

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094b.3
(337 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
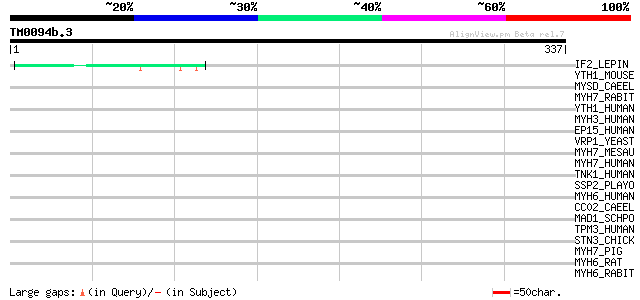
Score E
Sequences producing significant alignments: (bits) Value
IF2_LEPIN (Q8F7K1) Translation initiation factor IF-2 44 6e-04
YTH1_MOUSE (P59326) YTH domain protein 1 (Dermatomyositis associ... 43 0.001
MYSD_CAEEL (P02567) Myosin heavy chain D (MHC D) 42 0.002
MYH7_RABIT (P04461) Myosin heavy chain, cardiac muscle beta isof... 42 0.002
YTH1_HUMAN (Q9BYJ9) YTH domain protein 1 (Dermatomyositis associ... 42 0.002
MYH3_HUMAN (P11055) Myosin heavy chain, fast skeletal muscle, em... 41 0.004
EP15_HUMAN (P42566) Epidermal growth factor receptor substrate 1... 41 0.004
VRP1_YEAST (P37370) Verprolin 41 0.005
MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isof... 41 0.005
MYH7_HUMAN (P12883) Myosin heavy chain, cardiac muscle beta isof... 41 0.005
TNK1_HUMAN (O95271) Tankyrase 1 (EC 2.4.2.30) (TANK1) (Tankyrase... 40 0.006
SSP2_PLAYO (Q01443) Sporozoite surface protein 2 precursor 40 0.006
MYH6_HUMAN (P13533) Myosin heavy chain, cardiac muscle alpha iso... 40 0.006
CC02_CAEEL (P17656) Cuticle collagen 2 precursor 40 0.006
MAD1_SCHPO (P87169) Spindle assembly checkpoint component mad1 (... 40 0.008
TPM3_HUMAN (P06753) Tropomyosin alpha 3 chain (Tropomyosin 3) (T... 40 0.011
STN3_CHICK (O93388) Stathmin 3 (SCG10-like protein) (Neuroplasti... 40 0.011
MYH7_PIG (P79293) Myosin heavy chain, cardiac muscle beta isofor... 40 0.011
MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isofo... 40 0.011
MYH6_RABIT (P04460) Myosin heavy chain, cardiac muscle alpha iso... 40 0.011
>IF2_LEPIN (Q8F7K1) Translation initiation factor IF-2
Length = 880
Score = 43.9 bits (102), Expect = 6e-04
Identities = 40/143 (27%), Positives = 56/143 (38%), Gaps = 34/143 (23%)
Query: 4 KPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLAS 63
K + + G S+P A SPK+ V E DP + P V P P+ DS P+
Sbjct: 39 KKLIIKKKGDDPSTPSPAASPKKETVAESDPSSKPP-------VMPLPLPDDSGQSPIVR 91
Query: 64 DHPSEVNASQTPDAP------RPVTLAEGQRSPSSIRPPIPSPVK------------VTL 105
PS + ++ ++P RP + ++ S PP SP +
Sbjct: 92 PAPSSHSPAKREESPGKQDAGRPPRDKDTRQGGGSSYPPSRSPFQKEDSNIIVSRPIQRT 151
Query: 106 GPSAPNS---------PGVGGEG 119
GPS PNS PG GG G
Sbjct: 152 GPSRPNSGGGYQGNRGPGQGGGG 174
>YTH1_MOUSE (P59326) YTH domain protein 1 (Dermatomyositis
associated with cancer putative autoantigen-1 homolog)
(DACA-1 homolog)
Length = 559
Score = 42.7 bits (99), Expect = 0.001
Identities = 44/163 (26%), Positives = 67/163 (40%), Gaps = 13/163 (7%)
Query: 11 GGKRQSSPVS---ACSPKRSKVGEPDPQTTPLTGT-GGGFVAPDPVVTDSACDPLASDHP 66
GG + PVS + + SK +P P+ +G GG + P P+ + + P
Sbjct: 220 GGTNVNMPVSKPTSWAAIASKPAKPQPKMKTKSGPIVGGALPPPPIKHNMDIGTWDNKGP 279
Query: 67 S-EVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPG-VGGEGENLSA 124
