

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0105.17
(488 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
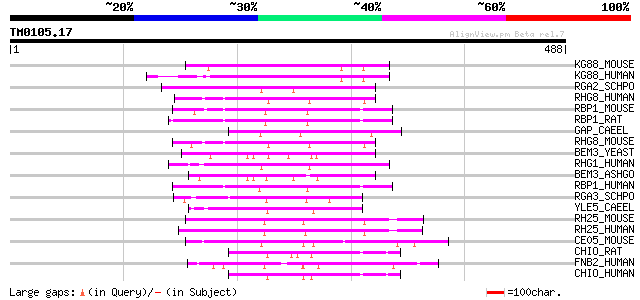
Score E
Sequences producing significant alignments: (bits) Value
KG88_MOUSE (P59281) Protein KIAA1688 homolog (Fragment) 70 2e-11
KG88_HUMAN (Q9C0H5) Protein KIAA1688 70 2e-11
RGA2_SCHPO (Q10164) Probable Rho-type GTPase-activating protein 2 67 1e-10
RHG8_HUMAN (Q9NSG0) Rho-GTPase-activating protein 8 (PP610) 60 1e-08
RBP1_MOUSE (Q62172) RalA binding protein 1 (RalBP1) (Ral interac... 59 2e-08
RBP1_RAT (Q62796) RalA binding protein 1 (RalBP1) (Ral interacti... 59 3e-08
GAP_CAEEL (P34288) GTPase-activating protein GAP (CeGAP) 59 3e-08
RHG8_MOUSE (Q9CXP4) Rho-GTPase-activating protein 8 57 8e-08
BEM3_YEAST (P32873) GTPase-activating protein BEM3 57 1e-07
RHG1_HUMAN (Q07960) Rho-GTPase-activating protein 1 (GTPase-acti... 56 2e-07
BEM3_ASHGO (Q74ZH7) GTPase-activating protein BEM3 54 7e-07
RBP1_HUMAN (Q15311) RalA binding protein 1 (RalBP1) (Ral interac... 54 1e-06
RGA3_SCHPO (O14014) Probable Rho-type GTPase-activating protein 3 53 2e-06
YLE5_CAEEL (P46941) Hypothetical protein C38D4.5 in chromosome III 53 2e-06
RH25_MOUSE (Q8BYW1) Rho-GTPase-activating protein 25 52 3e-06
RH25_HUMAN (P42331) Rho-GTPase-activating protein 25 52 3e-06
CE05_MOUSE (Q8K2H3) Protein C5orf5 homolog 51 6e-06
CHIO_RAT (Q03070) Beta-chimaerin (Beta-chimerin) (Rho-GTPase-act... 50 1e-05
FNB2_HUMAN (O75044) Formin binding protein 2 (srGAP2) 50 2e-05
CHIO_HUMAN (P52757) Beta-chimaerin (Beta-chimerin) (Rho-GTPase-a... 49 2e-05
>KG88_MOUSE (P59281) Protein KIAA1688 homolog (Fragment)
Length = 1097
Score = 69.7 bits (169), Expect = 2e-11
Identities = 53/188 (28%), Positives = 89/188 (47%), Gaps = 8/188 (4%)
Query: 155 SAKVFGVSAKSMQCSYDER--GNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFV 212
S +FG + + + ER +P + + + + G + EGIFR+ D + +
Sbjct: 898 SPSMFGSALQEVMSMQKERYPDRQLPWVQTRLSEEVLALNGDQTEGIFRVPGDIDEVNAL 957
Query: 213 RCQLNRGLVPRGVE-VHCLSGLIKAWFRELPTGVLDSLTPEQ-VMHCNSEEDCTNLVKLL 270
+ Q+++ VP G+E H + L+K W+REL ++ EQ + H S E +V L
Sbjct: 958 KLQVDQWKVPTGLEDPHVPASLLKLWYRELEEPLIPHEFYEQCIAHYESPEAAVAVVHAL 1017
Query: 271 PSTEAALLDWAINLMADVVE--HEQFNKMNARNIAMVFAPN--MTQMVDPLTALIHAVQV 326
P +L + I + V+ + KM+ N+AMV APN Q DP + +
Sbjct: 1018 PRINRMVLCYLIRFLQVFVQPANVAITKMDVSNLAMVMAPNCLRCQSDDPRVIFENTRKE 1077
Query: 327 MNFLKTLI 334
M+FL+ LI
Sbjct: 1078 MSFLRVLI 1085
>KG88_HUMAN (Q9C0H5) Protein KIAA1688
Length = 1083
Score = 69.7 bits (169), Expect = 2e-11
