKK 1143
>NFM_BOVIN (O77788) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M) (Fragment)
Length = 810
Score = 41.2 bits (95), Expect = 0.003
Identities = 38/168 (22%), Positives = 61/168 (35%), Gaps = 3/168 (1%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
EEE +E E E +E E S ++ +E E E+ E E + E
Sbjct: 414 EEEEGQEEEEEEEEAAKSDQAEEGGSEKEGSSEKEEGEQEEEGETEAEGEGEEAAAEAKE 473
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
+K E +VA + A + +P+ + VA P + S K+ E + E+ +
Sbjct: 474 EKKMEEKAEEVAPKE--ELAAEAKVEKPEKAKSPVAKSPTTK-SPTAKSPEAKSPEAKSP 530
Query: 244 DVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTPK 291
S + SP + + A P A P + A PK
Sbjct: 531 TAKSPTAKSPVAKSPTAKSPEAKSPEAKSPTAKSPTAKSPAAKSPAPK 578
Score = 35.0 bits (79), Expect = 0.22
Identities = 46/188 (24%), Positives = 71/188 (37%), Gaps = 27/188 (14%)
Query: 118 ECCVTAEEENKSLKENVASETENKSLMEH--VAA-------------ETENESLTKDVPE 162
E V+ +EE +KE A E E K E VAA E E E ++ E
Sbjct: 364 ELAVSVKEE---VKEEEAEEKEEKEEAEEEVVAAKKSPVKATAPELKEEEGEKEEEEGQE 420
Query: 163 ETENE---SLTEDVNESLTEDVGEMEKNESLTTDVASLSL---GDDALASPPPQPKTRRK 216
E E E + ++ E +E G EK E + G++A A + K K
Sbjct: 421 EEEEEEEAAKSDQAEEGGSEKEGSSEKEEGEQEEEGETEAEGEGEEAAAEAKEEKKMEEK 480
Query: 217 L--VAPQPAVRRSARIKTGEMEKNE-SLTTDVASLSLGDDALASPPPQPKTRRKLAAPQP 273
VAP+ + A+++ E K+ + + S + SP + T + A P
Sbjct: 481 AEEVAPKEELAAEAKVEKPEKAKSPVAKSPTTKSPTAKSPEAKSPEAKSPTAKSPTAKSP 540
Query: 274 AAPQPAVR 281
A P +
Sbjct: 541 VAKSPTAK 548
>AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3)
(Glucan 1,4-alpha-glucosidase) (1,4-alpha-D-glucan
glucohydrolase)
Length = 1367
Score = 41.2 bits (95), Expect = 0.003
Identities = 38/159 (23%), Positives = 54/159 (33%), Gaps = 9/159 (5%)
Query: 135 ASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDV 194
+S T S ++ TE+ S P + ES + V S TE S +T
Sbjct: 462 SSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTE 521
Query: 195 ASLSLGDDALASPPPQPKTRRKLVAP---QPAVRRSARIKTGEMEKNESLTTDVASLSLG 251
+S + +P P T AP SA + T ES +T V S +
Sbjct: 522 SS------SAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSTPVTSSTTE 575
Query: 252 DDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTP 290
+ P P T +AP P S+ TP
Sbjct: 576 SSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAPTP 614
Score = 40.8 bits (94), Expect = 0.004
Identities = 41/175 (23%), Positives = 60/175 (33%), Gaps = 13/175 (7%)
Query: 122 TAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDV 181
T E + + +S TE+ S + + ES + VP T + S TE + +T
Sbjct: 630 TTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVP--TPSSSTTESSSAPVTSST 687
Query: 182 GEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRS------ARIKTGEM 235
ES + V S + + P P T AP P S A + T
Sbjct: 688 -----TESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSS 742
Query: 236 EKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTP 290
ES + V S + + P P T +AP P S+ TP
Sbjct: 743 STTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTP 797
