

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.11
(245 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
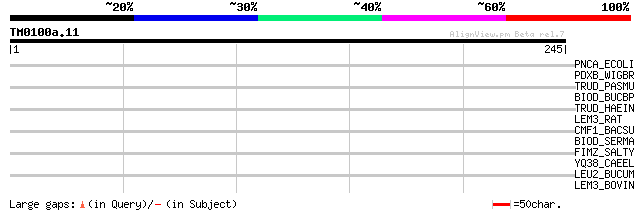
Score E
Sequences producing significant alignments: (bits) Value
PNCA_ECOLI (P21369) Pyrazinamidase/nicotinamidase (EC 3.5.1.-) (... 37 0.033
PDXB_WIGBR (Q8D2P6) Erythronate-4-phosphate dehydrogenase (EC 1.... 37 0.044
TRUD_PASMU (Q9CKK4) tRNA pseudouridine synthase D (EC 4.2.1.70) ... 34 0.37
BIOD_BUCBP (P59564) Dethiobiotin synthetase (EC 6.3.3.3) (Dethio... 33 0.63
TRUD_HAEIN (P44039) tRNA pseudouridine synthase D (EC 4.2.1.70) ... 31 2.4
LEM3_RAT (P98106) P-selectin precursor (Granule membrane protein... 31 3.1
CMF1_BACSU (P39145) ComF operon protein 1 31 3.1
BIOD_SERMA (P36572) Dethiobiotin synthetase (EC 6.3.3.3) (Dethio... 31 3.1
FIMZ_SALTY (P26319) Fimbriae Z protein 30 5.3
YQ38_CAEEL (Q09459) Hypothetical protein C09G5.8 in chromosome II 30 7.0
LEU2_BUCUM (Q9EVG8) 3-isopropylmalate dehydratase large subunit ... 30 7.0
LEM3_BOVIN (P42201) P-selectin precursor (Granule membrane prote... 30 7.0
>PNCA_ECOLI (P21369) Pyrazinamidase/nicotinamidase (EC 3.5.1.-) (EC
3.5.1.19) (PZAase) (Nicotine deamidase) (NAMase)
Length = 213
Score = 37.4 bits (85), Expect = 0.033
Identities = 38/158 (24%), Positives = 60/158 (37%), Gaps = 28/158 (17%)
Query: 30 LVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDSHHPNK- 88
L+LVD+ N FC GA LA E + ++ + RL + V+A D H N
Sbjct: 6 LLLVDLQNDFCAGGA--LAVPEGD----STVDVANRLIDWCQSRGEAVIASQDWHPANHG 59
Query: 89 -------------------PEDPYPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDG 129
P+ +P HC+ ++ + L P L D
Sbjct: 60 SFASQHGVEPYTPGQLDGLPQTFWPDHCVQNSEGAQLHPLLHQKAIAAVFHKGENPLVDS 119
Query: 130 YIGSAEEDG--SNVFVDWVKKNKIRTLLVVGICTDICV 165
Y + DW++ ++I L+V+G+ TD CV
Sbjct: 120 YSAFFDNGRRQKTSLDDWLRDHEIDELIVMGLATDYCV 157
>PDXB_WIGBR (Q8D2P6) Erythronate-4-phosphate dehydrogenase (EC
1.1.1.-)
Length = 378
Score = 37.0 bits (84), Expect = 0.044
Identities = 19/44 (43%), Positives = 26/44 (58%), Gaps = 1/44 (2%)
Query: 130 YIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTM 173
+IGSA +V VDW+KKNKI G C + V ++V S+M
Sbjct: 61 FIGSATSGKDHVDVDWLKKNKINFDFAPG-CNSVAVAEYVFSSM 103
>TRUD_PASMU (Q9CKK4) tRNA pseudouridine synthase D (EC 4.2.1.70)
(Pseudouridylate synthase) (Uracil hydrolyase)
Length = 335
Score = 33.9 bits (76), Expect = 0.37
Identities = 18/44 (40%), Positives = 24/44 (53%), Gaps = 2/44 (4%)
Query: 100 GTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSNVFV 143
G D NL ALRW + E NV R+K F Y+ +A + N+ V
Sbjct: 162 GRDGHNLTQALRWAQGEINVKDRKKRSF--YLSAARSEVFNLVV 203
>BIOD_BUCBP (P59564) Dethiobiotin synthetase (EC 6.3.3.3)
(Dethiobiotin synthase) (DTB synthetase) (DTBS)
Length = 227
Score = 33.1 bits (74), Expect = 0.63
Identities = 41/146 (28%), Positives = 62/146 (42%), Gaps = 26/146 (17%)
