

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.3
(244 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
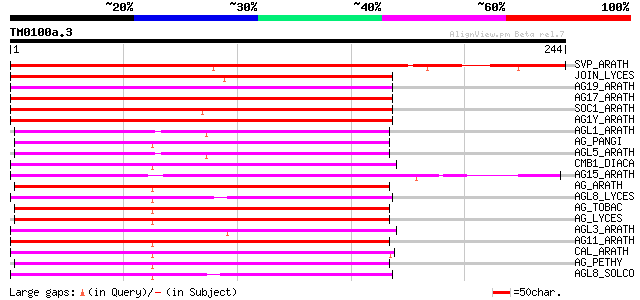
Score E
Sequences producing significant alignments: (bits) Value
SVP_ARATH (Q9FVC1) SHORT VEGETATIVE PHASE protein 181 1e-45
JOIN_LYCES (Q9FUY6) MADS-box JOINTLESS protein (LeMADS) 174 2e-43
AG19_ARATH (O82743) Agamous-like MADS box protein AGL19 119 5e-27
AG17_ARATH (Q38840) Agamous-like MADS box protein AGL17 118 1e-26
SOC1_ARATH (O64645) SUPPRESSOR OF CONSTANS OVEREXPRESSION 1 prot... 117 3e-26
AG1Y_ARATH (Q9SZJ6) Agamous-like MADS box protein At4g37940 116 4e-26
AGL1_ARATH (P29381) Agamous-like MADS box protein AGL1 (Protein ... 114 2e-25
AG_PANGI (Q40872) Floral homeotic protein AGAMOUS (GAG2) 114 3e-25
AGL5_ARATH (P29385) Agamous-like MADS box protein AGL5 113 4e-25
CMB1_DIACA (Q39685) MADS box protein CMB1 112 8e-25
AG15_ARATH (Q38847) Agamous-like MADS box protein AGL15 111 1e-24
AG_ARATH (P17839) Floral homeotic protein AGAMOUS 111 2e-24
AGL8_LYCES (Q40170) Agamous-like MADS box protein AGL8 homolog (... 111 2e-24
AG_TOBAC (Q43585) Floral homeotic protein AGAMOUS (NAG1) 110 2e-24
AG_LYCES (Q40168) Floral homeotic protein AGAMOUS (TAG1) 110 3e-24
AGL3_ARATH (P29383) Agamous-like MADS box protein AGL3 110 3e-24
AG11_ARATH (Q38836) Agamous-like MADS box protein AGL11 110 4e-24
CAL_ARATH (Q39081) Transcription factor CAULIFLOWER (Agamous-lik... 109 5e-24
AG_PETHY (Q40885) Floral homeotic protein AGAMOUS (pMADS3) 109 5e-24
AGL8_SOLCO (O22328) Agamous-like MADS box protein AGL8 homolog 109 5e-24
>SVP_ARATH (Q9FVC1) SHORT VEGETATIVE PHASE protein
Length = 240
Score = 181 bits (460), Expect = 1e-45
Identities = 107/252 (42%), Positives = 162/252 (63%), Gaps = 22/252 (8%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQI+KIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLFE+ SS
Sbjct: 1 MAREKIQIRKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + S +K PS +Q E+ + + K++ +K+H LRQ+ GE+LQGL
Sbjct: 61 SMKEVLERHNLQSKNLEKLDQPSLELQLVENSDHARMSKEIADKSHRLRQMRGEELQGLD 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K +K M EIS L++K ++L++EN++L+Q L + N
Sbjct: 121 IEELQQLEKALETGLTRVIETKSDKIMSEISELQKKGMQLMDENKRLRQQGTQLTE--EN 178
Query: 180 EYL----CSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTI---SSSSYLLEED 232
E L C+ + ++GG +S++ + Y + QS ES +S+ ++ +
Sbjct: 179 ERLGMQICNNVHAHGGAESENAAV------------YEEGQSSESITNAGNSTGAPVDSE 226
Query: 233 GSDTSLKLGLVY 244
SDTSL+LGL Y
Sbjct: 227 SSDTSLRLGLPY 238
>JOIN_LYCES (Q9FUY6) MADS-box JOINTLESS protein (LeMADS)
Length = 265
Score = 174 bits (440), Expect = 2e-43
Identities = 90/169 (53%), Positives = 125/169 (73%), Gaps = 1/169 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF+Y+SS
Sbjct: 1 MAREKIQIKKIDNSTARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFDYSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSN-DTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L D +S +K PS +Q +SN L K++ EK+H LRQ+ GE+LQGL
Sbjct: 61 SMKQILERRDLHSKNLEKLDQPSLELQLVENSNYSRLSKEISEKSHRLRQMRGEELQGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+ +LQ+LE L+ L+ V K +K M+EI+ L++K + L+EEN+KL+Q
Sbjct: 121 IEELQQLERSLETGLSRVIERKGDKIMREINQLQQKGMHLMEENEKLRQ 169
>AG19_ARATH (O82743) Agamous-like MADS box protein AGL19
Length = 219
Score = 119 bits (299), Expect = 5e-27
Identities = 66/168 (39%), Positives = 99/168 (58%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + ++K+I+N +SRQVTFSKRR GL KKA ELS LCDA++AL++FS +KL+E++SS