+ + +A Q +P+ + P ++PP P P P P V N +
Sbjct: 280 APKASAPQQTPSPQAAPQPQQVAQPLPVQPP-PLVQPQYQSPQQPLQPRWVAPRNRNAAF 338
Query: 125 AQPTGPSSSS------PPYPFSFLLSHPYLEKLKGAPSQNPQD 161
Q G +S S P + SHP LEKLK A S NP++
Sbjct: 339 GQSGGANSDSNSVGNAQPTSAPSVESHPVLEKLKAAHSYNPKE 381
>MYSD_CAEEL (P02567) Myosin heavy chain D (MHC D)
Length = 1938
Score = 42.4 bits (98), Expect = 0.002
Identities = 27/91 (29%), Positives = 50/91 (54%), Gaps = 7/91 (7%)
Query: 194 RQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKLL 253
+QEVE L++ ++A + +A + AAK+ +I L++E+ Q+E + + + KLL
Sbjct: 951 QQEVENLKKSIEAVDGNLAKSLEEKAAKENQIHSLQDEMNSQDETIGKINKEK----KLL 1006
Query: 254 ADYEAQAATLKEKIRSEAAHAAAKERVRGSL 284
+ Q L + +++E A A R+RG L
Sbjct: 1007 EENNRQ---LVDDLQAEEAKQAQANRLRGKL 1034
Score = 33.1 bits (74), Expect = 1.0
Identities = 26/88 (29%), Positives = 47/88 (52%), Gaps = 2/88 (2%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L QE +ELQ LD E + ++E + E+ +R E+E + + E + + T K
Sbjct: 1540 LEQEKDELQHALDEAEAALEAEESKVLRLQIEVQQIRSEIEKRIQE-KEEEFENTRKNHQ 1598
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERV 280
A E+ A+L+ + +S+A A AK+++
Sbjct: 1599 RA-LESIQASLETEAKSKAELARAKKKL 1625
Score = 32.7 bits (73), Expect = 1.3
Identities = 20/89 (22%), Positives = 40/89 (44%), Gaps = 7/89 (7%)
Query: 196 EVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKLLAD 255
++ L LDA + + A EK KE+ D+ +++ + ++ E + +L
Sbjct: 1206 QISALTNTLDALQKSKAKIEKEKGVLQKELDDINAQVDQETKSRVEQE-------RLAKQ 1258
Query: 256 YEAQAATLKEKIRSEAAHAAAKERVRGSL 284
YE Q A L++K+ ++ +G L
Sbjct: 1259 YEIQVAELQQKVDEQSRQIGEYTSTKGRL 1287
>MYH7_RABIT (P04461) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta) (Beta isomyosin) (Fragment)
Length = 736
Score = 42.4 bits (98), Expect = 0.002
Identities = 26/68 (38%), Positives = 39/68 (57%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL+RD+D E+T+A EK A + ++ +L EE+ G +E +A+ LT K K
Sbjct: 507 LEDECSELKRDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAK----LTKKKKA 562
Query: 253 LADYEAQA 260
L + QA
Sbjct: 563 LQEAHQQA 570
Score = 30.0 bits (66), Expect = 8.6
Identities = 39/168 (23%), Positives = 64/168 (37%), Gaps = 22/168 (13%)
Query: 191 VGLRQEVEELQRDLDAKEITVASQEK----------FLAAKDKEIADLREELEGQNEALA 240
V L QE +LQ + A++ +A E+ L AK KE+ + E+ E N L
Sbjct: 442 VSLLQEKNDLQLQVQAEQDNLADAEERCDQLIKNKIQLEAKVKEMNERLEDEEEMNAELT 501
Query: 241 EAQSDLTFKCK-----------LLADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAKAQ 289
+ L +C LA E + + K+++ A + + L K +
Sbjct: 502 AKKRKLEDECSELKRDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAKLTKKKK 561
Query: 290 VKHLHPTINLDPLGAFKRVTPAGLEGPQDPPEVADVDLGDAVEQEKKI 337
LD L A + + + D DL ++EQEKK+
Sbjct: 562 ALQEAHQQALDDLQAEEDKVNTLTKAKVKLEQQVD-DLEGSLEQEKKV 608
>YTH1_HUMAN (Q9BYJ9) YTH domain protein 1 (Dermatomyositis
associated with cancer putative autoantigen-1) (DACA-1)
Length = 559
