

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
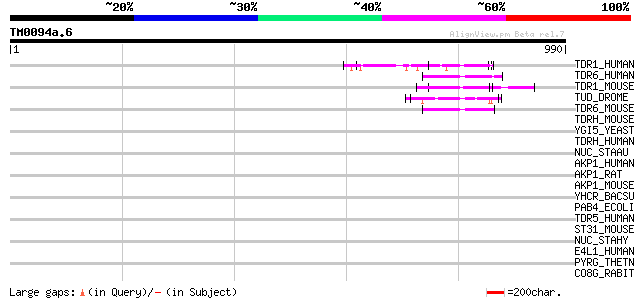
Query= TM0094a.6
(990 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TDR1_HUMAN (Q9BXT4) Tudor domain containing protein 1 63 4e-09
TDR6_HUMAN (O60522) Tudor domain containing protein 6 (Colon can... 56 4e-07
TDR1_MOUSE (Q99MV1) Tudor domain containing protein 1 54 3e-06
TUD_DROME (P25823) Maternal tudor protein 52 6e-06
TDR6_MOUSE (P61407) Tudor domain containing protein 6 48 1e-04
TDRH_MOUSE (Q80VL1) Tudor and KH domain containing protein 45 0.001
YGI5_YEAST (P53153) Hypothetical 32.1 kDa protein in MAD1-SCY1 i... 43 0.004
TDRH_HUMAN (Q9Y2W6) Tudor and KH domain containing protein 42 0.006
NUC_STAAU (P00644) Thermonuclease precursor (EC 3.1.31.1) (TNase... 42 0.006
AKP1_HUMAN (Q92667) A kinase anchor protein 1, mitochondrial pre... 41 0.014
AKP1_RAT (O88884) A kinase anchor protein 1, mitochondrial precu... 41 0.018
AKP1_MOUSE (O08715) A kinase anchor protein 1, mitochondrial pre... 41 0.018
YHCR_BACSU (P54602) Hypothetical protein yhcR precursor 38 0.12
PAB4_ECOLI (P22997) Protein parB 38 0.15
TDR5_HUMAN (Q8NAT2) Tudor domain containing protein 5 37 0.26
ST31_MOUSE (Q99MW1) Serine/threonine-protein kinase 31 (EC 2.7.1... 37 0.26
NUC_STAHY (P43270) Thermonuclease precursor (EC 3.1.31.1) (TNase... 35 0.99
E4L1_HUMAN (Q9H4G0) Band 4.1-like protein 1 (Neuronal protein 4.... 35 1.3
PYRG_THETN (Q8R720) CTP synthase (EC 6.3.4.2) (UTP--ammonia liga... 34 1.7
CO8G_RABIT (Q28679) Complement component C8 gamma chain precursor 34 1.7
>TDR1_HUMAN (Q9BXT4) Tudor domain containing protein 1
Length = 777
Score = 62.8 bits (151), Expect = 4e-09
Identities = 72/276 (26%), Positives = 124/276 (44%), Gaps = 36/276 (13%)
Query: 596 VLSGHRFKLLIPK----ETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVE 651
+LS R +IPK +I L+GV+ P G ++ EAI LM++ + + + ++V
Sbjct: 181 ILSLMRLCPIIPKLLELPMQAIKCVLAGVK-PSLGI-WTPEAICLMKKLVQNKIITVKVV 238
Query: 652 TVDRNGTFLGSLWESRT---NVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQK 708
N + + + +S T +V+ LL+AG A + S +D+ K+
Sbjct: 239 DKLENSSLVELIDKSETPHVSVSKVLLDAGFAVGEQSMVTDK----------PSDVKETS 288
Query: 709 LKIWENFVEGE-EVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGD---QKIASIQQQ 764
+ + VEG+ VE + + V+V + G+FY + + +K+ + +
Sbjct: 289 VPLG---VEGKVNPLEWTWVELGVDQTVDVVVCVIYSPGEFYCHVLKEDALKKLNDLNKS 345
Query: 765 LAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNT-PRGPVESPKDIFEVFYID 823
LA ++ P F + G +F D SWYRA+V P G V+ V ++D
Sbjct: 346 LAEHCQQKLP--NGFKAEIGQPCCAFFAGDGSWYRALVKEILPNGHVK-------VHFVD 396
Query: 824 YGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKS 859
YGN E+V +LR + + P C LA I+S
Sbjct: 397 YGNIEEVTADELRMISSTFLNLPFQGIRCQLADIQS 432
Score = 52.0 bits (123), Expect = 8e-06
Identities = 64/260 (24%), Positives = 107/260 (40%), Gaps = 51/260 (19%)
