

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0102b.4
(278 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
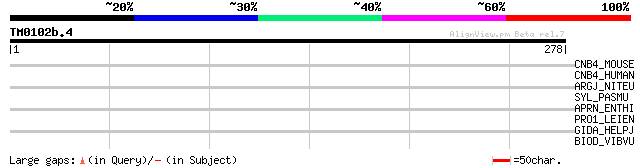
CNB4_MOUSE (Q8VEG4) Protein C14orf114 homolog 35 0.15
CNB4_HUMAN (Q9NVH0) Protein C14orf114 35 0.26
ARGJ_NITEU (Q82SU1) Arginine biosynthesis bifunctional protein a... 32 1.7
SYL_PASMU (P57923) Leucyl-tRNA synthetase (EC 6.1.1.4) (Leucine-... 32 2.2
APRN_ENTHI (P20301) Antigenic protein NP1 (Non-pathogenic protei... 31 2.9
PRO1_LEIEN (P13865) Probable transport protein (LTP) 31 3.8
GIDA_HELPJ (Q9ZML9) Glucose inhibited division protein A 31 3.8
BIOD_VIBVU (Q8D8N2) Dethiobiotin synthetase (EC 6.3.3.3) (Dethio... 30 8.5
>CNB4_MOUSE (Q8VEG4) Protein C14orf114 homolog
Length = 496
Score = 35.4 bits (80), Expect = 0.15
Identities = 30/98 (30%), Positives = 44/98 (44%), Gaps = 5/98 (5%)
Query: 7 EGSKVDRGRMTALELWSGPEIPKEEWMLRSLVREMKNLELIPS--LREAFMVEGHITIQV 64
E +V G L S P KEE L +RE N ++I L EA +E I +
Sbjct: 373 ERRQVRSGARALLNAESLPAHRKEE--LLHALREFYNTDIITEEMLHEAASLETRIYNE- 429
Query: 65 RYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKP 102
Y+ ++ H E L L+Q WR F+D+++P
Sbjct: 430 SYIPHGLKVVQRHTEGGLRSLMQLESRWRQHFLDSMQP 467
>CNB4_HUMAN (Q9NVH0) Protein C14orf114
Length = 496
Score = 34.7 bits (78), Expect = 0.26
Identities = 28/98 (28%), Positives = 46/98 (46%), Gaps = 5/98 (5%)
Query: 7 EGSKVDRGRMTALELWSGPEIPKEEWMLRSLVREMKNLELIPS--LREAFMVEGHITIQV 64
E +V G L S P KEE L +RE N +++ L+EA +E I+ +
Sbjct: 373 ERRQVRSGARALLNAESLPTQRKEE--LLQALREFYNTDVVTEEMLQEAASLETRISNE- 429
Query: 65 RYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKP 102
Y+ ++ H + L L+Q WR F+D+++P
Sbjct: 430 NYVPHGLKVVQCHSQGGLRSLMQLESRWRQHFLDSMQP 467
>ARGJ_NITEU (Q82SU1) Arginine biosynthesis bifunctional protein argJ
[Includes: Glutamate N-acetyltransferase (EC 2.3.1.35)
(Ornithine acetyltransferase) (Ornithine transacetylase)
(OATase); Amino-acid acetyltransferase (EC 2.3.1.1)
(N-acetylglutamate syn
Length = 409
Score = 32.0 bits (71), Expect = 1.7
Identities = 15/37 (40%), Positives = 24/37 (64%)
Query: 161 VRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPN 197
+ TT + ++R+I I GTT TI+ I +G ++HPN
Sbjct: 154 IMTTDIVPKGVSRQIQINGTTVTITGIAKGSGMIHPN 190
>SYL_PASMU (P57923) Leucyl-tRNA synthetase (EC 6.1.1.4)
(Leucine--tRNA ligase) (LeuRS)
Length = 860
Score = 31.6 bits (70), Expect = 2.2
Identities = 23/81 (28%), Positives = 34/81 (41%), Gaps = 7/81 (8%)
Query: 168 FITLARKILIKGTTYTISIIEEGEALL-----HPNCCCREESSVEDDVFSGWSLNGVTVT 222
F L +K L+ T T++ E +L H CC R ++ VE W +
Sbjct: 140 FTELYKKGLVYKKTSTVNWCPNDETVLANEQVHEGCCWRCDTPVEQKEIPQWFIKITDYA 199
Query: 223 PELSESGGEDDDGLWPVETKS 243
+L GG D LWP + K+
Sbjct: 200 EQL--LGGLDHLPLWPDQVKT 218