Score = 42.0 bits (97), Expect = 0.002
Identities = 45/164 (27%), Positives = 66/164 (39%), Gaps = 15/164 (9%)
Query: 11 GGKRQSSPVS---ACSPKRSKVGEPDPQTTPLTG-TGGGFVAPDPVVTDSACDPLASDHP 66
GG + PVS + + SK +P P+ +G GG + P P+ + D D+
Sbjct: 220 GGTNVNMPVSKPTSWAAIASKPAKPQPKMKTKSGPVMGGGLPPPPIKHNM--DIGTWDNK 277
Query: 67 SEVNASQTPD-APRPVTLAEGQR--SPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLS 123
V + P AP P + Q+ P +PP + + P + V N +
Sbjct: 278 GPVPKAPVPQQAPSPQAAPQPQQVAQPLPAQPPALAQPQYQSPQQPPQTRWVAPRNRNAA 337
Query: 124 AAQPTGPSSSS------PPYPFSFLLSHPYLEKLKGAPSQNPQD 161
Q G S S P + SHP LEKLK A S NP++
Sbjct: 338 FGQSGGAGSDSNSPGNVQPNSAPSVESHPVLEKLKAAHSYNPKE 381
>MYH3_HUMAN (P11055) Myosin heavy chain, fast skeletal muscle,
embryonic (Muscle embryonic myosin heavy chain) (SMHCE)
Length = 1940
Score = 41.2 bits (95), Expect = 0.004
Identities = 25/68 (36%), Positives = 39/68 (56%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL++D+D E+T+A EK A + ++ +L EEL G +E +A+ LT + K
Sbjct: 944 LEDECSELKKDIDDLELTLAKVEKEKHATENKVKNLTEELSGLDETIAK----LTREKKA 999
Query: 253 LADYEAQA 260
L + QA
Sbjct: 1000 LQEAHQQA 1007
Score = 32.7 bits (73), Expect = 1.3
Identities = 23/71 (32%), Positives = 36/71 (50%), Gaps = 4/71 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNE---ALAEAQSDLTFK 249
L+ E EL R L+ KE V+ + A ++ +L+ +LE +N+ ALA A
Sbjct: 1281 LQTEAGELSRQLEEKESIVSQLSRSKQAFTQQTEELKRQLEEENKAKNALAHALQSSRHD 1340
Query: 250 CKLLAD-YEAQ 259
C LL + YE +
Sbjct: 1341 CDLLREQYEEE 1351
Score = 30.0 bits (66), Expect = 8.6
Identities = 21/75 (28%), Positives = 37/75 (49%), Gaps = 7/75 (9%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L+ +EEL+ +++A+ T A EK + +D ELE +E L EA + + +L
Sbjct: 1112 LQARIEELEEEIEAERATRAKTEK-------QRSDYARELEELSERLEEAGGVTSTQIEL 1164
Query: 253 LADYEAQAATLKEKI 267
EA+ L+ +
Sbjct: 1165 NKKREAEFLKLRRDL 1179
>EP15_HUMAN (P42566) Epidermal growth factor receptor substrate 15
(Protein Eps15) (AF-1p protein)
Length = 896
Score = 41.2 bits (95), Expect = 0.004
Identities = 24/95 (25%), Positives = 51/95 (53%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E+ +LQR+ + E + +E + + E+ DL++E++ +N L + Q+ +L
Sbjct: 338 LNNEIVDLQREKNNVEQDLKEKEDTIKQRTSEVQDLQDEVQRENTNLQKLQAQKQQVQEL 397
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
L + + Q A L+E+++ A + ++ SL A+
Sbjct: 398 LDELDEQKAQLEEQLKEVRKKCAEEAQLISSLKAE 432
>VRP1_YEAST (P37370) Verprolin
Length = 817
Score = 40.8 bits (94), Expect = 0.005
Identities = 35/120 (29%), Positives = 49/120 (40%), Gaps = 15/120 (12%)
Query: 28 KVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHP---------SEVNASQTPDAP 78
K E + ++ P+ G G A T S+ P S P ++ A P
Sbjct: 45 KKAETNDRSAPIVGGGVVSSASGSSGTVSSKGPSMSAPPIPGMGAPQLGDILAGGIPKLK 104
Query: 79 RPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
A + SPS+ PPIP V P PN+P + + A P+ PSSS+PP P
Sbjct: 105 HINNNASTKPSPSASAPPIPGAVPSVAAPPIPNAP------LSPAPAVPSIPSSSAPPIP 158