Query: 619 GVRCP-----GRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLWESRTNVALT 673
G+RC R + +SEEAI + + ++ V V NG V LT
Sbjct: 422 GIRCQLADIQSRNKHWSEEAITRFQMCVAGIKLQARVVEVTENGI----------GVELT 471
Query: 674 LLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAK---------KQKLKIWENFVEGEEVSNG 724
L ++ SD + + HL+ ++ K K +L++ V+G + ++
Sbjct: 472 DLSTCYPRII----SDVLIDEHLVLKSASPHKDLPNDRLVNKHELQV---HVQGLQATSS 524
Query: 725 AN----VESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGA-- 778
A +E + ++ V E++ FY G + Q++L L + A
Sbjct: 525 AEQWKTIELPVDKTIQANVLEIISPNLFYALPKG---MPENQEKLCMLTAELLEYCNAPK 581
Query: 779 ----FSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQ 834
+ P+ GD + +D WYRA+V+ T S D+ EV Y DYGN E + +
Sbjct: 582 SRPPYRPRIGDACCAKYTSDDFWYRAVVLGT------SDTDV-EVLYADYGNIETLPLCR 634
Query: 835 LRPLDQSVSAAPGLAQLCSL 854
++P+ S A P CSL
Sbjct: 635 VQPITSSHLALPFQIIRCSL 654
Score = 48.1 bits (113), Expect = 1e-04
Identities = 37/115 (32%), Positives = 53/115 (45%), Gaps = 8/115 (6%)
Query: 748 FYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPR 807
F Q +K+A +Q L+ + P F P GDI F D WYRA V+
Sbjct: 107 FCQQLQSGRKLAELQASLSKY-CDQLPPRSDFYPAIGDICCAQFSEDDQWYRASVL---- 161
Query: 808 GPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSPSL 862
+ ++ V Y+DYGN E ++ +L P+ + P A C LA +K PSL
Sbjct: 162 --AYASEESVLVGYVDYGNFEILSLMRLCPIIPKLLELPMQAIKCVLAGVK-PSL 213
>TDR6_HUMAN (O60522) Tudor domain containing protein 6 (Colon cancer
antigen NY-CO-45) (Fragment)
Length = 753
Score = 56.2 bits (134), Expect = 4e-07
Identities = 39/143 (27%), Positives = 70/143 (48%), Gaps = 10/143 (6%)
Query: 737 VIVTEVLGGGKFYVQTVGDQ-KIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADK 795
V V+ + FYVQ + D+ +I+ + ++L ++ + +G ++GD++ F D
Sbjct: 39 VYVSHINDLSDFYVQLIEDEAEISHLSERLNSVKTRPEYYVGP-PLQRGDMICAVFPEDN 97
Query: 796 SWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLA 855
WYRA++ + P D+ V +IDYGN V +++ LD + PGL CSL
Sbjct: 98 LWYRAVIKE------QQPNDLLSVQFIDYGNVSVVHTNKIGRLDLVNAILPGLCIHCSLQ 151
Query: 856 YIKSPSLEEDFGQEAAEYLSELT 878
+ P + ++ Y S+ T
Sbjct: 152 GFEVP--DNKNSKKMMHYFSQRT 172
>TDR1_MOUSE (Q99MV1) Tudor domain containing protein 1
Length = 928
Score = 53.5 bits (127), Expect = 3e-06
Identities = 34/132 (25%), Positives = 63/132 (46%), Gaps = 7/132 (5%)
Query: 726 NVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVL--GAFSPKK 783
+++ K+ +K VTE FY+Q + + ++ Q +L A V+ + P K
Sbjct: 7 SLQLKKTMEIKGTVTEFKHPSNFYIQLYSSEVLENMNQLSTSLKETYANVVPEDGYLPVK 66
Query: 784 GDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVS 843
G++ + + D++W RA+V V+ + V YIDYGN+E + + PL + +
Sbjct: 67 GEVCVAKYTVDQTWNRAIVQ-----AVDVLQRKAHVLYIDYGNEEMIPIDSVHPLSRGLD 121
Query: 844 AAPGLAQLCSLA 855
P A C ++
Sbjct: 122 LFPPSAIKCCVS 133
Score = 52.4 bits (124), Expect = 6e-06