Score = 40.4 bits (93), Expect = 0.005
Identities = 41/188 (21%), Positives = 62/188 (32%), Gaps = 13/188 (6%)
Query: 115 IPCECCVTAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVN 174
+P T E + + + +E+ + + ++ TE+ S P + ES + V
Sbjct: 485 VPTPSSSTTESSSAPVTSST-TESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVT 543
Query: 175 ESLTEDVG------EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRS- 227
S TE ES +T V S + + P P T AP P S
Sbjct: 544 SSTTESSSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSST 603
Query: 228 -----ARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRR 282
A T ES + V S + + P P T +AP P
Sbjct: 604 TESSSAPAPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTES 663
Query: 283 SARIKTTP 290
S+ TP
Sbjct: 664 SSAPVPTP 671
Score = 33.9 bits (76), Expect = 0.49
Identities = 37/170 (21%), Positives = 60/170 (34%), Gaps = 7/170 (4%)
Query: 121 VTAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTED 180
VT+ S +S TE+ S + + ES + V T ES + V S TE
Sbjct: 371 VTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSST-TESSSAPVTSSTTES 429
Query: 181 VGEMEKNESLTTDVASL-SLGDDALASPPPQPKTRRKLVAPQPAV-----RRSARIKTGE 234
+ + + A + S ++ ++P P P + + P SA + T
Sbjct: 430 SSAPVTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPS 489
Query: 235 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSA 284
ES + V S + + P P T +AP P S+
Sbjct: 490 SSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSS 539
Score = 32.0 bits (71), Expect = 1.9
Identities = 39/162 (24%), Positives = 55/162 (33%), Gaps = 13/162 (8%)
Query: 135 ASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDV 194
+S TE+ S + + ES + VP T + S TE + S ES + V
Sbjct: 571 SSTTESSSAPVPTPSSSTTESSSAPVP--TPSSSTTE--SSSAPAPTPSSSTTESSSAPV 626
Query: 195 ASLSLGDDALASPPPQPKTRRKLVAPQP------AVRRSARIKTGEMEKNESLTTDVASL 248
S + + P P T AP P SA + T ES + V S
Sbjct: 627 TSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSS 686
Query: 249 SLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTP 290
+ +S P T +AP P S+ TP
Sbjct: 687 TTES---SSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTP 725
Score = 30.8 bits (68), Expect = 4.2
Identities = 34/155 (21%), Positives = 55/155 (34%), Gaps = 12/155 (7%)
Query: 121 VTAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTED 180
VT+ S +S TE+ S + + ES + VP T + S TE + S
Sbjct: 683 VTSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVP--TPSSSTTE--SSSAPVP 738
Query: 181 VGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNES 240
ES + V S + + P P T AP P S + S
Sbjct: 739 TPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPS 798
Query: 241 LTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAA 275
+T +S+ +P P P + + + P++
Sbjct: 799 SSTTESSV--------APVPTPSSSSNITSSAPSS 825
>MAP4_BOVIN (P36225) Microtubule-associated protein 4 (MAP 4)
(Microtubule-associated protein-U) (MAP-U)
Length = 1072
Score = 39.7 bits (91), Expect = 0.009
Identities = 51/196 (26%), Positives = 79/196 (40%), Gaps = 20/196 (10%)