Score = 34.3 bits (77), Expect = 0.46
Identities = 35/147 (23%), Positives = 51/147 (33%), Gaps = 30/147 (20%)
Query: 5 PRVAEVGGKR--QSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLA 62
P +AE+ +R + + S K P PL + P + S PL
Sbjct: 274 PFLAEINARRSERGAVEGVSSTKIQTENHKSPSQPPLPSSA-------PPIPTSHAPPLP 326
Query: 63 SDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPS-------------PVKVTLGPSA 109
P + AP+ T A P+ PP+P+ PV TL P
Sbjct: 327 PTAPPPPSLPNVTSAPKKATSA-----PAPPPPPLPAAMSSASTNSVKATPVPPTLAPPL 381
Query: 110 PNSPGVGGEGENLSAAQPTGPSSSSPP 136
PN+ V N +++ P P PP
Sbjct: 382 PNTTSV---PPNKASSMPAPPPPPPPP 405
Score = 30.4 bits (67), Expect = 6.6
Identities = 34/148 (22%), Positives = 48/148 (31%), Gaps = 18/148 (12%)
Query: 5 PRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASD 64
P V K S+P P + + + T P P T + +
Sbjct: 336 PNVTSAPKKATSAPAPPPPPLPAAMSSASTNSVKATPVPPTLAPPLPNTTS-----VPPN 390
Query: 65 HPSEVNASQTPDAPRPVTLAEGQR-SPSSIR----PPIPSPVKVTLGPSAP--------N 111
S + A P P P + S SSI PP P P T PSAP N
Sbjct: 391 KASSMPAPPPPPPPPPGAFSTSSALSASSIPLAPLPPPPPPSVATSVPSAPPPPPTLTTN 450
Query: 112 SPGVGGEGENLSAAQPTGPSSSSPPYPF 139
P + +S++ + + P PF
Sbjct: 451 KPSASSKQSKISSSSSSSAVTPGGPLPF 478
>MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1934
Score = 40.8 bits (94), Expect = 0.005
Identities = 25/68 (36%), Positives = 39/68 (56%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL+RD+D E+T+A EK A + ++ +L EE+ G +E +A+ LT + K
Sbjct: 942 LEDECSELKRDIDDLELTLAKVEKDKHATENKVKNLTEEMAGLDEIIAK----LTKEKKA 997
Query: 253 LADYEAQA 260
L + QA
Sbjct: 998 LQEAHQQA 1005
Score = 32.0 bits (71), Expect = 2.3
Identities = 27/99 (27%), Positives = 43/99 (43%), Gaps = 11/99 (11%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA----QSDLTF 248
L+ +EEL+ +L+A+ A EK + DL ELE +E L EA +
Sbjct: 1110 LQARIEELEEELEAERTARAKVEKLRS-------DLSRELEEISERLEEAGGATSVQIEM 1162
Query: 249 KCKLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
K A+++ L+E A AAA + +A+
Sbjct: 1163 NKKREAEFQKMRRDLEEATLQHEATAAALRKKHADSVAE 1201
Score = 32.0 bits (71), Expect = 2.3
Identities = 26/95 (27%), Positives = 44/95 (45%), Gaps = 6/95 (6%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEA---LAEAQSDLTFK 249
L+ E EL R LD KE ++ + +++ DL+ +LE + +A LA A
Sbjct: 1279 LQTENGELSRQLDEKEALISQLTRGKLTYTQQLEDLKRQLEEEVKAKNTLAHALQSARHD 1338
Query: 250 CKLLADY---EAQAATLKEKIRSEAAHAAAKERVR 281
C LL + E +A + + S+A A+ R +
Sbjct: 1339 CDLLREQYEEETEAKAELQCVLSKANSEVAQWRTK 1373
>MYH7_HUMAN (P12883) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1935
Score = 40.8 bits (94), Expect = 0.005
Identities = 25/68 (36%), Positives = 39/68 (56%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL+RD+D E+T+A EK A + ++ +L EE+ G +E +A+ LT + K
Sbjct: 943 LEDECSELKRDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAK----LTKEKKA 998
Query: 253 LADYEAQA 260
L + QA