Identities = 40/139 (28%), Positives = 62/139 (43%), Gaps = 13/139 (9%)
Query: 727 VESKQQEVLKVIVTEVLGGGKFYVQTVGD---QKIASIQQQLAALNLKEAPVLGAFSPKK 783
VE E + V+V + G+FY + D +K+ + Q LA ++ P F +
Sbjct: 458 VEFTVDETVDVVVCMMYSPGEFYCHFLKDDALEKLDDLNQSLADYCAQKPP--NGFKAEI 515
Query: 784 GDIVLCYFHADKSWYRAMVVNT-PRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSV 842
G +F D +WYRA+V P G V+ V ++DYGN E+V QL+ +
Sbjct: 516 GRPCCAFFSGDGNWYRALVKEILPSGNVK-------VHFVDYGNVEEVTTDQLQAILPQF 568
Query: 843 SAAPGLAQLCSLAYIKSPS 861
P C L I+ P+
Sbjct: 569 LLLPFQGMQCWLVDIQPPN 587
Score = 48.9 bits (115), Expect = 7e-05
Identities = 51/188 (27%), Positives = 80/188 (42%), Gaps = 20/188 (10%)
Query: 748 FYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPR 807
F Q K+A +Q+ L+ P F P GD+ F D WYRA V+
Sbjct: 261 FCQQLQSGHKLAELQESLSEYCGHVIP-RSDFYPTIGDVCCAQFSEDDQWYRASVL---- 315
Query: 808 GPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFG 867
+ ++ V Y+DYGN E ++ +L P+ + P A C LA +K PSL +
Sbjct: 316 --AYASEESVLVGYVDYGNFEILSLKRLCPIIPKLLDLPMQALNCVLAGVK-PSL-GIWT 371
Query: 868 QEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQ 927
EA + E+ V+ R + V GT ++ + +V +S + A++
Sbjct: 372 PEAVCVMKEM-----------VQNRMVTVRVVGMLGTRALVELIDKSVAPHVSASKALID 420
Query: 928 EGLARMEK 935
G A EK
Sbjct: 421 SGFAIKEK 428
Score = 45.1 bits (105), Expect = 0.001
Identities = 26/77 (33%), Positives = 37/77 (47%), Gaps = 7/77 (9%)
Query: 778 AFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRP 837
A+ P+ GD + D WYRA+V+ T V +V Y DYGN E + S+++P
Sbjct: 736 AYRPRTGDACCAKYTNDDFWYRAIVLETSESDV-------KVLYADYGNIETLPLSRVQP 788
Query: 838 LDQSVSAAPGLAQLCSL 854
+ S P CSL
Sbjct: 789 IPASHLELPFQIIRCSL 805
>TUD_DROME (P25823) Maternal tudor protein
Length = 2515
Score = 52.4 bits (124), Expect = 6e-06
Identities = 50/170 (29%), Positives = 78/170 (45%), Gaps = 24/170 (14%)
Query: 715 FVEGEEVSNG--ANVESKQQEVL----KVIVTEVLGGGKFYVQTVGDQKIASIQQQLAAL 768
+++GE+V+ A+ ++ E L ++ V G F++Q D K +L L
Sbjct: 1771 YIDGEDVAKKLIADGFARPLEYLASGCSCYISHVNGICDFFIQLERDSKAL----ELIEL 1826
Query: 769 NLKEAPVLGAFSP-KKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQ 827
L++ L +KG IV F D+ WYRA + E P +EV +IDYGN
Sbjct: 1827 YLRKKDTLKPLEGFEKGLIVAALFEDDELWYRAQLQK------ELPDSRYEVLFIDYGNT 1880
Query: 828 EQVAYSQLRPLDQSVSAAPGLAQLCSL----AYIK-SPSLEEDFGQEAAE 872
S+ L + +++ P L++ CSL AYI SP E F + E
Sbjct: 1881 STT--SKCLMLSEEIASLPSLSKKCSLQLPDAYISWSPEAEAKFAELTGE 1928
Score = 45.8 bits (107), Expect = 6e-04
Identities = 47/177 (26%), Positives = 74/177 (41%), Gaps = 16/177 (9%)
Query: 706 KQKLKIWENFVEGEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQL 765
K+ LKI+E+ + V + K+ E + I++ FYVQ + A + +
Sbjct: 2138 KRSLKIYEHLQK--LVQAELKLIQKRNENSECIISYGNSPKSFYVQMKHNS--ADLDLIV 2193
Query: 766 AAL-NLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDY 824
