

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0095c.6
(384 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
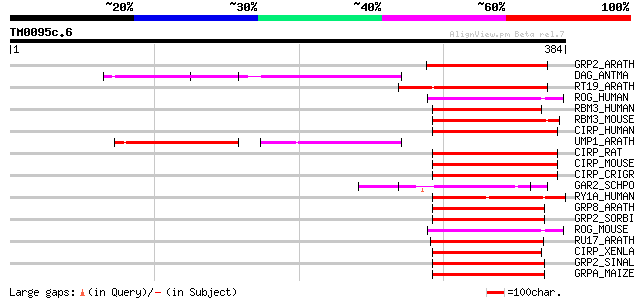
Score E
Sequences producing significant alignments: (bits) Value
GRP2_ARATH (Q9SVM8) Glycine-rich RNA-binding protein 2, mitochon... 75 3e-13
DAG_ANTMA (Q38732) DAG protein, chloroplast precursor 73 1e-12
RT19_ARATH (P39697) 40S ribosomal protein S19, mitochondrial pre... 72 2e-12
ROG_HUMAN (P38159) Heterogeneous nuclear ribonucleoprotein G (hn... 66 2e-10
RBM3_HUMAN (P98179) Putative RNA-binding protein 3 (RNA binding ... 66 2e-10
RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding ... 65 2e-10
CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein (Glycine-... 65 2e-10
UMP1_ARATH (Q9LKA5) Unknown mitochondrial protein At3g15000 65 3e-10
CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-ri... 65 3e-10
CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein (Glycine-... 65 3e-10
CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein (Glycine-... 65 3e-10
GAR2_SCHPO (P41891) Protein gar2 65 4e-10
RY1A_HUMAN (Q15414) RNA binding motif protein, Y chromosome, fam... 64 6e-10
GRP8_ARATH (Q03251) Glycine-rich RNA-binding protein 8 (CCR1 pro... 63 1e-09
GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2 63 1e-09
ROG_MOUSE (O35479) Heterogeneous nuclear ribonucleoprotein G (hn... 63 1e-09
RU17_ARATH (Q42404) U1 small nuclear ribonucleoprotein 70 kDa (U... 62 2e-09
CIRP_XENLA (O93235) Cold-inducible RNA-binding protein (Glycine-... 62 2e-09
GRP2_SINAL (P49311) Glycine-rich RNA-binding protein GRP2A 62 2e-09
GRPA_MAIZE (P10979) Glycine-rich RNA-binding, abscisic acid-indu... 61 4e-09
>GRP2_ARATH (Q9SVM8) Glycine-rich RNA-binding protein 2,
mitochondrial precursor (AtGRP2)
Length = 158
Score = 75.1 bits (183), Expect = 3e-13
Identities = 38/84 (45%), Positives = 56/84 (66%)
Query: 289 LRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGA 348
L + KLF+ GLS+ T + +LR AF FG++V KVI+D+ + RS+G+ F+ + E AA A
Sbjct: 32 LMSTKLFIGGLSWGTDDASLRDAFAHFGDVVDAKVIVDRETGRSRGFGFVNFNDEGAATA 91
Query: 349 ALKEMNGKIINGWMIVVDVAKTNP 372
A+ EM+GK +NG I V+ A P
Sbjct: 92 AISEMDGKELNGRHIRVNPANDRP 115
>DAG_ANTMA (Q38732) DAG protein, chloroplast precursor
Length = 230
Score = 72.8 bits (177), Expect = 1e-12
Identities = 38/93 (40%), Positives = 55/93 (58%), Gaps = 1/93 (1%)
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
P +N HW+++M+ P ++ Q+ID Y+ TL TV+GS +EA+ +Y S T+ GF
Sbjct: 82 PGCDYN-HWLIVMEFPKDPAPTREQMIDTYLNTLATVLGSMEEAKKNMYAFSTTTYTGFQ 140
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
C + EE S + LPGVL V PD + KDY
Sbjct: 141 CTVTEETSEKFKGLPGVLWVLPDSYIDVKNKDY 173
Score = 72.0 bits (175), Expect = 2e-12
Identities = 42/146 (28%), Positives = 76/146 (51%), Gaps = 9/146 (6%)
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSK 185
C I S+ A+ P V ++ D +++S + + + +SG+ R ++ P +
Sbjct: 37 CTRPHPIKSRSAAYPTVRALT-DGEYSSRRNNNNNNSGEE--------RETIMLPGCDYN 87
Query: 186 HWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDC 245
HWL+ M+ P T+ Q++D Y L V+G+ ++A+ +Y S T GF C + E+
Sbjct: 88 HWLIVMEFPKDPAPTREQMIDTYLNTLATVLGSMEEAKKNMYAFSTTTYTGFQCTVTEET 147
Query: 246 AQDLAGVPGVSSVQPDDNFESENKNY 271
++ G+PGV V PD + +NK+Y
Sbjct: 148 SEKFKGLPGVLWVLPDSYIDVKNKDY 173
>RT19_ARATH (P39697) 40S ribosomal protein S19, mitochondrial
precursor
Length = 212
Score = 72.4 bits (176), Expect = 2e-12