Identities = 60/220 (27%), Positives = 98/220 (44%), Gaps = 28/220 (12%)
Query: 121 EVRHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTI 180
E+RH + F P SA +V G+ + Y ER +P +
Sbjct: 874 EIRHAKNAVFS----------------PSMFGSALQEVMGMQRER----YPER--QLPWV 911
Query: 181 LLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNRGLVPRGVE-VHCLSGLIKAWFR 239
+ + + G + EGIFR+ D + ++ Q+++ VP G+E H + L+K W+R
Sbjct: 912 QTRLSEEVLALNGDQTEGIFRVPGDIDEVNALKLQVDQWKVPTGLEDPHVPASLLKLWYR 971
Query: 240 ELPTGVLDSLTPEQ-VMHCNSEEDCTNLVKLLPSTEAALLDWAINLMADVVE--HEQFNK 296
EL ++ EQ + H +S E +V LP +L + I + V+ + K
Sbjct: 972 ELEEPLIPHEFYEQCIAHYDSPEAAVAVVHALPRINRMVLCYLIRFLQVFVQPANVAVTK 1031
Query: 297 MNARNIAMVFAPN--MTQMVDPLTALIHAVQVMNFLKTLI 334
M+ N+AMV APN Q DP + + M+FL+ LI
Sbjct: 1032 MDVSNLAMVMAPNCLRCQSDDPRVIFENTRKEMSFLRVLI 1071
>RGA2_SCHPO (Q10164) Probable Rho-type GTPase-activating protein 2
Length = 1275
Score = 66.6 bits (161), Expect = 1e-10
Identities = 52/202 (25%), Positives = 92/202 (44%), Gaps = 14/202 (6%)
Query: 134 NGFLGLPTELQPEVPQKVPSASAKVFGVSA-KSMQCSYDERGNSVPTILLMMQNRLYSEG 192
N L + E QK + +FG+ +++ S + +P ++ L S
Sbjct: 1038 NVMLSPSSTTSAEPLQKHIVRKSGIFGLPLNEAVNISTQFNDSGLPIVVYRCIEYLESCR 1097
Query: 193 GLKAEGIFRINADNSQEEFVRCQLNRGL------VPRGVEVHCLSGLIKAWFRELPTGVL 246
K EGI+R++ S + ++ Q N G+ +VH ++GL+K + R LPT +L
Sbjct: 1098 AEKEEGIYRLSGSASTIKHLKEQFNEGVDYDLLSSDEEFDVHVIAGLLKLYLRNLPTNLL 1157
Query: 247 DS-------LTPEQVMHCNSEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNA 299
D+ L P + + +++ LP ALLD ++ + ++ E+ NKMN
Sbjct: 1158 DTSMHKLFELLPNVPNDSAALGELCDVISKLPPENFALLDSLLHHLRRIIAFEKVNKMNI 1217
Query: 300 RNIAMVFAPNMTQMVDPLTALI 321
RN+ +VF+P + D LI
Sbjct: 1218 RNVCIVFSPTLNIPSDIFMMLI 1239
>RHG8_HUMAN (Q9NSG0) Rho-GTPase-activating protein 8 (PP610)
Length = 718
Score = 60.1 bits (144), Expect = 1e-08
Identities = 50/188 (26%), Positives = 86/188 (45%), Gaps = 15/188 (7%)
Query: 146 EVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINAD 205
+ P P + FGVS + ++ +G +P +L L E GL+ EG+FR +A
Sbjct: 465 KTPPPRPPLPTQQFGVSLQYLKDK--NQGELIPPVLRFTVTYL-REKGLRTEGLFRRSAS 521
Query: 206 NSQEEFVRCQLNRGLVPRGVE---VHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEED 262
++ N+G + +H + ++K + RELP +L EQ++ E
Sbjct: 522 VQTVREIQRLYNQGKPVNFDDYGDIHIPAVILKTFLRELPQPLLTFQAYEQILGITCVES 581
Query: 263 ------CTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNM---TQM 313
C +++ LP +L + + + V FNKMN+ N+A VF N+ +Q
Sbjct: 582 SLRVTGCRQILRSLPEHNYVVLRYLMGFLHAVSRESIFNKMNSSNLACVFGLNLIWPSQG 641