Sbjct: 1 MVRGKTEMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALVIFSPRSKLYEFSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S + + Q D L KK+E+ R+L GE + ++
Sbjct: 61 SIAATIERYQRRIKEIGNNHKRNDNSQQARDETSGLTKKIEQLEISKRKLLGEGIDACSI 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+LQ+LE L RSL+ + K + +EI LK +E L++EN+ LK+
Sbjct: 121 EELQQLENQLDRSLSRIRAKKYQLLREEIEKLKAEERNLVKENKDLKE 168
>AG17_ARATH (Q38840) Agamous-like MADS box protein AGL17
Length = 227
Score = 118 bits (295), Expect = 1e-26
Identities = 66/168 (39%), Positives = 102/168 (60%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I+KID+ +SRQVTFSKRRKGL KKA+EL+ LCDA++ LI+FS T+KL+++ASS
Sbjct: 1 MGRGKIVIQKIDDSTSRQVTFSKRRKGLIKKAKELAILCDAEVCLIIFSNTDKLYDFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S + + + P+ ++F +TLR+++ RQL G +L GL++
Sbjct: 61 SVKSTIERFNTAKMEEQELMNPASEVKFWQREAETLRQELHSLQENYRQLTGVELNGLSV 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+LQ +E L+ SL + +++ EI L RK + EN +L +
Sbjct: 121 KELQNIESQLEMSLRGIRMKREQILTNEIKELTRKRNLVHHENLELSR 168
>SOC1_ARATH (O64645) SUPPRESSOR OF CONSTANS OVEREXPRESSION 1 protein
(Agamous-like MADS box protein AGL20)
Length = 214
Score = 117 bits (292), Expect = 3e-26
Identities = 67/169 (39%), Positives = 103/169 (60%), Gaps = 1/169 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + Q+K+I+N +SRQVTFSKRR GL KKA ELS LCDA+++LI+FS KL+E+ASS
Sbjct: 1 MVRGKTQMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVSLIIFSPKGKLYEFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPS-PTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
+ + ++ T P S MQ + KK+E+ R+L GE + +
Sbjct: 61 NMQDTIDRYLRHTKDRVSTKPVSEENMQHLKYEAANMMKKIEQLEASKRKLLGEGIGTCS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+ +LQ++E+ L++S+ + K + F ++I LK+KE L EN+KL +
Sbjct: 121 IEELQQIEQQLEKSVKCIRARKTQVFKEQIEQLKQKEKALAAENEKLSE 169
>AG1Y_ARATH (Q9SZJ6) Agamous-like MADS box protein At4g37940
Length = 228
Score = 116 bits (291), Expect = 4e-26
Identities = 64/168 (38%), Positives = 103/168 (61%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I++ID+ +SRQVTFSKRRKGL KKA+EL+ LCDA++ LI+FS+T KL+++ASS
Sbjct: 1 MGRGKIVIQRIDDSTSRQVTFSKRRKGLIKKAKELAILCDAEVGLIIFSSTGKLYDFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S ++ + + + P+ ++F LR+++ RQ+ GE L GL++
Sbjct: 61 SMKSVIDRYNKSKIEQQQLLNPASEVKFWQREAAVLRQELHALQENHRQMMGEQLNGLSV 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
++L LE ++ SL + K++ QEI L +K + +EN L +
Sbjct: 121 NELNSLENQIEISLRGIRMRKEQLLTQEIQELSQKRNLIHQENLDLSR 168
>AGL1_ARATH (P29381) Agamous-like MADS box protein AGL1 (Protein
Shatterproof 1)
Length = 248
Score = 114 bits (285), Expect = 2e-25
Identities = 64/168 (38%), Positives = 100/168 (59%), Gaps = 5/168 (2%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSR 62
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL++FS +L+EYA++S
Sbjct: 18 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFSTRGRLYEYANNSV 77
Query: 63 GGLLPLPDANSAT*DKTTPPSPT---MQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
G + A D PPS T Q+ LR+++ + + R + GE L L
Sbjct: 78 RG--TIERYKKACSDAVNPPSVTEANTQYYQQEASKLRRQIRDIQNSNRHIVGESLGSLN 135
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE L++ ++ V K+E + EI ++++E+EL N L+
Sbjct: 136 FKELKNLEGRLEKGISRVRSKKNELLVAEIEYMQKREMELQHNNMYLR 183
>AG_PANGI (Q40872) Floral homeotic protein AGAMOUS (GAG2)
Length = 242