Sbjct: 999 LQEAHQQA 1006
Score = 34.7 bits (78), Expect = 0.35
Identities = 26/95 (27%), Positives = 44/95 (45%), Gaps = 6/95 (6%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQ---NEALAEAQSDLTFK 249
L+ E EL R LD KE ++ + +++ DL+ +LE + ALA A
Sbjct: 1280 LQTENGELSRQLDEKEALISQLTRGKLTYTQQLEDLKRQLEEEVKAKNALAHALQSARHD 1339
Query: 250 CKLLADY---EAQAATLKEKIRSEAAHAAAKERVR 281
C LL + E +A +++ S+A A+ R +
Sbjct: 1340 CDLLREQYEEETEAKAELQRVLSKANSEVAQWRTK 1374
Score = 32.3 bits (72), Expect = 1.7
Identities = 26/99 (26%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA----QSDLTF 248
L+++++ELQ ++ E + S+ A +K +DL ELE +E L EA +
Sbjct: 1104 LQKKLKELQARIEELEEELESERTARAKVEKLRSDLSRELEEISERLEEAGGATSVQIEM 1163
Query: 249 KCKLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
K A+++ L+E A AAA + +A+
Sbjct: 1164 NKKREAEFQKMRRDLEEATLQHEATAAALRKKHADSVAE 1202
>TNK1_HUMAN (O95271) Tankyrase 1 (EC 2.4.2.30) (TANK1) (Tankyrase I)
(TNKS-1) (TRF1-interacting ankyrin-related ADP-ribose
polymerase)
Length = 1327
Score = 40.4 bits (93), Expect = 0.006
Identities = 38/129 (29%), Positives = 47/129 (35%), Gaps = 16/129 (12%)
Query: 32 PDPQTTPLT-GTGGGFVAPDPVVTDSACDPLASDH-----PSEVNASQTPDAPRPVTLAE 85
P P PL+ G G P T S P AS P + PD PR +
Sbjct: 28 PPPPPPPLSPGLAPGTTPASP--TASGLAPFASPRHGLALPEGDGSRDPPDRPRSPDPVD 85
Query: 86 GQRSPSSIR--------PPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPY 137
G S+ P +P+ + APN G G S++ PT SSSSP
Sbjct: 86 GTSCCSTTSTICTVAAAPVVPAVSTSSAAGVAPNPAGSGSNNSPSSSSSPTSSSSSSPSS 145
Query: 138 PFSFLLSHP 146
P S L P
Sbjct: 146 PGSSLAESP 154
>SSP2_PLAYO (Q01443) Sporozoite surface protein 2 precursor
Length = 827
Score = 40.4 bits (93), Expect = 0.006
Identities = 32/134 (23%), Positives = 47/134 (34%), Gaps = 3/134 (2%)
Query: 4 KPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLAS 63
KP E + +P +P + EP P P+P + +P
Sbjct: 414 KPNPNEPSNPNKPNPNEPSNPNKPNPNEPSNPNKPNPNEPSNPNKPNPNEPLNPNEPSNP 473
Query: 64 DHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPI-PSPVKVTLGPSAPNSPGVGGEGENL 122
+ PS NA P+ P PS+ P P+ PS PN P E L
Sbjct: 474 NEPSNPNAPSNPNEPSNPNEPSNPNEPSNPNEPSNPNEPSNPKKPSNPNEP--SNPNEPL 531
Query: 123 SAAQPTGPSSSSPP 136
+ +P+ P+ S P
Sbjct: 532 NPNEPSNPNEPSNP 545
Score = 40.0 bits (92), Expect = 0.008
Identities = 34/152 (22%), Positives = 61/152 (39%), Gaps = 19/152 (12%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
S+P +PK K P+ + P +P+ + +P +P+E + + P
Sbjct: 507 SNPNEPSNPK--KPSNPNEPSNP----------NEPLNPNEPSNPNEPSNPNEPSNPEEP 554
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
P+ + +P P PS K P P +P E L+ +P+ P S+P
Sbjct: 555 SNPKEPSNPNEPSNPEEPNPEEPSNPKEPSNPEEPINP------EELNPKEPSNPEESNP 608
Query: 136 PYPFSFLLSHPYLEKLKGAPSQNPQDVAQEAV 167
P + S+P E + ++NP + E +
Sbjct: 609 KEPINPEESNP-KEPINPEDNENPLIIQDEPI 639
Score = 38.1 bits (87), Expect = 0.032
Identities = 24/81 (29%), Positives = 37/81 (45%), Gaps = 2/81 (2%)