Query: 314 VDPLTALI 321
V L+AL+
Sbjct: 642 VSSLSALV 649
>RBP1_MOUSE (Q62172) RalA binding protein 1 (RalBP1) (Ral
interacting protein 1) (Dinitrophenyl S-glutathione
ATPase) (DNP-SG ATPase)
Length = 647
Score = 59.3 bits (142), Expect = 2e-08
Identities = 52/205 (25%), Positives = 91/205 (44%), Gaps = 17/205 (8%)
Query: 144 QPEVPQKVPSASAKVFGV---SAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIF 200
+PEVPQ + +FGV A YD G +P + + + + G+K EG++
Sbjct: 174 EPEVPQMDAPSVKPIFGVPLVDAVERTMMYD--GVRLPAVFRECVDYM-EKHGMKCEGVY 230
Query: 201 RINADNSQEEFVRCQLNRGLVPR--GVEVHCLSGLIKAWFRELPTGVLDS-LTPEQVMHC 257
R++ S+ + ++ +R P E + ++ L+K + R+LP +L L P C
Sbjct: 231 RVSGIKSKVDELKAAYDREESPNLEEYEPNTVASLLKQYLRDLPENLLTKELMPRFEEAC 290
Query: 258 NSE------EDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMT 311
++ L++ LP LL W I + V+ E KMN +NI++V +P T
Sbjct: 291 GKTTEMEKVQEFQRLLRELPECNHLLLSWLIVHLDHVIAKELETKMNIQNISIVLSP--T 348
Query: 312 QMVDPLTALIHAVQVMNFLKTLILK 336
+ + V T++LK
Sbjct: 349 VQISNRVLYVLFTHVQELFGTVVLK 373
>RBP1_RAT (Q62796) RalA binding protein 1 (RalBP1) (Ral interacting
protein 1) (Cytocentrin) (Dinitrophenyl S-glutathione
ATPase) (DNP-SG ATPase)
Length = 646
Score = 58.9 bits (141), Expect = 3e-08
Identities = 53/207 (25%), Positives = 95/207 (45%), Gaps = 14/207 (6%)
Query: 140 PTELQPEVPQKVPSASAKVFGVS-AKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEG 198
PT+ +PEVPQ + +FG A +++ + G +P + + + + G+K EG
Sbjct: 171 PTQ-EPEVPQTDAPSLRPIFGAPFADAVERTMMYDGIRLPAVFRECVDYM-EKHGMKCEG 228
Query: 199 IFRINADNSQEEFVRCQLNRGLVPR--GVEVHCLSGLIKAWFRELPTGVLDS-LTPEQVM 255
I+R++ S+ + ++ +R P E + ++ L+K + R+LP +L L P
Sbjct: 229 IYRVSGIKSKVDELKAAYDREESPNLEEYEPNTVASLLKQYLRDLPENLLTKELMPRFEE 288
Query: 256 HCNSE------EDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPN 309
C ++ L++ LP LL W I M V+ E KMN +NI++V +P
Sbjct: 289 ACGRTTEVEKVQEFQRLLRELPEYNHLLLSWLIVHMDHVIAKELETKMNIQNISIVLSP- 347
Query: 310 MTQMVDPLTALIHAVQVMNFLKTLILK 336
T + + V T++LK
Sbjct: 348 -TVQISNRVLYVLFTHVQELFGTVLLK 373
>GAP_CAEEL (P34288) GTPase-activating protein GAP (CeGAP)
Length = 1317
Score = 58.9 bits (141), Expect = 3e-08
Identities = 50/173 (28%), Positives = 81/173 (45%), Gaps = 21/173 (12%)
Query: 193 GLKAEGIFRINADNSQEEFVRCQL-NRG-----------LVPRGVEVHCLSGLIKAWFRE 240
G+ GI+RI + + ++ L NRG L PR +V+ +S L+K + R+
Sbjct: 689 GMDTVGIYRIPGNTAAVNALKESLSNRGFDSVDLSKVESLDPRWRDVNVVSSLLKMFLRK 748
Query: 241 LPTGVL-DSLTPEQV------MHCNSEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQ 293
LP +L D L P + H N NL++ LP L + I ++++ +H