Score = 114 bits (284), Expect = 3e-25
Identities = 63/166 (37%), Positives = 100/166 (59%), Gaps = 1/166 (0%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS- 61
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS +L+EYA++S
Sbjct: 19 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYANNSV 78
Query: 62 RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLH 121
+G + A + + + ++ QF LR+++ R + GE L LT+
Sbjct: 79 KGTIERYKKACTDSPNTSSVSEANAQFYQQEASKLRQEISSIQKNNRNMMGESLGSLTVR 138
Query: 122 QLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
L+ LE L++ ++ + K+E EI +++KE++L NQ L+
Sbjct: 139 DLKGLETKLEKGISRIRSKKNELLFAEIEYMQKKEIDLHNNNQYLR 184
>AGL5_ARATH (P29385) Agamous-like MADS box protein AGL5
Length = 246
Score = 113 bits (283), Expect = 4e-25
Identities = 63/168 (37%), Positives = 100/168 (59%), Gaps = 5/168 (2%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSR 62
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL++FS +L+EYA++S
Sbjct: 18 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFSTRGRLYEYANNSV 77
Query: 63 GGLLPLPDANSAT*DKTTPPSPT---MQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
G + A D PP+ T Q+ LR+++ + + R + GE L L
Sbjct: 78 RG--TIERYKKACSDAVNPPTITEANTQYYQQEASKLRRQIRDIQNLNRHILGESLGSLN 135
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE L++ ++ V K E + EI ++++E+EL +N L+
Sbjct: 136 FKELKNLESRLEKGISRVRSKKHEMLVAEIEYMQKREIELQNDNMYLR 183
>CMB1_DIACA (Q39685) MADS box protein CMB1
Length = 233
Score = 112 bits (280), Expect = 8e-25
Identities = 68/171 (39%), Positives = 100/171 (57%), Gaps = 1/171 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCDA++ALIVFS KL+E+ S+
Sbjct: 1 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIVFSNRGKLYEFCST 60
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S L S +T+ PS + L+ KV+ R L GEDL L+
Sbjct: 61 SCMNKTLERYQRCSYGSLETSQPSKETESSYQEYLKLKAKVDVLQRSHRNLLGEDLGELS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
+L++LE L +SL + +K + + +++ L++KE L E N+ LK L
Sbjct: 121 TKELEQLEHQLDKSLRQIRSIKTQHMLDQLADLQKKEEMLFESNRALKTKL 171
>AG15_ARATH (Q38847) Agamous-like MADS box protein AGL15
Length = 268
Score = 111 bits (278), Expect = 1e-24
Identities = 80/244 (32%), Positives = 125/244 (50%), Gaps = 31/244 (12%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N +SRQVTFSKRR GL KKA+ELS LCDA++A+IVFS + KLFEY+S+
Sbjct: 1 MGRGKIEIKRIENANSRQVTFSKRRSGLLKKARELSVLCDAEVAVIVFSKSGKLFEYSST 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
+ S + + + + + D L+ ++ + + QL G+ L LT
Sbjct: 61 G------MKQTLSRYGNHQSSSASKAEEDCAEVDILKDQLSKLQEKHLQLQGKGLNPLTF 114
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLL--F 178
+LQ LE+ L +L +V K+ ++ + KE EN+ L++ + L L F
Sbjct: 115 KELQSLEQQLYHALITVRERKERLLTNQLEESRLKEQRAELENETLRRQVQELRSFLPSF 174
Query: 179 NEYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSL 238
Y+ S I + ID ++ L+ + S L+ SDT+L
Sbjct: 175 THYVPSYIKCF-AIDPKNALINH----------------------DSKCSLQNTDSDTTL 211
Query: 239 KLGL 242
+LGL
Sbjct: 212 QLGL 215
>AG_ARATH (P17839) Floral homeotic protein AGAMOUS
Length = 252
Score = 111 bits (277), Expect = 2e-24
Identities = 63/166 (37%), Positives = 103/166 (61%), Gaps = 1/166 (0%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS- 61
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+ +L+EY+++S
Sbjct: 19 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNNSV 78
Query: 62 RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLH 121