L +LK+ + P +CY D +YR + + V P FEVF +DY
Sbjct: 2194 KTLQSLKKEKLKKLIDPTTNSNGVCYSQEDACYYRCSIKS-----VLDPSQGFEVFLLDY 2248
Query: 825 GNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYI----KSPSLEEDFGQEAAEYLSEL 877
GN + ++ L Q + P LA C L+ I LEE F ++ EL
Sbjct: 2249 GN--TLVVPEVWQLPQEIEPIPSLALHCQLSKIPMDVSDEKLEEAFAALLEQHFGEL 2303
Score = 41.2 bits (95), Expect = 0.014
Identities = 33/100 (33%), Positives = 47/100 (47%), Gaps = 9/100 (9%)
Query: 780 SPKKGDI--VLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRP 837
S KK D+ + +D +WYRA + +S FEVFYIDYGN E++ ++
Sbjct: 1352 SLKKFDVGQICAVRSSDGNWYRARISGK-----DSNAACFEVFYIDYGNTEEIKRDDIKA 1406
Query: 838 LD-QSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSE 876
LD + A G A +L I PS + +E L E
Sbjct: 1407 LDAKFYEHASGFAVEINLP-IGRPSNDTKLKARISEILEE 1445
>TDR6_MOUSE (P61407) Tudor domain containing protein 6
Length = 2134
Score = 48.1 bits (113), Expect = 1e-04
Identities = 34/129 (26%), Positives = 59/129 (45%), Gaps = 8/129 (6%)
Query: 737 VIVTEVLGGGKFYVQTVGDQ-KIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADK 795
V V+ + FY+Q + D+ +I ++ ++L + + G + GD++ F D
Sbjct: 1315 VYVSHINDLSDFYIQLIEDEAEINNLSERLNDVRTRPQYHTGP-QWQSGDVICAVFPEDN 1373
Query: 796 SWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLA 855
WYRA+V+ + P + V +IDYGN V ++ L + P L CSL
Sbjct: 1374 LWYRALVME------QQPNGLLSVQFIDYGNMSVVHTNRTGRLGPVDAVLPALCLHCSLW 1427
Query: 856 YIKSPSLEE 864
+ P +E
Sbjct: 1428 GLSVPVCKE 1436
Score = 37.0 bits (84), Expect = 0.26
Identities = 21/61 (34%), Positives = 32/61 (52%), Gaps = 5/61 (8%)
Query: 794 DKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCS 853
D WYRA+++ T R P+ +V ++DYG +E V+ S LR L P + C+
Sbjct: 322 DGQWYRALLLETFR-----PQRCAQVLHVDYGRKELVSCSSLRYLLPEYFRMPVVTYPCA 376
Query: 854 L 854
L
Sbjct: 377 L 377
Score = 34.7 bits (78), Expect = 1.3
Identities = 28/95 (29%), Positives = 40/95 (41%), Gaps = 9/95 (9%)
Query: 785 DIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSA 844
D + C + +YRA V V+ VF +D GN E V + +R L
Sbjct: 546 DDLCCVKWKENGYYRATVTRLDSKSVD-------VFLVDRGNSENVDWCDVRMLLPQFRQ 598
Query: 845 APGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTL 879
P LA C+LA I L + + QEA + + L
Sbjct: 599 LPILALKCTLADIW--PLGKTWSQEATSFFKKTVL 631
>TDRH_MOUSE (Q80VL1) Tudor and KH domain containing protein
Length = 560
Score = 44.7 bits (104), Expect = 0.001
Identities = 38/146 (26%), Positives = 64/146 (43%), Gaps = 8/146 (5%)
Query: 733 EVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFH 792
E L+V V+ F++Q +G + + + E + + GDIV
Sbjct: 306 EYLEVYVSASEHPNHFWIQIIGSRSLQLDKLVSEMTQHYENSLPEDLTVHVGDIVAAPLS 365
Query: 793 ADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLC 852
+ SWYRA V+ T +E+ +++++D+G+ A LR L + P A C
Sbjct: 366 TNGSWYRARVLGT----LENGN--LDLYFVDFGDNGDCALKDLRALRSDFLSLPFQAIEC 419
Query: 853 SLAYIKSPSLEEDFGQEAAEYLSELT 878