Sbjct: 749 LPEPLLTDKLYPFFIDANRISTHHNRLHKLRNLLRKLPRPHYDTLRFLIVHLSEITKHSD 808
Query: 294 FNKMNARNIAMVFAPNMTQMVDP--LTALIHAVQVMNFLKTLILKTLRERDES 344
NKM RN+A++F P++ + D T + H ++TLI L DES
Sbjct: 809 VNKMECRNLALMFGPSIVRPSDDNMATMVTHMSDQCKIIETLIHYNLWMFDES 861
>RHG8_MOUSE (Q9CXP4) Rho-GTPase-activating protein 8
Length = 425
Score = 57.4 bits (137), Expect = 8e-08
Identities = 54/194 (27%), Positives = 90/194 (45%), Gaps = 19/194 (9%)
Query: 144 QPEVPQKVPSASAKV----FGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGI 199
QP P K P + FGVS + ++ +G +P +L L E GL+ EG+
Sbjct: 174 QPPPPTKTPPPRPPLPTQQFGVSLQYLRDK--NQGELIPPVLRWTVTYL-REKGLRTEGL 230
Query: 200 FRINADNSQEEFVRCQLNRGLVPRGVE---VHCLSGLIKAWFRELPTGVLDSLTPEQVMH 256
FR +A V+ ++G + +H + ++K + RELP +L EQ++
Sbjct: 231 FRRSASAQTVRQVQRLYDQGKPVNFDDYGDMHLPAVILKTFLRELPQPLLTFQAYEQILG 290
Query: 257 CNSEED------CTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNM 310
S E C +++ LP A+L + + + +V NKMN+ N+A VF N+
Sbjct: 291 ITSVESSLRVTHCRLILRSLPEHNYAVLRYLMGFLHEVSLESISNKMNSSNLACVFGLNL 350
Query: 311 ---TQMVDPLTALI 321
+Q V L+AL+
Sbjct: 351 IWPSQGVASLSALV 364
>BEM3_YEAST (P32873) GTPase-activating protein BEM3
Length = 1128
Score = 56.6 bits (135), Expect = 1e-07
Identities = 52/203 (25%), Positives = 95/203 (46%), Gaps = 33/203 (16%)
Query: 152 PSASAKVFGVSAKS-MQCSYDERGN--SVPTILLMMQNRLYSEGGLKAEGIFRINADNS- 207
P S +FG S ++ ++ S + N +P+++ LY G++ EGIFR++ ++
Sbjct: 898 PHVSTAIFGSSLETCLRLSSHKYQNVYDLPSVVYRCLEYLYKNRGIQEEGIFRLSGSSTV 957
Query: 208 ----QEEFVR------CQLNRGLVPRGVE--------VHCLSGLIKAWFRELPT---GVL 246
QE F + C+ N + + E V+ +SGL+K + R+LP G
Sbjct: 958 IKTLQERFDKEYDVDLCRYNESIEAKDDEASPSLYIGVNTVSGLLKLYLRKLPHLLFGDE 1017
Query: 247 DSLTPEQVMHCNSEEDCT------NLVK--LLPSTEAALLDWAINLMADVVEHEQFNKMN 298
L+ ++V+ N L++ L+P +L+ L+ + E+ +FNKMN
Sbjct: 1018 QFLSFKRVVDENHNNPVQISLGFKELIESGLVPHANLSLMYALFELLVRINENSKFNKMN 1077
Query: 299 ARNIAMVFAPNMTQMVDPLTALI 321
RN+ +VF+P + + L I
Sbjct: 1078 LRNLCIVFSPTLNIPISMLQPFI 1100
>RHG1_HUMAN (Q07960) Rho-GTPase-activating protein 1
(GTPase-activating protein rhoOGAP) (Rho-related small
GTPase protein activator) (CDC42 GTPase-activating
protein) (p50-rhoGAP)
Length = 439
Score = 56.2 bits (134), Expect = 2e-07
Identities = 52/205 (25%), Positives = 89/205 (43%), Gaps = 14/205 (6%)
Query: 140 PTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEG-GLKAEG 198
P +P + P + + FGVS + +Q ++ P +++ + Y + L EG
Sbjct: 224 PATAPKPMPPRPPLPNQQ-FGVSLQHLQ---EKNPEQEPIPIVLRETVAYLQAHALTTEG 279
Query: 199 IFRINADNSQEEFVRCQLNRGL---VPRGVEVHCLSGLIKAWFRELPTGVLD-SLTPEQV 254