>APRN_ENTHI (P20301) Antigenic protein NP1 (Non-pathogenic protein
1) (Fragment)
Length = 640
Score = 31.2 bits (69), Expect = 2.9
Identities = 23/100 (23%), Positives = 42/100 (42%), Gaps = 14/100 (14%)
Query: 95 EFVDNLKPWSSSFAPINRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
+++DN+ WS+ PI+ + GI L + T + + D KI + L
Sbjct: 15 QWLDNISRWSNDRMPIDSI------GIDLGLQTTQPY-----SINDTFKIGSPIGGMIYL 63
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALL 194
F + + F + R +I Y I++ EE ++L
Sbjct: 64 RFDTTFTNSFYVTFYNVGRASII---NYNITMNEEWNSVL 100
>PRO1_LEIEN (P13865) Probable transport protein (LTP)
Length = 567
Score = 30.8 bits (68), Expect = 3.8
Identities = 23/86 (26%), Positives = 40/86 (45%), Gaps = 3/86 (3%)
Query: 115 WVRCTGIPLHMWT-MDCFKHLL--LQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITL 171
W R G+ ++T + F L L +G I+ D D + V+ L V +T + +T+
Sbjct: 246 WKRTIGVMFQVFTTLGIFVAALMGLALGQSIRFDHDGDQKVMARMQGLCVFSTLFSLLTV 305
Query: 172 ARKILIKGTTYTISIIEEGEALLHPN 197
I+ + + EEG A L+P+
Sbjct: 306 VLGIVTRESRAKFDGGEEGRAELNPS 331
>GIDA_HELPJ (Q9ZML9) Glucose inhibited division protein A
Length = 621
Score = 30.8 bits (68), Expect = 3.8
Identities = 30/135 (22%), Positives = 55/135 (40%), Gaps = 23/135 (17%)
Query: 57 EGHITIQVRYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSSFAPINRVTWV 116
+GH+T +V L ++T+H +L SKG V
Sbjct: 54 KGHLTKEVDVLGGAMGIITDHSGLQY-RVLNASKG----------------------PAV 90
Query: 117 RCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARKIL 176
R T + M T F L+ + + ++ +E ++LE ++ TT +N A+K++
Sbjct: 91 RGTRAQIDMDTYRIFARNLVLNTPNLSVSQEMTESLILENDEVVGVTTNINNTYRAKKVI 150
Query: 177 IKGTTYTISIIEEGE 191
I T+ ++ GE
Sbjct: 151 ITTGTFLKGVVHIGE 165
>BIOD_VIBVU (Q8D8N2) Dethiobiotin synthetase (EC 6.3.3.3)
(Dethiobiotin synthase) (DTB synthetase) (DTBS)
Length = 227
Score = 29.6 bits (65), Expect = 8.5
Identities = 23/75 (30%), Positives = 36/75 (47%), Gaps = 14/75 (18%)
Query: 74 LTNHQESNLVELLQGSKGWRSEFVDNLKPWSSSFAPINRVTWVRCTGIPLHMWT---MDC 130
L H+E+ V L++G+ GWR D + S + TWV+ +P+ + + C
Sbjct: 103 LAQHKENADVVLVEGAGGWRVPVSD-----TDSLS-----TWVKQEDLPVVLVVGIKLGC 152
Query: 131 FKHLLLQVGDVIKID 145
H LL +VIK D
Sbjct: 153 LSHALL-TAEVIKAD 166
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,976,456
Number of Sequences: 164201
Number of extensions: 1353812
Number of successful extensions: 2938
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 2936
Number of HSP's gapped (non-prelim): 8
length of query: 278
length of database: 59,974,054
effective HSP length: 109
effective length of query: 169
effective length of database: 42,076,145
effective search space: 7110868505
effective search space used: 7110868505
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0102b.4