+G + A S + + Q+ + LR+++ + RQL GE + ++
Sbjct: 79 KGTIERYKKAISDNSNTGSVAEINAQYYQQESAKLRQQIISIQNSNRQLMGETIGSMSPK 138
Query: 122 QLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE L+RS+ + K+E EI ++++EV+L +NQ L+
Sbjct: 139 ELRNLEGRLERSITRIRSKKNELLFSEIDYMQKREVDLHNDNQILR 184
>AGL8_LYCES (Q40170) Agamous-like MADS box protein AGL8 homolog
(TM4)
Length = 227
Score = 111 bits (277), Expect = 2e-24
Identities = 67/175 (38%), Positives = 99/175 (56%), Gaps = 12/175 (6%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+K+I+N +RQVTFSKRR GL KKA E+S LCDA++ LIVFS KLFEYA+
Sbjct: 1 MGRGRVQLKRIENKINRQVTFSKRRSGLLKKAHEISVLCDAEVGLIVFSTKGKLFEYAND 60
Query: 61 S-------RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGE 113
S R + D T+P S T++ L+ ++E + GE
Sbjct: 61 SCMERILERYERYSFAEKQLVPTDHTSPVSWTLEHRK-----LKARLEVLQRNQKHYVGE 115
Query: 114 DLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
DL+ L++ +LQ LE L +L + K++ + IS L++K+ L E+N +L +
Sbjct: 116 DLESLSMKELQNLEHQLDSALKHIRSRKNQLMHESISVLQKKDRALQEQNNQLSK 170
>AG_TOBAC (Q43585) Floral homeotic protein AGAMOUS (NAG1)
Length = 248
Score = 110 bits (276), Expect = 2e-24
Identities = 60/166 (36%), Positives = 101/166 (60%), Gaps = 1/166 (0%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS- 61
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+ +L+EYA++S
Sbjct: 19 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANNSV 78
Query: 62 RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLH 121
+ + A S + + + Q+ LR ++ ++ R + GE L L+L
Sbjct: 79 KATIERYKKACSDSSNTGSISEANAQYYQQEASKLRAQIGNLQNQNRNMLGESLAALSLR 138
Query: 122 QLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
L+ LE+ +++ ++ + K+E EI ++++E++L NQ L+
Sbjct: 139 DLKNLEQKIEKGISKIRSKKNELLFAEIEYMQKREIDLHNNNQYLR 184
>AG_LYCES (Q40168) Floral homeotic protein AGAMOUS (TAG1)
Length = 248
Score = 110 bits (275), Expect = 3e-24
Identities = 60/166 (36%), Positives = 101/166 (60%), Gaps = 1/166 (0%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS- 61
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL+VFS +L+EYA++S
Sbjct: 19 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVVFSNRGRLYEYANNSV 78
Query: 62 RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLH 121
+ + A S + + + Q+ LR ++ ++ R + GE L G+ L
Sbjct: 79 KATIERYKKACSDSSNTGSVSEANAQYYQQEASKLRAQIGNLMNQNRNMMGEALAGMKLK 138
Query: 122 QLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE+ +++ ++ + K+E EI ++++EV+L NQ L+
Sbjct: 139 ELKNLEQRIEKGISKIRSKKNELLFAEIEYMQKREVDLHNNNQYLR 184
>AGL3_ARATH (P29383) Agamous-like MADS box protein AGL3
Length = 258
Score = 110 bits (275), Expect = 3e-24
Identities = 63/172 (36%), Positives = 96/172 (55%), Gaps = 2/172 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R ++++K+I+N +RQVTF+KRR GL KKA ELS LCDA+IAL++FS KL+E+ SS
Sbjct: 1 MGRGKVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEIALLIFSNRGKLYEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSND--TLRKKVEEKTHELRQLNGEDLQGL 118
G + + P + D L+ +VE H R L GE+L +
Sbjct: 61 PSGMARTVDKYRKHSYATMDPNQSAKDLQDKYQDYLKLKSRVEILQHSQRHLLGEELSEM 120
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
+++L+ LE + SL + K + ++S LK KE L+E N+ L++ L
Sbjct: 121 DVNELEHLERQVDASLRQIRSTKARSMLDQLSDLKTKEEMLLETNRDLRRKL 172
>AG11_ARATH (Q38836) Agamous-like MADS box protein AGL11
Length = 230
Score = 110 bits (274), Expect = 4e-24
Identities = 61/168 (36%), Positives = 104/168 (61%), Gaps = 1/168 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS +L+EYA++