SLA I E++ +EA + LT
Sbjct: 420 SLARIAPTG--EEWEEEALDEFDRLT 443
>YGI5_YEAST (P53153) Hypothetical 32.1 kDa protein in MAD1-SCY1
intergenic region
Length = 274
Score = 43.1 bits (100), Expect = 0.004
Identities = 24/93 (25%), Positives = 48/93 (50%), Gaps = 1/93 (1%)
Query: 271 ADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESA-KDLALELVENGYA 329
A PFG +A + + R+L + V + +D+++ + V Y D KDL+LE++++G A
Sbjct: 167 AQPFGNEALIWLQNRILGKKVWVKPLSIDQYNRCVARVSYWDWFGGWKDLSLEMLKDGLA 226
Query: 330 KYVEWSANMMEEEAKRRLKTAELEAKKSRLRMW 362
E N ++ + + + E A+ + +W
Sbjct: 227 VVYEGKVNTEFDDREDKYRYYEFLARSRKKGLW 259
>TDRH_HUMAN (Q9Y2W6) Tudor and KH domain containing protein
Length = 606
Score = 42.4 bits (98), Expect = 0.006
Identities = 40/146 (27%), Positives = 65/146 (44%), Gaps = 8/146 (5%)
Query: 733 EVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFH 792
E L+V V+ F++Q VG + + + E V + GDIV
Sbjct: 306 EYLEVYVSASEHPNHFWIQIVGSRSLQLDKLVNEMTQHYENSVPEDLTVHVGDIVAAPLP 365
Query: 793 ADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLC 852
+ SWYRA V+ G +E+ +++++D+G+ LR L + P A C
Sbjct: 366 TNGSWYRARVL----GTLENGN--LDLYFVDFGDNGDCPLKDLRALRSDFLSLPFQAIEC 419
Query: 853 SLAYIKSPSLEEDFGQEAAEYLSELT 878
SLA I +PS + + +EA + LT
Sbjct: 420 SLARI-APS-GDQWEEEALDEFDRLT 443
>NUC_STAAU (P00644) Thermonuclease precursor (EC 3.1.31.1) (TNase)
(Micrococcal nuclease) (Staphylococcal nuclease)
Length = 231
Score = 42.4 bits (98), Expect = 0.006
Identities = 50/183 (27%), Positives = 81/183 (43%), Gaps = 28/183 (15%)
Query: 541 LTAESRALSGRKGIH--SAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLS 598
L++ + A G++ ++DP V + TS KK LH+ PA + +
Sbjct: 57 LSSSANASQTDNGVNRSGSEDPTVY-----SATSTKK-------LHKE---PATLIKAID 101
Query: 599 GHRFKLLIPKETCSIAFALSGV---RCPGRG-EPYSEEAIALMRRKIMQRDVEIEVE--- 651
G KL+ + + L + P +G E Y EA A + K+++ +IEVE
Sbjct: 102 GDTVKLMYKGQPMTFRLLLVDTPETKHPKKGVEKYGPEASAFTK-KMVENAKKIEVEFDK 160
Query: 652 --TVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKL 709
D+ G L ++ V L+ GLAK+ + + E HL ++E AKK+KL
Sbjct: 161 GQRTDKYGRGLAYIYADGKMVNEALVRQGLAKVAYVYKPNNTHEQHLR-KSEAQAKKEKL 219
Query: 710 KIW 712
IW
Sbjct: 220 NIW 222
Score = 33.9 bits (76), Expect = 2.2
Identities = 32/115 (27%), Positives = 54/115 (46%), Gaps = 19/115 (16%)
Query: 260 RLAVSSSTET-----AADPFGPDAKFFTEMRVLN-RDVRIVLEG---VDKFSNLIGSVYY 310
RL + + ET + +GP+A FT+ V N + + + + DK+ + +Y
Sbjct: 117 RLLLVDTPETKHPKKGVEKYGPEASAFTKKMVENAKKIEVEFDKGQRTDKYGRGLAYIY- 175
Query: 311 PDGESAKDLALELVENGYAK--YVEWSANMMEEEAKRRLKTAELEAKKSRLRMWT 363
DG+ + LV G AK YV N E+ L+ +E +AKK +L +W+
Sbjct: 176 ADGKMVNEA---LVRQGLAKVAYVYKPNNTHEQH----LRKSEAQAKKEKLNIWS 223
>AKP1_HUMAN (Q92667) A kinase anchor protein 1, mitochondrial
precursor (Protein kinase A anchoring protein 1) (PRKA1)
(A-kinase anchor protein 149 kDa) (AKAP 149) (Dual
