

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
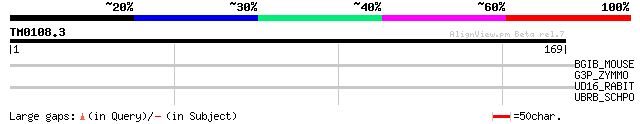
Query= TM0108.3
(169 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BGIB_MOUSE (P97402) N-acetyllactosaminide beta-1,6-N-acetylgluco... 32 0.91
G3P_ZYMMO (P09316) Glyceraldehyde-3-phosphate dehydrogenase (EC ... 31 1.5
UD16_RABIT (Q28611) UDP-glucuronosyltransferase 1-6 precursor, m... 29 4.5
UBRB_SCHPO (O13731) Ubr11 protein 29 5.9
>BGIB_MOUSE (P97402) N-acetyllactosaminide
beta-1,6-N-acetylglucosaminyl-transferase (EC 2.4.1.150)
(N-acetylglucosaminyltransferase) (I-branching enzyme)
(IGNT) (Large I antigen-forming
beta-1,6-N-acetylglucosaminyltransferase)
Length = 400
Score = 31.6 bits (70), Expect = 0.91
Identities = 15/38 (39%), Positives = 18/38 (46%), Gaps = 4/38 (10%)
Query: 6 PPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFC 43
PP SW N+ W EA GG C G++V G C
Sbjct: 312 PPNASWTGNLRAVKWMDMEAKHGG----CQGHYVHGIC 345
>G3P_ZYMMO (P09316) Glyceraldehyde-3-phosphate dehydrogenase (EC
1.2.1.12) (GAPDH)
Length = 337
Score = 30.8 bits (68), Expect = 1.5
Identities = 21/79 (26%), Positives = 35/79 (43%), Gaps = 4/79 (5%)
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLI--PPYSEIVRHIRRSIPHGMKVVFD 117
+ VD+ ++ C+ I T+ A AH + + LI P ++ R + + H D
Sbjct: 89 KLGVDIVME--CTGIFTNTEKASAHLTAGAKKVLISAPAKGDVDRTVVYGVNHKDLTADD 146
Query: 118 LIPREANMVADCLAQNAHV 136
I A+ +CLA HV
Sbjct: 147 KIVSNASCTTNCLAPVLHV 165
>UD16_RABIT (Q28611) UDP-glucuronosyltransferase 1-6 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*6) (UGT1-06)
(UGT1.6) (UGT1A6)
Length = 531
Score = 29.3 bits (64), Expect = 4.5
Identities = 16/40 (40%), Positives = 21/40 (52%)
Query: 119 IPREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQD 158
IPR +D ++ V NF V+LL PL C Y Y+D
Sbjct: 192 IPRCYTKFSDQMSFPQRVVNFLVNLLEVPLFYCLYSKYED 231
>UBRB_SCHPO (O13731) Ubr11 protein
Length = 2052
Score = 28.9 bits (63), Expect = 5.9
Identities = 13/44 (29%), Positives = 23/44 (51%), Gaps = 2/44 (4%)
Query: 91 QPLIPPYSEIVRHIRRSIPHGMKVVFDL--IPREANMVADCLAQ 132
Q +IP I+ H++ PHG ++F + + +N V+ C Q
Sbjct: 762 QGVIPQQRAILSHVQWDFPHGKNILFVMQRVAMLSNTVSSCFTQ 805
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.141 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,118,472
Number of Sequences: 164201
Number of extensions: 735673
Number of successful extensions: 1662
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1661
Number of HSP's gapped (non-prelim): 4
length of query: 169
length of database: 59,974,054
effective HSP length: 102
effective length of query: 67
effective length of database: 43,225,552
effective search space: 2896111984
effective search space used: 2896111984
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0108.3