IFR +A+ V+ + N GL + E+H + ++K + RELP +L L P V
Sbjct: 280 IFRRSANTQVVREVQQKYNMGLPVDFDQYNELHLPAVILKTFLRELPEPLLTFDLYPHVV 339
Query: 255 MHCNSEED-----CTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPN 309
N +E +++ LP +L + + + H NKM N+A+VF PN
Sbjct: 340 GFLNIDESQRVPATLQVLQTLPEENYQVLRFLTAFLVQISAHSDQNKMTNTNLAVVFGPN 399
Query: 310 MTQMVDPLTALIHAVQVMNFLKTLI 334
+ D L + F K L+
Sbjct: 400 LLWAKDAAITLKAINPINTFTKFLL 424
>BEM3_ASHGO (Q74ZH7) GTPase-activating protein BEM3
Length = 1013
Score = 54.3 bits (129), Expect = 7e-07
Identities = 51/200 (25%), Positives = 94/200 (46%), Gaps = 38/200 (19%)
Query: 158 VFGVSAKS--MQCSYDERGN-SVPTILLMMQNRLYSEGGLKAEGIFRINADNS-----QE 209
VFG +S S+ +G +P+++ LY G++ EGIFR++ +S QE
Sbjct: 790 VFGADLRSCLQLSSHPYQGKYEIPSVVFRTLEFLYKNRGIQEEGIFRLSGSSSLIKSLQE 849
Query: 210 EFVR------CQLNRGL-VPRG--------VEVHCLSGLIKAWFRELPTGVLDS---LTP 251
+F + C N + V G V+V+ +SGL+K + R+LP + +
Sbjct: 850 QFDKEYDVDLCNYNDKVSVTPGNENQGGLYVDVNTVSGLLKLYLRKLPHMIFGDAAYMDF 909
Query: 252 EQVMHCNSEEDCTNLVKL----------LPSTEAALLDWAINLMADVVEHEQFNKMNARN 301
++++ N ++ + L+ L + AL+ L+ + E+ ++NKMN RN
Sbjct: 910 KRIVERNGDD--SKLIALEFRALVNSGRIAKEYVALMYALFELLVKITENSKYNKMNLRN 967
Query: 302 IAMVFAPNMTQMVDPLTALI 321
+ +VF+P + V+ L I
Sbjct: 968 LCIVFSPTLNIPVNILHPFI 987
>RBP1_HUMAN (Q15311) RalA binding protein 1 (RalBP1) (Ral
interacting protein 1) (76-kDa Ral-interacting protein)
(Dinitrophenyl S-glutathione ATPase) (DNP-SG ATPase)
Length = 654
Score = 53.5 bits (127), Expect = 1e-06
Identities = 49/203 (24%), Positives = 90/203 (44%), Gaps = 13/203 (6%)
Query: 144 QPEVPQKVPSASAKVFGVS-AKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRI 202
+PEVPQ +FG+ A +++ + G +P + + + + G+K EGI+R+
Sbjct: 174 EPEVPQIDVPNLKPIFGIPLADAVERTMMYDGIRLPAVFRECIDYV-EKYGMKCEGIYRV 232
Query: 203 NADNSQEEFVRCQLNR--GLVPRGVEVHCLSGLIKAWFRELPTGVLDS-LTPEQVMHCNS 259
+ S+ + ++ +R E + ++ L+K + R+LP +L L P C
Sbjct: 233 SGIKSKVDELKAAYDREESTNLEDYEPNTVASLLKQYLRDLPENLLTKELMPRFEEACGR 292
Query: 260 E------EDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQM 313
++ L+K LP L+ W I M V+ E KMN +NI++V +P T
Sbjct: 293 TTETEKVQEFQRLLKELPECNYLLISWLIVHMDHVIAKELETKMNIQNISIVLSP--TVQ 350
Query: 314 VDPLTALIHAVQVMNFLKTLILK 336
+ + V ++LK
Sbjct: 351 ISNRVLYVFFTHVQELFGNVVLK 373
>RGA3_SCHPO (O14014) Probable Rho-type GTPase-activating protein 3
Length = 969
Score = 53.1 bits (126), Expect = 2e-06
Identities = 42/178 (23%), Positives = 86/178 (47%), Gaps = 16/178 (8%)
Query: 145 PEVPQKVP--SASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRI 202