Query: 59 DPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLG-PSAPNSPGVGG 117
+P ++P+ N P+ P P +R+P+ +P P+P K PS PN P
Sbjct: 371 NPNNPNNPNNPNDPSNPNNPNPKKRNPKRRNPNKPKPNKPNPNKPNPNEPSNPNKPN-PN 429
Query: 118 EGENLSAAQPTGPSSSSPPYP 138
E N + P PS+ + P P
Sbjct: 430 EPSNPNKPNPNEPSNPNKPNP 450
>MYH6_HUMAN (P13533) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1939
Score = 40.4 bits (93), Expect = 0.006
Identities = 26/89 (29%), Positives = 49/89 (54%), Gaps = 5/89 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL++D+D E+T+A EK A + ++ +L EE+ G +E +A+ LT + K
Sbjct: 945 LEDECSELKKDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAK----LTKEKKA 1000
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERVR 281
L + QA ++ + ++ +K +V+
Sbjct: 1001 LQEAHQQALD-DLQVEEDKVNSLSKSKVK 1028
Score = 33.1 bits (74), Expect = 1.0
Identities = 25/95 (26%), Positives = 46/95 (48%), Gaps = 6/95 (6%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNE---ALAEAQSDLTFK 249
L+ E EL R L+ KE ++ + + +++ DL+ +LE + + ALA A
Sbjct: 1282 LQTENGELARQLEEKEALISQLTRGKLSYTQQMEDLKRQLEEEGKAKNALAHALQSARHD 1341
Query: 250 CKLLADY---EAQAATLKEKIRSEAAHAAAKERVR 281
C LL + E +A +++ S+A A+ R +
Sbjct: 1342 CDLLREQYEEETEAKAELQRVLSKANSEVAQWRTK 1376
Score = 30.4 bits (67), Expect = 6.6
Identities = 26/95 (27%), Positives = 41/95 (42%), Gaps = 11/95 (11%)
Query: 197 VEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA----QSDLTFKCKL 252
+EEL+ +L+A+ A EK + DL ELE +E L EA + K
Sbjct: 1117 IEELEEELEAERTARAKVEKLRS-------DLSRELEEISERLEEAGGATSVQIEMNKKR 1169
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
A+++ L+E A AAA + +A+
Sbjct: 1170 EAEFQKMRRDLEEATLQHEATAAALRKKHADSVAE 1204
>CC02_CAEEL (P17656) Cuticle collagen 2 precursor
Length = 301
Score = 40.4 bits (93), Expect = 0.006
Identities = 33/115 (28%), Positives = 37/115 (31%), Gaps = 2/115 (1%)
Query: 25 KRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLA 84
K K G+P P G G AP VT C P P + P P P
Sbjct: 115 KPGKPGKPGAPGAP-GNPGKGASAPCEPVTQPPCQPCPGGPPGPAGPAGPPGPPGPDGNP 173
Query: 85 EGQRSPSSIRPP-IPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
PS P P P AP +PG GE P P + PP P
Sbjct: 174 GSPAGPSGPGPAGPPGPAGPAGNDGAPGAPGGPGEPGASEQGGPGEPGPAGPPGP 228
>MAD1_SCHPO (P87169) Spindle assembly checkpoint component mad1
(Mitotic arrest deficient protein 1)
Length = 689
Score = 40.0 bits (92), Expect = 0.008
Identities = 35/139 (25%), Positives = 72/139 (51%), Gaps = 27/139 (19%)
Query: 194 RQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQ---------- 243
++E+EE++ + +A ++ + + L AK KEIA+L+ ++E ++AL+E
Sbjct: 118 QKEIEEVRNEKEATQVKI---HELLDAKWKEIAELKTQIEKNDQALSEKNHEVMVSNQAL 174
Query: 244 ----SDLTFKCKLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAKAQ----VKHLHP 295
++LT KL AD Q T +++ AAA+++++ + Q +K +
Sbjct: 175 QMKDTNLTNLEKLFADSREQLETKCKEL------AAAEQQLQELSVHNQQLEESIKQVSS 228