Query: 109 VPVFEIIPCECCVTAEEEN--KSLKENVASETENKSLMEHVAAETENESLTKD---VPEE 163
VPV ++ + E++ KSL++ S + E V A + SL D V EE
Sbjct: 536 VPVKDMETAQTQEATSEDSQLKSLQDEGQSAVPLMTSPEAVVAMGQKHSLPTDEDSVLEE 595
Query: 164 TENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDALASP----PPQPKTRRK-LV 218
E + + +E +E G + E T S G+D A P PP P+ + K L
Sbjct: 596 LEQKKPSSQTSELPSETSGVAKPEEGPPTGSVS---GNDITAPPNKELPPSPEKKTKPLA 652
Query: 219 APQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQP 278
QPA +++ KT + SL A +LG ++ P + A P P
Sbjct: 653 TTQPAKTSTSKAKT----QPTSLPKQTAPTTLGG---SNKKPMSLASGSVPAAPPKRPAA 705
Query: 279 AVRRSARIKTTPKKFK 294
A R + + + K K
Sbjct: 706 ATSRPSTLPSKDTKPK 721
>MAPB_MOUSE (P14873) Microtubule-associated protein 1B (MAP 1B)
(MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]
Length = 2464
Score = 38.9 bits (89), Expect = 0.015
Identities = 38/172 (22%), Positives = 71/172 (41%), Gaps = 17/172 (9%)
Query: 91 LKNEIKTGFAKQHNFKGCVPVFEIIPCECCVTAEEENKSLKENVASETENKSLMEHVAAE 150
+K EIKT ++ K P E++ E +++ K KE V E + + E
Sbjct: 638 VKKEIKTKLEEKKEEK---PKKEVVKKEDKTPLKKDEKPRKEEVKKEIKKEIKKE----- 689
Query: 151 TENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQ 210
E + L K+V +ET + ++V + ++V + EK S + S +
Sbjct: 690 -ERKELKKEVKKETPLKDAKKEVKKEEKKEVKKEEKEPKKEIKKISKDIKKSTPQSDTKK 748
Query: 211 PKTRRKLVAPQPAVRRSARIKTGEME--------KNESLTTDVASLSLGDDA 254
P + VA + + + G+++ K E TT+ A+ ++G A
Sbjct: 749 PSALKPKVAKKEESTKKEPLAAGKLKDKGKVKVIKKEGKTTEAAATAVGTAA 800
>NFH_HUMAN (P12036) Neurofilament triplet H protein (200 kDa
neurofilament protein) (Neurofilament heavy polypeptide)
(NF-H)
Length = 1026
Score = 38.5 bits (88), Expect = 0.020
Identities = 35/136 (25%), Positives = 55/136 (39%), Gaps = 7/136 (5%)
Query: 145 EHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDAL 204
E + ++E T V E+TE +TE+V E ++ E E E + G++
Sbjct: 442 EKIKVVEKSEKETVIVEEQTEETQVTEEVTEEEEKEAKEEEGKEEEGGEEEEAEGGEEET 501
Query: 205 ASPPPQPKTRRKLVAPQPA---VRRSARIKTGEMEKNESLTTDVASLSLGDDAL---ASP 258
SPP + + A P + A K+ E E+ +S +V S A
Sbjct: 502 KSPPAEEAASPEKEAKSPVKEEAKSPAEAKSPEKEEAKS-PAEVKSPEKAKSPAKEEAKS 560
Query: 259 PPQPKTRRKLAAPQPA 274
PP+ K+ K A PA
Sbjct: 561 PPEAKSPEKEEAKSPA 576
>BNI1_YEAST (P41832) BNI1 protein (Synthetic lethal 39)
Length = 1953
Score = 38.5 bits (88), Expect = 0.020
Identities = 36/136 (26%), Positives = 52/136 (37%), Gaps = 13/136 (9%)
Query: 131 KENVASETENKSLMEHVAAETEN--ESLTKDVPEETENESLTEDVN--ESLTEDVGEMEK 186
++N + KSL E+ A + + KD+ + EN VN E T D + K
Sbjct: 1159 EQNEDEGLDKKSLPENSTASAASAFDKAEKDMRQHVENGKQGRVVNHEEDKTADFSAVSK 1218
Query: 187 NESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVA 246
+ T LS L+S PP P P P A++ +EK + D
Sbjct: 1219 LNN-TDGAEDLSTQSSVLSSQPPPP--------PPPPPPVPAKLFGESLEKEKKSEDDTV 1269
Query: 247 SLSLGDDALASPPPQP 262
D+ A PPP P
Sbjct: 1270 KQETTGDSPAPPPPPP 1285
>FA5_BOVIN (Q28107) Coagulation factor V precursor (Activated protein
C cofactor)
Length = 2211
Score = 38.1 bits (87), Expect = 0.026