Query: 26 TVNGLVLVDIIN--GFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDS 83
TV+ ++L+++ G+ T G +A + N A L R F L S
Sbjct: 17 TVSSIILLELARRYGYNTSGYKPIAAGCRTNDFKQY-NSDAVLLRKFSTVKL-------S 68
Query: 84 HHPNKPEDPYPPHCIAGTDESN--------LVPALRWLENETNVTLRRKECFDGYIGSAE 135
+ P P C T+E N L L+ L+ ++N L +G IG
Sbjct: 69 YQEVNPYLFIDPVCPFFTNEKNHMNVCMNKLSSGLQRLKKKSNWIL-----IEG-IGGLH 122
Query: 136 EDGSNVFV--DWVKKNKIRTLLVVGI 159
SN FV DW+KK ++T+LV+GI
Sbjct: 123 TPFSNEFVMSDWIKKENLKTILVIGI 148
>TRUD_HAEIN (P44039) tRNA pseudouridine synthase D (EC 4.2.1.70)
(Pseudouridylate synthase) (Uracil hydrolyase)
Length = 339
Score = 31.2 bits (69), Expect = 2.4
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 2/44 (4%)
Query: 100 GTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSNVFV 143
G D NL ALRW + E V R+K F Y+ +A + N+ V
Sbjct: 166 GRDGHNLTQALRWAQGEIKVKDRKKRSF--YLSAARSEIFNLVV 207
>LEM3_RAT (P98106) P-selectin precursor (Granule membrane protein
140) (GMP-140) (PADGEM) (CD62P antigen)
(Leukocyte-endothelial cell adhesion molecule 3)
(LECAM3)
Length = 768
Score = 30.8 bits (68), Expect = 3.1
Identities = 28/113 (24%), Positives = 43/113 (37%), Gaps = 11/113 (9%)
Query: 100 GTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGI 159
GT+++ A W +NE N ++C + YI S G + D + R L
Sbjct: 105 GTNKTLTAEAENWADNEPNNKRNNQDCVEIYIKSNSAPGK--WNDEPCFKRKRALCYTAS 162
Query: 160 CTDICV------LDFVCSTMSAKNRGFLKPLENVVVYSHGCATFDVPLEVARN 206
C D+ ++ + S + GF P Y C FD+P V N
Sbjct: 163 CQDMSCNSQGERIETIGSYTCSCYPGFYGP---ECEYVQECGKFDIPQHVLMN 212
>CMF1_BACSU (P39145) ComF operon protein 1
Length = 463
Score = 30.8 bits (68), Expect = 3.1
Identities = 18/81 (22%), Positives = 34/81 (41%), Gaps = 2/81 (2%)
Query: 80 FLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGS 139
+ +H + Y C+ S VP W E + K +DG + S ++ +
Sbjct: 70 YFSFYHSSGKNKLYCRSCVMMGRVSEEVPLYSWKEENESNWKSIKLTWDGKLSSGQQKAA 129
Query: 140 NVFVDWVKKNKIRTLLVVGIC 160
NV ++ + K + LL+ +C
Sbjct: 130 NVLIEAISKKE--ELLIWAVC 148
>BIOD_SERMA (P36572) Dethiobiotin synthetase (EC 6.3.3.3)
(Dethiobiotin synthase) (DTB synthetase) (DTBS)
Length = 226
Score = 30.8 bits (68), Expect = 3.1
Identities = 31/109 (28%), Positives = 46/109 (41%), Gaps = 10/109 (9%)
Query: 57 SEMINESARLA-RLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGT-----DESNLVPAL 110
SEM E R L + N V D +P +P PH ++ D + L L
Sbjct: 43 SEMTAEGLRNGDALALQANSGVALDYDEVNPYVFAEPTSPHIVSADEGRPIDAARLSDGL 102
Query: 111 RWLENETNVTLRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGI 159
R LE + L E G+ + + F DWV++ ++ +LVVGI
Sbjct: 103 RRLEQRADWVL--VEGAGGWFTPLSAEYT--FADWVRQEQLPVILVVGI 147
>FIMZ_SALTY (P26319) Fimbriae Z protein
Length = 210
Score = 30.0 bits (66), Expect = 5.3
Identities = 30/141 (21%), Positives = 56/141 (39%), Gaps = 10/141 (7%)
Query: 98 IAGTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSNVFVDWVKK-----NKIR 152
+ GTD L+ ++ ++ T + + Y G A G+N FV K N ++