PE+P ++P S +FG +S++ G+ +P ++ M + + + G L+ EGI+RI
Sbjct: 762 PEMPTRMPPPGPSPTMFG---RSLENQLKIEGSVLPQVIAMCVSCVDAHG-LEVEGIYRI 817
Query: 203 NADNSQEEFVRCQLNRGLVPRG---VEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNS 259
+ SQ + + G + ++ + ++K + LP V+ E+++
Sbjct: 818 SGSASQVRVLVDEFENGSIRMEHLTSDLFACTSVLKTYLHRLPEPVIPGTQYEELLEAEK 877
Query: 260 ----EEDCTNLVKLLPSTEAALLD---WAINLMADVVEHEQFNKMNARNIAMVFAPNM 310
EE +V+++ + A L + I + V +H + N MN++N++ VFAP +
Sbjct: 878 IEKEEEKIERVVEVMKTLHPAHLSVFRFLIAHLGRVCKHAEKNLMNSKNVSTVFAPTL 935
>YLE5_CAEEL (P46941) Hypothetical protein C38D4.5 in chromosome III
Length = 837
Score = 52.8 bits (125), Expect = 2e-06
Identities = 40/163 (24%), Positives = 73/163 (44%), Gaps = 14/163 (8%)
Query: 158 VFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLN 217
VFG S S C ++ NS+ + + + GL+ +GI+R++ + S + +RCQ +
Sbjct: 607 VFG-STLSAICQHE---NSLVPKFIRVITEVIESKGLETDGIYRVSGNLSAVQKIRCQAD 662
Query: 218 RGLVPRGV---EVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTN-------LV 267
+ V ++H L+G +K +FREL + ++ + T L+
Sbjct: 663 QDNYKALVSEEDIHVLTGALKLFFRELTDPLFPISLHKEYTSAMQMPNATTRFKKFEELL 722
Query: 268 KLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNM 310
LP+ L + + V H N+M N+A+VF P +
Sbjct: 723 SRLPNENRETLKMLLRHLNRVASHSSQNRMQQHNLAIVFGPTL 765
>RH25_MOUSE (Q8BYW1) Rho-GTPase-activating protein 25
Length = 648
Score = 52.0 bits (123), Expect = 3e-06
Identities = 48/224 (21%), Positives = 93/224 (41%), Gaps = 20/224 (8%)
Query: 155 SAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRC 214
S VFG + G + IL+ E G+ EGIFR+ ++ + +R
Sbjct: 154 SGAVFGQRLDETVAYEQKFGPHLVPILVEKCAEFILEHGVSEEGIFRLPGQDNLVKQLRD 213
Query: 215 QLNRGLVP---RGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHC---------NSEED 262
+ G P R +VH ++ L+K + R+LP V+ E + C ++++
Sbjct: 214 AFDAGERPSFDRDTDVHTVASLLKLYLRDLPEPVVPWSQYEGFLLCGQLMNADEAKAQQE 273
Query: 263 CTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNM--TQMVDPLTAL 320
+ LP LL + + ++ + NKM+ N+A V N+ +++ DP +
Sbjct: 274 LVKQLSTLPRDNYNLLSYICRFLHEIQLNCAVNKMSVDNLATVIGVNLIRSKVEDPAVIM 333
Query: 321 IHAVQVMNFLKTLILKTLRERDESMAKARQLSSLLNSPSCKGDS 364
Q+ + +I RD + + + ++ P+ K D+
Sbjct: 334 RGTPQIQRVMTMMI------RDHEVLFPKSKDAPISPPAQKNDA 371
>RH25_HUMAN (P42331) Rho-GTPase-activating protein 25
Length = 638
Score = 52.0 bits (123), Expect = 3e-06
Identities = 49/229 (21%), Positives = 94/229 (40%), Gaps = 20/229 (8%)
Query: 149 QKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQ 208
++V VFG + G + IL+ E G EGIFR+ ++