Identities = 38/153 (24%), Positives = 69/153 (44%), Gaps = 21/153 (13%)
Query: 129 SLKENVASETENKSLMEH-VAAETENESLTKDVPE-----ETENESLTEDVN-------- 174
SL +++ E+ + L + ++ + ESL+ D+ + + ESL+ D+
Sbjct: 1224 SLSPDLSQESLSPDLGQTALSPDPSQESLSPDLGQTALSPDPSQESLSPDLGQTALSPDP 1283
Query: 175 --ESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQ----PKTRRKLVAPQPAVRRSA 228
ESL+ D+G+ + L+ + S LG AL+ P Q P + ++P P+ + S
Sbjct: 1284 GQESLSPDLGQTSLSPDLSQESLSPDLGQTALSPDPSQESLSPDLGQTALSPDPS-QESL 1342
Query: 229 RIKTGEMEKNESLTTDVASLSLGDDALASPPPQ 261
G+ + L + S LG AL+ P Q
Sbjct: 1343 SPDLGQTSLSPDLGQESLSPDLGQTALSPDPSQ 1375
Score = 32.3 bits (72), Expect = 1.4
Identities = 29/109 (26%), Positives = 49/109 (44%), Gaps = 11/109 (10%)
Query: 175 ESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQ----PKTRRKLVAPQPAVRRSARI 230
ESL+ D+G+ + L+ + S LG AL+ P Q P + ++P P+ + S
Sbjct: 1214 ESLSPDLGQTSLSPDLSQESLSPDLGQTALSPDPSQESLSPDLGQTALSPDPS-QESLSP 1272
Query: 231 KTGEMEKN-----ESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPA 274
G+ + ESL+ D+ SL D L+ P + +P P+
Sbjct: 1273 DLGQTALSPDPGQESLSPDLGQTSLSPD-LSQESLSPDLGQTALSPDPS 1320
>SHK1_RAT (Q9WV48) SH3 and multiple ankyrin repeat domains protein 1
(Shank1) (GKAP/SAPAP interacting protein) (SPANK-1)
(Synamon) (Somatostatin receptor interacting protein)
(SSTR interacting protein) (SSTRIP)
Length = 2167
Score = 37.7 bits (86), Expect = 0.034
Identities = 30/95 (31%), Positives = 39/95 (40%), Gaps = 2/95 (2%)
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
+E + S + L+ G PP P T AP PAV + S +
Sbjct: 1577 LEFSNSFEKPESPLTPGPPHPLPDPPSPATPLP-AAPPPAVAAAPPTLDSTASSLTSYDS 1635
Query: 244 DVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQP 278
+VA+L+ G A PP P AAP P APQP
Sbjct: 1636 EVATLTQGAPAAPGDPPAPGPPAP-AAPAPPAPQP 1669
>POLS_SINDV (P03316) Structural polyprotein (P130) [Contains: Coat
protein C (EC 3.4.21.-) (Capsid protein C); Spike
glycoprotein E3; Spike glycoprotein E2; 6 kDa peptide;
Spike glycoprotein E1]
Length = 1245
Score = 37.7 bits (86), Expect = 0.034
Identities = 29/100 (29%), Positives = 42/100 (42%), Gaps = 25/100 (25%)
Query: 214 RRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPP-PQPKTRRKLAAP- 271
RR+ AP PA + G + + LTT V++L +G PP P+P R+K AP
Sbjct: 25 RRRQAAPMPA-------RNGLASQIQQLTTAVSALVIGQATRPQPPRPRPPPRQKKQAPK 77
Query: 272 ----------------QPAAPQPAVRRSARIKTTPKKFKD 295
QPA P+P R+ +K + D
Sbjct: 78 QPPKPKKPKTQEKKKKQPAKPKPGKRQRMALKLEADRLFD 117
>MAPB_RAT (P15205) Microtubule-associated protein 1B (MAP 1B)
(Neuraxin) [Contains: MAP1 light chain LC1]
Length = 2459
Score = 37.7 bits (86), Expect = 0.034
Identities = 40/175 (22%), Positives = 71/175 (39%), Gaps = 17/175 (9%)
Query: 91 LKNEIKTGFAKQHNFKGCVPVFEIIPCECCVTAEEENKSLKENVASETENKSLMEHVAAE 150
+K EIKT K K P E+ E +++ K KE E + + E
Sbjct: 637 VKKEIKT---KPEEKKEEKPKKEVAKKEDKTPLKKDEKPKKEEAKKEIKKEIKKE----- 688
Query: 151 TENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQ 210
E + L K+V +ET + ++V + ++V + EK S + S +