Identities = 37/103 (35%), Positives = 66/103 (63%), Gaps = 1/103 (0%)
Query: 270 NYEDSLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKIS 329
+++ ++VP SS + TK L++ GLS T E +L+ AF F + + +V+ +K++
Sbjct: 10 HWKQGVNVPVSSMLGSLRYMSTK-LYIGGLSPGTDEHSLKDAFSSFNGVTEARVMTNKVT 68
Query: 330 KRSKGYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNP 372
RS+GY F+ + +E++A +A+ MNG+ +NG+ I V+VAK P
Sbjct: 69 GRSRGYGFVNFISEDSANSAISAMNGQELNGFNISVNVAKDWP 111
>ROG_HUMAN (P38159) Heterogeneous nuclear ribonucleoprotein G (hnRNP
G) (RNA binding motif protein, X chromosome)
(Glycoprotein p43)
Length = 391
Score = 65.9 bits (159), Expect = 2e-10
Identities = 35/94 (37%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 290 RTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAA 349
R KLF+ GL+ T+EK L A F +G +V+V ++ D+ + +S+G+AF+ + + A A
Sbjct: 6 RPGKLFIGGLNTETNEKALEAVFGKYGRIVEVLLMKDRETNKSRGFAFVTFESPADAKDA 65
Query: 350 LKEMNGKIINGWMIVVDVAKTNPPRFNRGHARPP 383
++MNGK ++G I V+ A P F G PP
Sbjct: 66 ARDMNGKSLDGKAIKVEQA--TKPSFESGRRGPP 97
>RBM3_HUMAN (P98179) Putative RNA-binding protein 3 (RNA binding
motif protein 3) (RNPL)
Length = 157
Score = 65.9 bits (159), Expect = 2e-10
Identities = 33/76 (43%), Positives = 49/76 (64%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GL+F T E+ L F FG + +V V+ D+ ++RS+G+ FI +T E A A++
Sbjct: 7 KLFVGGLNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHASVAMRA 66
Query: 353 MNGKIINGWMIVVDVA 368
MNG+ ++G I VD A
Sbjct: 67 MNGESLDGRQIRVDHA 82
>RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding
motif protein 3)
Length = 153
Score = 65.5 bits (158), Expect = 2e-10
Identities = 37/88 (42%), Positives = 55/88 (62%), Gaps = 1/88 (1%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GL+F T E+ L F FG + +V V+ D+ ++RS+G+ FI +T E A A++
Sbjct: 7 KLFVGGLNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHASDAMRA 66
Query: 353 MNGKIINGWMIVVDVAKTNPPRFNRGHA 380
MNG+ ++G I VD A + R +RG A
Sbjct: 67 MNGESLDGRQIRVDHAGKS-ARGSRGGA 93
>CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 65.5 bits (158), Expect = 2e-10
Identities = 33/87 (37%), Positives = 55/87 (62%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GLSF T+E++L F +G++ +V V+ D+ ++RS+G+ F+ + + A A+
Sbjct: 7 KLFVGGLSFDTNEQSLEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 353 MNGKIINGWMIVVDVAKTNPPRFNRGH 379
MNGK ++G I VD A + +RG+
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGY 93
>UMP1_ARATH (Q9LKA5) Unknown mitochondrial protein At3g15000
Length = 395
Score = 65.1 bits (157), Expect = 3e-10
Identities = 32/86 (37%), Positives = 52/86 (60%), Gaps = 1/86 (1%)
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+V+++ PP G ++ ++ID Y+KTL ++GSE EA+M +Y S ++ F + E++
Sbjct: 99 HWLVVVE-PPQGEPTRDEIIDSYIKTLAQIVGSEDEARMKIYSVSTRCYYAFGALVSEDL 157
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S +L L V V PD + KDY
Sbjct: 158 SHKLKELSNVRWVLPDSYLDVRNKDY 183
Score = 57.0 bits (136), Expect = 8e-08
Identities = 31/98 (31%), Positives = 54/98 (54%), Gaps = 1/98 (1%)
Query: 174 RTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKT 233
+ +L + +HWLV ++ P E T+ +I+D Y + L +++G+E +A+M IY VS +
Sbjct: 87 KETILLDGCDFEHWLVVVEPPQGEP-TRDEIIDSYIKTLAQIVGSEDEARMKIYSVSTRC 145
Query: 234 NFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
+ F + ED + L + V V PD + NK+Y
Sbjct: 146 YYAFGALVSEDLSHKLKELSNVRWVLPDSYLDVRNKDY 183
>CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-rich
RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 65.1 bits (157), Expect = 3e-10
Identities = 33/87 (37%), Positives = 54/87 (61%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GLSF T+E+ L F +G++ +V V+ D+ ++RS+G+ F+ + + A A+
Sbjct: 7 KLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 353 MNGKIINGWMIVVDVAKTNPPRFNRGH 379
MNGK ++G I VD A + +RG+
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGY 93
>CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 65.1 bits (157), Expect = 3e-10
Identities = 33/87 (37%), Positives = 54/87 (61%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GLSF T+E+ L F +G++ +V V+ D+ ++RS+G+ F+ + + A A+
Sbjct: 7 KLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 353 MNGKIINGWMIVVDVAKTNPPRFNRGH 379
MNGK ++G I VD A + +RG+
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGY 93
>CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 65.1 bits (157), Expect = 3e-10
Identities = 33/87 (37%), Positives = 54/87 (61%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLFV GLSF T+E+ L F +G++ +V V+ D+ ++RS+G+ F+ + + A A+
Sbjct: 7 KLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 353 MNGKIINGWMIVVDVAKTNPPRFNRGH 379
MNGK ++G I VD A + +RG+
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGY 93
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 64.7 bits (156), Expect = 4e-10
Identities = 42/137 (30%), Positives = 69/137 (49%), Gaps = 6/137 (4%)
Query: 242 DEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVPNSSEAS------QEASLRTKKLF 295
D + + D + G S D ES +++ + + P S E + S T +F
Sbjct: 207 DSESSSDSSSESGDSDSSSDSESESSSEDEKKRKAEPASEERPAKITKPSQDSNETCTVF 266
Query: 296 VTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNG 355
V LS+ ++ L FE +G +V +VI+D S RSKGY ++++ T EAA AA+
Sbjct: 267 VGRLSWNVDDQWLGQEFEEYGTIVGARVIMDGQSGRSKGYGYVDFETPEAAKAAVAANGT 326
Query: 356 KIINGWMIVVDVAKTNP 372
K I+G M+ +D++ P
Sbjct: 327 KEIDGRMVNLDLSNPRP 343
Score = 44.3 bits (103), Expect = 5e-04
Identities = 28/91 (30%), Positives = 46/91 (49%), Gaps = 13/91 (14%)
Query: 270 NYEDSLSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKIS 329
N+ D LS P+ + +FV LSF +E L AF G G++ +++ D S
Sbjct: 356 NFGDQLSEPSDT------------VFVGNLSFNATEDDLSTAFGGCGDIQSIRLPTDPQS 403
Query: 330 KRSKGYAFIEYTTEEAAGAALKEMNGKIING 360
R KG+ ++ ++ ++A + EMNG I G
Sbjct: 404 GRLKGFGYVTFSDIDSAKKCV-EMNGHFIAG 433
>RY1A_HUMAN (Q15414) RNA binding motif protein, Y chromosome, family
1 member A1 (RNA-binding motif protein 1)
Length = 496
Score = 63.9 bits (154), Expect = 6e-10
Identities = 40/92 (43%), Positives = 56/92 (60%), Gaps = 2/92 (2%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLF+ GL+ T+EK L+A F G + +V +I D+ SK S+G+AFI + A A K+
Sbjct: 9 KLFIGGLNRETNEKMLKAVFGKHGPISEVLLIKDRTSK-SRGFAFITFENPADAKNAAKD 67
Query: 353 MNGKIINGWMIVVDVAKTNPPRFNRGHARPPA 384
MNGK ++G I V+ AK P + G RPPA
Sbjct: 68 MNGKSLHGKAIKVEQAK-KPSFQSGGRRRPPA 98
>GRP8_ARATH (Q03251) Glycine-rich RNA-binding protein 8 (CCR1
protein)
Length = 169
Score = 63.2 bits (152), Expect = 1e-09
Identities = 29/78 (37%), Positives = 54/78 (69%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
+ FV GL++ T+++ L+ F FG+++ K+I D+ S RS+G+ F+ + E+A A++E
Sbjct: 7 RCFVGGLAWATNDEDLQRTFSQFGDVIDSKIINDRESGRSRGFGFVTFKDEKAMRDAIEE 66
Query: 353 MNGKIINGWMIVVDVAKT 370
MNGK ++G +I V+ A++
Sbjct: 67 MNGKELDGRVITVNEAQS 84
>GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2
Length = 168
Score = 63.2 bits (152), Expect = 1e-09