Sbjct: 59 LPGTDGFTLLKRIKSIQEHTRILFLSSKSEAFYAGRAIRAGANGFVSKRKDLNDIYNAVK 118
Query: 153 TLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVV-----YSHGCATFDVPLEVARNT 207
+L L+F+ +T + K PL N V ++G + ++ ++ +
Sbjct: 119 MILSGYSFFPSETLNFISNTRTPKGGHHDMPLSNREVTVLRYLANGMSNKEIAEQLLLSN 178
Query: 208 KGALAHPQEFLHHMGLYMAKE 228
K AH +GL+ E
Sbjct: 179 KTISAHKANIFSKLGLHSIVE 199
>YQ38_CAEEL (Q09459) Hypothetical protein C09G5.8 in chromosome II
Length = 1531
Score = 29.6 bits (65), Expect = 7.0
Identities = 22/73 (30%), Positives = 35/73 (47%), Gaps = 3/73 (4%)
Query: 55 QISEMIN-ESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWL 113
++SEM + ES R + EK P S H ++P PP I E L+ L+
Sbjct: 409 EMSEMSDDESGRSTPVIEEKKKPRRKSRKSSHQEPSKNPIPPPRIPDQTEKVLLDKLKVA 468
Query: 114 ENETNVTLRRKEC 126
EN+ + + ++EC
Sbjct: 469 END--LAMLQEEC 479
>LEU2_BUCUM (Q9EVG8) 3-isopropylmalate dehydratase large subunit (EC
4.2.1.33) (Isopropylmalate isomerase) (Alpha-IPM
isomerase) (IPMI) (Fragment)
Length = 444
Score = 29.6 bits (65), Expect = 7.0
Identities = 36/142 (25%), Positives = 58/142 (40%), Gaps = 31/142 (21%)
Query: 69 LFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNV--------- 119
LF EK+L FL S ++ + CI D SNL P + W N V
Sbjct: 254 LFWEKSLDYWKFLKSD-----KNAHFDKCIT-LDISNLAPQITWGTNPDQVISIDEKIPD 307
Query: 120 -----TLRRKECFDG---YIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCS 171
+L +KE Y+G SN ++ + +++ +G CT+ + D +
Sbjct: 308 YNNINSLLKKESAKSACEYMGLK----SNTYLTNISIDRV----FIGSCTNARIEDLRAA 359
Query: 172 TMSAKNRGFLKPLENVVVYSHG 193
+ KNR K ++ +VV G
Sbjct: 360 SKILKNRKIAKHVKAIVVPGSG 381
>LEM3_BOVIN (P42201) P-selectin precursor (Granule membrane protein
140) (GMP-140) (PADGEM) (CD62P antigen)
(Leukocyte-endothelial cell adhesion molecule 3)
(LECAM3)
Length = 646
Score = 29.6 bits (65), Expect = 7.0
Identities = 32/125 (25%), Positives = 45/125 (35%), Gaps = 21/125 (16%)
Query: 100 GTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGS-NVFVDWVKKNKIRTLLVVG 158
GT ++ A W +NE N ++C + YI S G N W +K R L
Sbjct: 105 GTKKTLTEEAENWADNEPNNKRNNQDCVEIYIKSLSAPGKWNDEPCWKRK---RALCYRA 161
Query: 159 ICTDI----------CVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATFDVPLEVARNTK 208
C D+ + ++ CS GF P Y C FD+P V N
Sbjct: 162 SCQDMSCSKQGECIETIGNYTCSCYP----GFYGP---ECEYVRECGEFDLPQHVHMNCS 214
Query: 209 GALAH 213
L +
Sbjct: 215 HPLGN 219
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,187,438
Number of Sequences: 164201
Number of extensions: 1205080
Number of successful extensions: 2683
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2677
Number of HSP's gapped (non-prelim): 12
length of query: 245
length of database: 59,974,054
effective HSP length: 107
effective length of query: 138
effective length of database: 42,404,547
effective search space: 5851827486
effective search space used: 5851827486
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0100a.11