Sbjct: 140 RRVAGTPCGVFGQRLDETVAYEQKFGPHLVPILVEKCAEFILEHGRNEEGIFRLPGQDNL 199
Query: 209 EEFVRCQLNRGLVP---RGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCN------- 258
+ +R + G P R +VH ++ L+K + R+LP V+ E + C
Sbjct: 200 VKQLRDAFDAGERPSFDRDTDVHTVASLLKLYLRDLPEPVVPWSQYEGFLLCGQLTNADE 259
Query: 259 --SEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNM--TQMV 314
++++ + +LP +LL + + ++ + NKM+ N+A V N+ +++
Sbjct: 260 AKAQQELMKQLSILPRDNYSLLSYICRFLHEIQLNCAVNKMSVDNLATVIGVNLIRSKVE 319
Query: 315 DPLTALIHAVQVMNFLKTLILKTLRERDESMAKARQLSSLLNSPSCKGD 363
DP + Q+ + +I RD + + L+ P+ K D
Sbjct: 320 DPAVIMRGTPQIQRVMTMMI------RDHEVLFPKSKDIPLSPPAQKND 362
>CE05_MOUSE (Q8K2H3) Protein C5orf5 homolog
Length = 851
Score = 51.2 bits (121), Expect = 6e-06
Identities = 53/248 (21%), Positives = 112/248 (44%), Gaps = 18/248 (7%)
Query: 155 SAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRC 214
++K+FG+ +Q N VP I+ + + + GGL+ +G+F++N + E++R
Sbjct: 17 ASKIFGIPLDELQQG-GHPDNEVPFIVRHVVDYIEEHGGLEQQGLFQVNGNAETVEWLRQ 75
Query: 215 QLNRGL---VPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHC----NSEEDCTN-- 265
+ + G + + +V L++ + +ELP V+ ++ N+E++
Sbjct: 76 RYDSGEEVDLVKEADVPSAISLLRFFLQELPEPVIPGSLHIHLLQLSQDYNNEDEFGRKL 135
Query: 266 --LVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHA 323
L++ LP +LL + +A+V H + +A ++A VF P++ + + +
Sbjct: 136 RFLLQQLPPVNYSLLKFLCRFLANVASHHE-EIWSANSLAAVFGPDVFHIYTDVEDMKEQ 194
Query: 324 VQVMNFLKTLILKTLR--ERDESMAKARQLSSL---LNSPSCKGDSHPFKDNREESSAQP 378
V + L+ E +E + LSS+ +N S + + ++ EE +
Sbjct: 195 EIVSRIMAGLLENYYEFFENEEEDFSSNDLSSITEQVNELSEEEEEDEKLEHIEELPEEG 254
Query: 379 VDTCATMP 386
V+ A MP
Sbjct: 255 VEKSAGMP 262
>CHIO_RAT (Q03070) Beta-chimaerin (Beta-chimerin)
(Rho-GTPase-activating protein 3)
Length = 295
Score = 50.4 bits (119), Expect = 1e-05
Identities = 43/164 (26%), Positives = 81/164 (49%), Gaps = 16/164 (9%)
Query: 193 GLKAEGIFRINADNSQEEFVRCQLNRGLVPRGV------EVHCLSGLIKAWFRELPTGVL 246
GLK+EG++R++ E V+ +R + +++ ++G +K +FR+LP ++
Sbjct: 132 GLKSEGLYRVSGFTEHIEDVKMAFDRDGEKADISANIYPDINIITGALKLYFRDLPIPII 191
Query: 247 --DSLTP--EQVMHCNSEEDCT---NLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNA 299
D+ T E N++E ++ LLP L + + + V +E+ N MNA
Sbjct: 192 TYDTYTKFIEAAKISNADERLEAVHEVLMLLPPAHYETLRYLMIHLKKVTMNEKDNLMNA 251
Query: 300 RNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTLRERDE 343
N+ +VF P T M P + + + M + K LI++ L E ++
Sbjct: 252 ENLGIVFGP--TLMRPPEDSTLTTLHDMRYQK-LIVQILIENED 292
>FNB2_HUMAN (O75044) Formin binding protein 2 (srGAP2)
