

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0096b.3
(223 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
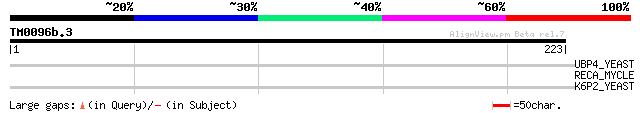
Score E
Sequences producing significant alignments: (bits) Value
UBP4_YEAST (P32571) Ubiquitin carboxyl-terminal hydrolase 4 (EC ... 34 0.31
RECA_MYCLE (P35901) RecA protein (Recombinase A) [Contains: Mle ... 29 7.7
K6P2_YEAST (P16862) 6-phosphofructokinase beta subunit (EC 2.7.1... 29 7.7
>UBP4_YEAST (P32571) Ubiquitin carboxyl-terminal hydrolase 4 (EC
3.1.2.15) (Ubiquitin thiolesterase 4)
(Ubiquitin-specific processing protease 4)
(Deubiquitinating enzyme 4) (Vacuole biogenesis protein
SSV7)
Length = 926
Score = 33.9 bits (76), Expect = 0.31
Identities = 17/37 (45%), Positives = 23/37 (61%), Gaps = 4/37 (10%)
Query: 28 VVDDVVYHLQLLSGNTNGLSIQTRGMLSPHIHHRMQD 64
+ DD VY ++GNT+GLS+Q +SP I H M D
Sbjct: 338 IKDDSVY----INGNTSGLSLQHLPKMSPSIRHSMDD 370
>RECA_MYCLE (P35901) RecA protein (Recombinase A) [Contains: Mle
recA intein]
Length = 711
Score = 29.3 bits (64), Expect = 7.7
Identities = 18/63 (28%), Positives = 33/63 (51%), Gaps = 1/63 (1%)
Query: 48 IQTRGMLSPHIHHRMQDVGALVELDPGTPVPEVLALDSSLILPDALKFVS-VMTKAIRDL 106
+Q + L +I H +++ TP+PE++ L ++ L D KF+S KA+ L
Sbjct: 355 LQWKKALMGNIRHTVRENSMGASFIDFTPLPELVELQRAVYLGDGKKFLSEEYLKALTPL 414
Query: 107 IIA 109
++A
Sbjct: 415 VLA 417
>K6P2_YEAST (P16862) 6-phosphofructokinase beta subunit (EC
2.7.1.11) (Phosphofructokinase 2) (Phosphohexokinase)
(6PF-1-K beta subunit)
Length = 959
Score = 29.3 bits (64), Expect = 7.7
Identities = 18/64 (28%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 62 MQDVGALVELDPGTPVPEVLALDSSLILPDALKFVSVMTKAIRDLIIAKQVGVVFDGTDE 121
++ V A++E P TP P + ++ ++ ++ V +TKA+ + I AK D
Sbjct: 501 LEAVNAVLESTPDTPSPLIAVNENKIVRKPLMESVK-LTKAVAEAIQAKDFKRAMSLRDT 559
Query: 122 EAIE 125
E IE
Sbjct: 560 EFIE 563
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.142 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,547,062
Number of Sequences: 164201
Number of extensions: 1105025
Number of successful extensions: 2448
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 2447
Number of HSP's gapped (non-prelim): 3
length of query: 223
length of database: 59,974,054
effective HSP length: 106
effective length of query: 117
effective length of database: 42,568,748
effective search space: 4980543516
effective search space used: 4980543516
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0096b.3