Sbjct: 689 -EKKELKKEVKKETPLKDAKKEVKKDEKKEVKKEEKEPKKEIKKISKDIKKSTPLSDTKK 747
Query: 211 PKTRRKLVAPQPAVRRSARIKTGEME--------KNESLTTDVASLSLGDDALAS 257
P + VA + + I G+++ K E TT+ A+ ++G A+A+
Sbjct: 748 PAALKPKVAKKEEPTKKEPIAAGKLKDKGKVKVIKKEGKTTEAAATAVGTAAVAA 802
>IF38_SCHPO (O14164) Probable eukaryotic translation initiation
factor 3 93 kDa subunit (eIF3 p93)
Length = 918
Score = 37.7 bits (86), Expect = 0.034
Identities = 33/128 (25%), Positives = 62/128 (47%), Gaps = 14/128 (10%)
Query: 122 TAEEENKSLKENVASETENKSLMEHV----------AAETENESLTKDVPEETENESLTE 171
++EEE+ S E+ +SE E++S V + ++E++S + + EETE+E +E
Sbjct: 42 SSEEESASSSESESSEEESESEESEVEVPKKKAVAASEDSESDSESSEEEEETESEEDSE 101
Query: 172 DVNESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIK 231
+ES +E E E E ++ S D++ S P + + +PA + R +
Sbjct: 102 VSDESESESESESESEEESESEEES----DESERSGPSSFLKKPEKEEAKPAGLKFLRGE 157
Query: 232 TGEMEKNE 239
+ E +E
Sbjct: 158 SSEESSDE 165
>C5P2_MOUSE (Q8K389) CDK5 regulatory subunit associated protein 2
(CDK5 activator-binding protein C48) (Fragment)
Length = 454
Score = 37.7 bits (86), Expect = 0.034
Identities = 31/106 (29%), Positives = 54/106 (50%), Gaps = 6/106 (5%)
Query: 124 EEENKSLKENVASETEN--KSLMEHVAAETENESLTKDVPE-ETENESLTEDVNESLTED 180
++E++ L+E++A + E+ + +E+ + ENE L +D+ E E N+ LTE+V S E
Sbjct: 113 QKESEKLRESLARKAESLEQLQLEYTSVREENERLQRDIIEKERHNQELTEEVCSSRQEL 172
Query: 181 VGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRR 226
E+ +S LS D L S + K KL P +++
Sbjct: 173 SRVQEEAKSRQ---QLLSQKDKLLQSLQMELKVYEKLAEEHPRLQQ 215
>YDG9_SCHPO (Q10496) Hypothetical protein C26F1.09 in chromosome I
Length = 1031
Score = 37.4 bits (85), Expect = 0.045
Identities = 32/142 (22%), Positives = 56/142 (38%), Gaps = 6/142 (4%)
Query: 133 NVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTT 192
++ +E + SL ET N+ +PE E+E + NE+ + G + +
Sbjct: 319 SIFNEEQTNSL-----TETFNDLTLDHLPENVESEPVAGKENETAKNESGASDNDHKANV 373
Query: 193 DVASLSLGDDALASPPPQPKTR-RKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLG 251
V L +DA+ + T+ L AP P V S+ I ++N+ L+ +
Sbjct: 374 HVFVLKSSEDAITLNEEKIATQDDPLEAPTPIVASSSTIFLNSNQRNDELSASGSQEPHP 433
Query: 252 DDALASPPPQPKTRRKLAAPQP 273
D S P L+ +P
Sbjct: 434 KDGTNSTSSLPLDTNNLSNSEP 455
>YAFA_SCHPO (Q09863) Hypothetical protein C29E6.10c in chromosome I
Length = 1085
Score = 37.4 bits (85), Expect = 0.045
Identities = 34/134 (25%), Positives = 58/134 (42%), Gaps = 7/134 (5%)
Query: 124 EEENKSLKENVASETENKSL-MEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVG 182
E +KS K ++ + S+ + +V +++ N+ +T+D E S DV+E + E
Sbjct: 345 EGSDKSQKGIISDSPKLLSIPLNNVPSKSLNDDITQD-----ELNSSNADVDEEVIETTS 399
Query: 183 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEK-NESL 241
EKN V S+S G+ L P+T+ P P+ ++ SL
Sbjct: 400 LEEKNVDNQEFVTSISNGNQTLEDTSHSPQTQPPFQPPYPSKADEKNSYHSDLYNFGSSL 459
Query: 242 TTDVASLSLGDDAL 255
T L++ DD L
Sbjct: 460 TVKGGILTVADDLL 473
>RGP1_HUMAN (P46060) Ran GTPase-activating protein 1
Length = 587
Score = 37.4 bits (85), Expect = 0.045