Length = 1071
Score = 49.7 bits (117), Expect = 2e-05
Identities = 54/252 (21%), Positives = 116/252 (45%), Gaps = 38/252 (15%)
Query: 157 KVFGVSAKSMQCSYDERGNSV------PTILLMMQN--RLYSEGGLKAEGIFRINADNSQ 208
K G S ++ CS R ++V I L++++ R S GL+ EGIFR++ +
Sbjct: 476 KTLGESQRT-DCSLARRSSTVRKQDSSQAIPLVVESCIRFISRHGLQHEGIFRVSGSQVE 534
Query: 209 EEFVRCQLNRGLVP-----RGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMH----C-- 257
++ RG P ++ ++G++K +FR G+ L P+ + H C
Sbjct: 535 VNDIKNAFERGEDPLAGDQNDHDMDSIAGVLKLYFR----GLEHPLFPKDIFHDLMACVT 590
Query: 258 --NSEEDCTNLVKLL---PSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQ 312
N +E ++ K+L P T ++ + + + + + N M+ N+A+ F P++
Sbjct: 591 MDNLQERALHIRKVLLVLPKTTLIIMRYLFAFLNHLSQFSEENMMDPYNLAICFGPSLMS 650
Query: 313 MVDPLTALIHAVQVMNFLKTLILK------TLRERDESM-AKARQLSSLLNSPSCKGDSH 365
+ + + V +KT+I++ + RE + + ++ + +SP G++
Sbjct: 651 VPEGHDQVSCQAHVNELIKTIIIQHENIFPSPRELEGPVYSRGGSMEDYCDSP--HGETT 708
Query: 366 PFKDNREESSAQ 377
P +D+ ++ +A+
Sbjct: 709 PVEDSTQDVTAE 720
>CHIO_HUMAN (P52757) Beta-chimaerin (Beta-chimerin)
(Rho-GTPase-activating protein 3)
Length = 468
Score = 49.3 bits (116), Expect = 2e-05
Identities = 42/164 (25%), Positives = 80/164 (48%), Gaps = 16/164 (9%)
Query: 193 GLKAEGIFRINADNSQEEFVRCQLNRGLVPRGV------EVHCLSGLIKAWFRELPTGVL 246
GLK+EG++R++ E V+ +R + +++ ++G +K +FR+LP V+
Sbjct: 305 GLKSEGLYRVSGFTEHIEDVKMAFDRDGEKADISANVYPDINIITGALKLYFRDLPIPVI 364
Query: 247 DSLTPEQVMHC----NSEEDCT---NLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNA 299
T + + N++E ++ LLP L + + + V +E+ N MNA
Sbjct: 365 TYDTYSKFIDAAKISNADERLEAVHEVLMLLPPAHYETLRYLMIHLKKVTMNEKDNFMNA 424
Query: 300 RNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTLRERDE 343
N+ +VF P T M P + + + M + K LI++ L E ++
Sbjct: 425 ENLGIVFGP--TLMRPPEDSTLTTLHDMRYQK-LIVQILIENED 465
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 61,904,019
Number of Sequences: 164201
Number of extensions: 2876287
Number of successful extensions: 30609
Number of sequences better than 10.0: 540
Number of HSP's better than 10.0 without gapping: 425
Number of HSP's successfully gapped in prelim test: 123
Number of HSP's that attempted gapping in prelim test: 19409
Number of HSP's gapped (non-prelim): 4451
length of query: 488
length of database: 59,974,054
effective HSP length: 114
effective length of query: 374
effective length of database: 41,255,140
effective search space: 15429422360
effective search space used: 15429422360
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0105.17