Query: 296 TINLDPLGAFKRVTPAGLE 314
+I L+ + A +R+ + LE
Sbjct: 229 SIELEKINAEQRLQISELE 247
Score = 30.4 bits (67), Expect = 6.6
Identities = 24/78 (30%), Positives = 41/78 (51%), Gaps = 10/78 (12%)
Query: 199 ELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQN---EALAEAQSDLTFKCKLLAD 255
EL++ + + ++ EK AA+++ I E+L N E L E ++DL K +
Sbjct: 231 ELEKINAEQRLQISELEKLKAAQEERI----EKLSSNNRNVEILKEEKNDLESKLYRFEE 286
Query: 256 YEAQAATLK---EKIRSE 270
Y + ATL+ EKI++E
Sbjct: 287 YRDKVATLELENEKIQTE 304
>TPM3_HUMAN (P06753) Tropomyosin alpha 3 chain (Tropomyosin 3)
(Tropomyosin gamma) (hTM5)
Length = 284
Score = 39.7 bits (91), Expect = 0.011
Identities = 30/90 (33%), Positives = 48/90 (53%), Gaps = 6/90 (6%)
Query: 196 EVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKLLAD 255
E E+ Q + +K++ E LAA K++ +EL+ +EAL +AQ L K AD
Sbjct: 26 EAEQKQAEERSKQL-----EDELAAMQKKLKGTEDELDKYSEALKDAQEKLELAEKKAAD 80
Query: 256 YEAQAATLKEKIR-SEAAHAAAKERVRGSL 284
EA+ A+L +I+ E A+ER+ +L
Sbjct: 81 AEAEVASLNRRIQLVEEELDRAQERLATAL 110
>STN3_CHICK (O93388) Stathmin 3 (SCG10-like protein)
(Neuroplasticin-2)
Length = 180
Score = 39.7 bits (91), Expect = 0.011
Identities = 33/89 (37%), Positives = 46/89 (51%), Gaps = 6/89 (6%)
Query: 197 VEELQRDLDAKEITVASQE----KFLAAK-DKEIADLREELEGQNEALAEAQSDLTFKCK 251
+EELQR L+A E +QE K LA K + E L + LE N A+ L +K +
Sbjct: 82 LEELQRRLEAAEERRKTQEAQVLKQLAEKREHEREVLHKALEENNNFSRLAEEKLNYKME 141
Query: 252 LLADY-EAQAATLKEKIRSEAAHAAAKER 279
L + EA A L+E++R + HAA R
Sbjct: 142 LSREIREAHLAALRERLREKELHAAEVRR 170
>MYH7_PIG (P79293) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1935
Score = 39.7 bits (91), Expect = 0.011
Identities = 24/68 (35%), Positives = 39/68 (57%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
+ E EL+RD+D E+T+A EK A + ++ +L EE+ G +E +A+ LT + K
Sbjct: 943 VEDECSELKRDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAK----LTKEKKA 998
Query: 253 LADYEAQA 260
L + QA
Sbjct: 999 LQEAHQQA 1006
Score = 32.0 bits (71), Expect = 2.3
Identities = 27/99 (27%), Positives = 43/99 (43%), Gaps = 11/99 (11%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA----QSDLTF 248
L+ +EEL+ +L+A+ A EK + DL ELE +E L EA +
Sbjct: 1111 LQARIEELEEELEAERTARAKVEKLRS-------DLSRELEEISERLEEAGGATSVQIEM 1163
Query: 249 KCKLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
K A+++ L+E A AAA + +A+
Sbjct: 1164 NKKREAEFQKMRRDLEEATLQHEATAAALRKKHADSVAE 1202
Score = 32.0 bits (71), Expect = 2.3
Identities = 27/95 (28%), Positives = 44/95 (45%), Gaps = 6/95 (6%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQ---NEALAEAQSDLTFK 249
L+ E EL R LD KE ++ + +++ DL+ +LE + ALA A
Sbjct: 1280 LQTENGELSRQLDEKEALISQLTRGKLTYTQQLEDLKRQLEEEVKAKNALAHALQSARHA 1339
Query: 250 CKLLAD-YEAQAATLKE--KIRSEAAHAAAKERVR 281
LL + YE + T E ++ S+A A+ R +
Sbjct: 1340 ADLLREQYEEETETKAELQRVLSKANSEVAQWRTK 1374
>MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1938