Identities = 41/183 (22%), Positives = 83/183 (44%), Gaps = 17/183 (9%)
Query: 83 DLHVSFNKLKNEIKTGFAKQHNFKGCVPVFEI----IPCECCVTAEE--ENKSLKENVAS 136
+L++SF ++K + A+ K + ++ + E C +E E ++ + +AS
Sbjct: 297 ELNLSFCEIKRDAALAVAEAMADKAELEKLDLNGNTLGEEGCEQLQEVLEGFNMAKVLAS 356
Query: 137 ETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVAS 196
++++ E E E E ++ E+ E E E+ E + G+ EK+ + + +
Sbjct: 357 LSDDEDEEEEEEGEEEEEEAEEEEEEDEEEEEEEEEEEEEEPQQRGQGEKSATPSRKILD 416
Query: 197 LSLGDDA--LASPPPQ--------PKTRRKL-VAPQPAVRRSARIKTGEMEKNESLTTDV 245
+ G+ A L+SPPP P + L + P+ +V + + T + EK S V
Sbjct: 417 PNTGEPAPVLSSPPPADVSTFLAFPSPEKLLRLGPKSSVLIAQQTDTSDPEKVVSAFLKV 476
Query: 246 ASL 248
+S+
Sbjct: 477 SSV 479
>FKB4_DROME (P54397) 39 kDa FK506-binding nuclear protein (EC
5.2.1.8) (Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase)
Length = 357
Score = 37.4 bits (85), Expect = 0.045
Identities = 38/165 (23%), Positives = 72/165 (43%), Gaps = 22/165 (13%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
E++++ EN+ + K+ + E ENES +D E+T+++S + ES E+ GE
Sbjct: 96 EDDDQMTIENLLNSKAIKNSKKSEDDEDENESGEED-EEDTDDDSQIIEEYESFLEN-GE 153
Query: 184 MEKNESLTTDVASLSLGD----DALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNE 239
E ++ + D D D + QPK + ++P + ++S + + G +K E
Sbjct: 154 EEDDDDVDEDNEESGEEDEQDSDDSEAEEEQPKAKVAKLSPGASAKKSGKEQNGVAKKEE 213
Query: 240 SLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSA 284
A + K + + QP A +PA ++ A
Sbjct: 214 ----------------AKQQQKKKEKPEAKKEQPKAKEPAKQQPA 242
>BTBC_HUMAN (Q8IY92) BTB/POZ domain containing protein 12
Length = 1151
Score = 37.4 bits (85), Expect = 0.045
Identities = 42/157 (26%), Positives = 69/157 (43%), Gaps = 9/157 (5%)
Query: 132 ENVASETEN-KSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNE-S 189
EN S EN + L+ + A+ E E+ T + + ++E E+VNE+ E++ E +
Sbjct: 123 ENCESRAENFQELLRSMWADEEEEAET--LLKSKDHEEDQENVNEAEMEEIYEFAATQRK 180
Query: 190 LTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASL- 248
L + + G+DA P + + L Q + K EME E + A+
Sbjct: 181 LLQEERAAGAGEDADWLEGGSPVSGQLLAGVQVQKQWD---KVEEMEPLEPGRDEAATTW 237
Query: 249 -SLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSA 284
+G AL P Q R AP+ AP+ A+ S+
Sbjct: 238 EKMGQCALPPPQGQHSGARGAEAPEQEAPEEALGHSS 274
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.309 0.127 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,810,420
Number of Sequences: 164201
Number of extensions: 1507285
Number of successful extensions: 7818
Number of sequences better than 10.0: 274
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 243
Number of HSP's that attempted gapping in prelim test: 7408
Number of HSP's gapped (non-prelim): 585
length of query: 295
length of database: 59,974,054
effective HSP length: 109
effective length of query: 186
effective length of database: 42,076,145
effective search space: 7826162970
effective search space used: 7826162970
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0109.1