Sbjct: 1 MGRGKIEIKRIENSTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYANN 60
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
+ R + A S + + +T + + LR++++ + R L G+ L L+
Sbjct: 61 NIRSTIERYKKACSDSTNTSTVQEINAAYYQQESAKLRQQIQTIQNSNRNLMGDSLSSLS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+ +L+++E L+++++ + K E + EI +++E+EL EN L+
Sbjct: 121 VKELKQVENRLEKAISRIRSKKHELLLVEIENAQKREIELDNENIYLR 168
>CAL_ARATH (Q39081) Transcription factor CAULIFLOWER (Agamous-like
MADS box protein AGL10)
Length = 253
Score = 109 bits (273), Expect = 5e-24
Identities = 70/172 (40%), Positives = 100/172 (57%), Gaps = 3/172 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTFSKRR GL KKAQE+S LCDA+++LIVFS KLFEY+S
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRTGLLKKAQEISVLCDAEVSLIVFSHKGKLFEYSSE 60
Query: 61 S--RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGL 118
S L + A P S S L+ K+E R GE+L+ +
Sbjct: 61 SCMEKVLERYERYSYAERQLIAPDSHVNTNWSMEYSRLKAKIELLERNQRHYLGEELEPM 120
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL-KQV 169
+L LQ LE+ L+ +L + K++ + ++ L+RKE E+ EEN L KQ+
Sbjct: 121 SLKDLQNLEQQLETALKHIRSRKNQLMNESLNHLQRKEKEIQEENSMLTKQI 172
>AG_PETHY (Q40885) Floral homeotic protein AGAMOUS (pMADS3)
Length = 242
Score = 109 bits (273), Expect = 5e-24
Identities = 60/166 (36%), Positives = 99/166 (59%), Gaps = 1/166 (0%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS- 61
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+ +L+EYA++S
Sbjct: 19 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYANNSV 78
Query: 62 RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLH 121
+ + A S + + + Q+ LR ++ ++ R GE L L L
Sbjct: 79 KATIERYKKACSDSSNTGSIAEANAQYYQQEASKLRAQIGNLQNQNRNFLGESLAALNLR 138
Query: 122 QLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
L+ LE+ +++ ++ + K+E EI ++++E++L NQ L+
Sbjct: 139 DLRNLEQKIEKGISKIRAKKNELLFAEIEYMQKREIDLHNNNQYLR 184
>AGL8_SOLCO (O22328) Agamous-like MADS box protein AGL8 homolog
Length = 250
Score = 109 bits (273), Expect = 5e-24
Identities = 67/175 (38%), Positives = 99/175 (56%), Gaps = 12/175 (6%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+K+I+N +RQVTFSKRR GL KKA E+S LCDA++ LIVFS KLFEYA+
Sbjct: 1 MGRGRVQLKRIENKINRQVTFSKRRSGLLKKAHEISVLCDAEVGLIVFSTKGKLFEYATD 60
Query: 61 S-------RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGE 113
S R + D T+P S T++ N L+ ++E + GE
Sbjct: 61 SCMERLLERYERYSFAEKQLVPTDHTSPGSWTLE-----NAKLKARLEVLQRNEKLYVGE 115
Query: 114 DLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
DL+ L + +LQ LE L +L + K++ + IS L++++ L E+N +L +
Sbjct: 116 DLESLNMKELQNLEHQLASALKHIRSRKNQLMHESISVLQKQDRALQEQNNQLSK 170
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.136 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,591,372
Number of Sequences: 164201
Number of extensions: 939742
Number of successful extensions: 4234
Number of sequences better than 10.0: 176
Number of HSP's better than 10.0 without gapping: 87
Number of HSP's successfully gapped in prelim test: 89
Number of HSP's that attempted gapping in prelim test: 4020
Number of HSP's gapped (non-prelim): 260
length of query: 244
length of database: 59,974,054
effective HSP length: 107
effective length of query: 137
effective length of database: 42,404,547
effective search space: 5809422939
effective search space used: 5809422939
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0100a.3