Score = 39.7 bits (91), Expect = 0.011
Identities = 24/68 (35%), Positives = 39/68 (57%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL++D+D E+T+A EK A + ++ +L EE+ G +E +A+ LT + K
Sbjct: 944 LEDECSELKKDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAK----LTKEKKA 999
Query: 253 LADYEAQA 260
L + QA
Sbjct: 1000 LQEAHQQA 1007
Score = 31.2 bits (69), Expect = 3.9
Identities = 25/95 (26%), Positives = 45/95 (47%), Gaps = 6/95 (6%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNE---ALAEAQSDLTFK 249
L+ E EL R L+ KE + + + +++ DL+ +LE + + ALA A
Sbjct: 1281 LQTENGELARQLEEKEALIWQLTRGKLSYTQQMEDLKRQLEEEGKAKNALAHALQSARHD 1340
Query: 250 CKLLADY---EAQAATLKEKIRSEAAHAAAKERVR 281
C LL + E +A +++ S+A A+ R +
Sbjct: 1341 CDLLREQYEEEMEAKAELQRVLSKANSEVAQWRTK 1375
Score = 30.4 bits (67), Expect = 6.6
Identities = 26/95 (27%), Positives = 41/95 (42%), Gaps = 11/95 (11%)
Query: 197 VEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA----QSDLTFKCKL 252
+EEL+ +L+A+ A EK + DL ELE +E L EA + K
Sbjct: 1116 IEELEEELEAERTARAKVEKLRS-------DLTRELEEISERLEEAGGATSVQIEMNKKR 1168
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
A+++ L+E A AAA + +A+
Sbjct: 1169 EAEFQKMRRDLEEATLQHEATAAALRKKHADSVAE 1203
>MYH6_RABIT (P04460) Myosin heavy chain, cardiac muscle alpha
isoform (MyHC-alpha) (Alpha isomyosin) (Fragment)
Length = 465
Score = 39.7 bits (91), Expect = 0.011
Identities = 24/68 (35%), Positives = 39/68 (57%), Gaps = 4/68 (5%)
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L E EL++D+D E+T+A EK A + ++ +L EE+ G +E +A+ LT + K
Sbjct: 140 LEDECSELKKDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAK----LTKEKKA 195
Query: 253 LADYEAQA 260
L + QA
Sbjct: 196 LQEAHQQA 203
Score = 30.4 bits (67), Expect = 6.6
Identities = 26/95 (27%), Positives = 41/95 (42%), Gaps = 11/95 (11%)
Query: 197 VEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA----QSDLTFKCKL 252
+EEL+ +L+A+ A EK + DL ELE +E L EA + K
Sbjct: 312 IEELEEELEAERTARAKVEKLRS-------DLSRELEEISERLEEAGGATSVQIEMNKKR 364
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAK 287
A+++ L+E A A+A R +A+
Sbjct: 365 EAEFQKMRRDLEEATLQHEATASALRRKHADSVAE 399
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,893,291
Number of Sequences: 164201
Number of extensions: 2043056
Number of successful extensions: 10442
Number of sequences better than 10.0: 659
Number of HSP's better than 10.0 without gapping: 145
Number of HSP's successfully gapped in prelim test: 525
Number of HSP's that attempted gapping in prelim test: 8714
Number of HSP's gapped (non-prelim): 1857
length of query: 337
length of database: 59,974,054
effective HSP length: 111
effective length of query: 226
effective length of database: 41,747,743
effective search space: 9434989918
effective search space used: 9434989918
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0094b.3