Identities = 29/78 (37%), Positives = 54/78 (69%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
+ FV GL++ T+ +TL AF FG+++ KVI D+ + RS+G+ F+ +++E++ A++
Sbjct: 9 RCFVGGLAWATNNETLEQAFANFGQVIDSKVITDRETGRSRGFGFVTFSSEQSMLDAIEN 68
Query: 353 MNGKIINGWMIVVDVAKT 370
MNGK ++G I V+ A++
Sbjct: 69 MNGKELDGRNITVNQAQS 86
>ROG_MOUSE (O35479) Heterogeneous nuclear ribonucleoprotein G (hnRNP
G) (RNA binding motif protein, X chromosome)
Length = 388
Score = 62.8 bits (151), Expect = 1e-09
Identities = 34/94 (36%), Positives = 55/94 (58%), Gaps = 2/94 (2%)
Query: 290 RTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAA 349
R KLF+ GL+ T+EK L A F +G +V++ ++ D+ + +S+G+AF+ + A A
Sbjct: 6 RPGKLFIGGLNTETNEKALEAVFGKYGRIVEIILMKDRETNKSRGFAFVTLESPADAKDA 65
Query: 350 LKEMNGKIINGWMIVVDVAKTNPPRFNRGHARPP 383
++MNGK ++G I V+ A P F G PP
Sbjct: 66 ARDMNGKSLDGKAIKVEQA--TKPFFESGRRGPP 97
>RU17_ARATH (Q42404) U1 small nuclear ribonucleoprotein 70 kDa (U1
snRNP 70 kDa) (snRNP70) (U1-70K)
Length = 427
Score = 62.4 bits (150), Expect = 2e-09
Identities = 29/78 (37%), Positives = 51/78 (65%)
Query: 292 KKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALK 351
K LFV+ L++ +SE ++ FE +G + +V ++ D+++ + KGYAFIEY AA K
Sbjct: 138 KTLFVSRLNYESSESKIKREFESYGPIKRVHLVTDQLTNKPKGYAFIEYMHTRDMKAAYK 197
Query: 352 EMNGKIINGWMIVVDVAK 369
+ +G+ I+G ++VDV +
Sbjct: 198 QADGQKIDGRRVLVDVER 215
>CIRP_XENLA (O93235) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (XCIRP)
Length = 163
Score = 62.4 bits (150), Expect = 2e-09
Identities = 31/76 (40%), Positives = 48/76 (62%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
KLF+ GL+F T+E L AF +G + +V V+ D+ +KRS+G+ F+ + + A A+
Sbjct: 6 KLFIGGLNFETNEDCLEQAFTKYGRISEVVVVKDRETKRSRGFGFVTFENVDDAKDAMMA 65
Query: 353 MNGKIINGWMIVVDVA 368
MNGK ++G I VD A
Sbjct: 66 MNGKSVDGRQIRVDQA 81
>GRP2_SINAL (P49311) Glycine-rich RNA-binding protein GRP2A
Length = 169
Score = 62.0 bits (149), Expect = 2e-09
Identities = 30/78 (38%), Positives = 52/78 (66%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
+ FV GL++ T E++L AF FGELV K+I D+ + RS+G+ F+ + E++ A++
Sbjct: 9 RCFVGGLAWATDERSLETAFSQFGELVDSKIINDRETGRSRGFGFVTFKDEKSMKDAIEG 68
Query: 353 MNGKIINGWMIVVDVAKT 370
MNG+ ++G I V+ A++
Sbjct: 69 MNGQDLDGRSITVNEAQS 86
>GRPA_MAIZE (P10979) Glycine-rich RNA-binding, abscisic
acid-inducible protein
Length = 157
Score = 61.2 bits (147), Expect = 4e-09
Identities = 29/78 (37%), Positives = 53/78 (67%)
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
+ FV GL++ TS ++L AF +GE++ KVI D+ + RS+G+ F+ +++E + A++
Sbjct: 9 RCFVGGLAWATSNESLENAFASYGEILDSKVITDRETGRSRGFGFVTFSSENSMLDAIEN 68
Query: 353 MNGKIINGWMIVVDVAKT 370
MNGK ++G I V+ A++
Sbjct: 69 MNGKELDGRNITVNQAQS 86
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.130 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,655,017
Number of Sequences: 164201
Number of extensions: 1854418
Number of successful extensions: 5485
Number of sequences better than 10.0: 306
Number of HSP's better than 10.0 without gapping: 248
Number of HSP's successfully gapped in prelim test: 58
Number of HSP's that attempted gapping in prelim test: 4926
Number of HSP's gapped (non-prelim): 529
length of query: 384
length of database: 59,974,054
effective HSP length: 112
effective length of query: 272
effective length of database: 41,583,542
effective search space: 11310723424
effective search space used: 11310723424
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0095c.6