specificity A-kinase anchoring protein 1) (D-AKAP-1)
(Spermatid A-kinase anchor prote
Length = 903
Score = 41.2 bits (95), Expect = 0.014
Identities = 46/202 (22%), Positives = 83/202 (40%), Gaps = 21/202 (10%)
Query: 735 LKVIVTEVLGGGKFYVQTVGDQKIASI----QQQLAALNLKEAPVLGAFSPKKGDIVLCY 790
++VIV + G +VQ ++ QQ + P L +P + ++
Sbjct: 710 VEVIVVNQVNAGHLFVQQHTHPTFHALRSLDQQMYLCYSQPGIPTLP--TPVEITVICAA 767
Query: 791 FHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQ 850
AD +W+RA VV + E ++ E+ Y+DYG ++V LR + P
Sbjct: 768 PGADGAWWRAQVVAS----YEETNEV-EIRYVDYGGYKRVKVDVLRQIRSDFVTLPFQGA 822
Query: 851 LCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAV 910
L + S ++ F EA +SE+T AQV +G ++ +
Sbjct: 823 EVLLDSVMPLSDDDQFSPEADAAMSEMT--GNTALLAQVTSYSPTG--------LPLIQL 872
Query: 911 TLVAVDAEISVNAAMLQEGLAR 932
V D + +N ++++ GLA+
Sbjct: 873 WSVVGDEVVLINRSLVERGLAQ 894
>AKP1_RAT (O88884) A kinase anchor protein 1, mitochondrial
precursor (Protein kinase A anchoring protein 1) (PRKA1)
(A-kinase anchor protein 121 kDa) (AKAP 121) (Dual
specificity A-Kinase anchoring protein 1) (D-AKAP-1)
(Spermatid A-kinase anchor protein
Length = 854
Score = 40.8 bits (94), Expect = 0.018
Identities = 46/202 (22%), Positives = 83/202 (40%), Gaps = 21/202 (10%)
Query: 735 LKVIVTEVLGGGKFYVQTVGDQKIASI----QQQLAALNLKEAPVLGAFSPKKGDIVLCY 790
++VIV + G +VQ ++ QQ + P L +P + ++
Sbjct: 661 VEVIVVNQVNAGHLFVQQHTHPTFHALRSLDQQMYLCYSQPGIPTLP--TPVEITVICAA 718
Query: 791 FHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQ 850
AD +W+RA VV + E ++ E+ Y+DYG ++V LR + P
Sbjct: 719 PGADGAWWRAQVVAS----YEETNEV-EIRYVDYGGYKRVKVDVLRQIRSDFVTLPFQGA 773
Query: 851 LCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAV 910
L + S ++ F EA +SE+T AQV +G ++ +
Sbjct: 774 EVLLDSVVPLSDDDHFSPEADAAMSEMT--GNTALLAQVTSYSATG--------LPLIQL 823
Query: 911 TLVAVDAEISVNAAMLQEGLAR 932
V D + +N ++++ GLA+
Sbjct: 824 WSVVGDEVVLINRSLVERGLAQ 845
>AKP1_MOUSE (O08715) A kinase anchor protein 1, mitochondrial
precursor (Protein kinase A anchoring protein 1) (PRKA1)
(Dual specificity A-Kinase anchoring protein 1)
(D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP)
Length = 857
Score = 40.8 bits (94), Expect = 0.018
Identities = 46/202 (22%), Positives = 83/202 (40%), Gaps = 21/202 (10%)
Query: 735 LKVIVTEVLGGGKFYVQTVGDQKIASI----QQQLAALNLKEAPVLGAFSPKKGDIVLCY 790
++VIV + G +VQ ++ QQ + P L +P + ++
Sbjct: 664 VEVIVVNQVNAGHLFVQQHTHPTFHALRSLDQQMYLCYSQPGIPTLP--TPVEITVICAA 721
Query: 791 FHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQ 850
AD +W+RA VV + E ++ E+ Y+DYG ++V LR + P
Sbjct: 722 PGADGAWWRAQVVAS----YEETNEV-EIRYVDYGGYKRVKVDVLRQIRSDFVTLPFQGA 776
Query: 851 LCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAV 910
L + S ++ F EA +SE+T AQV +G ++ +
Sbjct: 777 EVLLDSVVPLSDDDHFSPEADAAMSEMT--GNTALLAQVTSYSATG--------LPLIQL 826
Query: 911 TLVAVDAEISVNAAMLQEGLAR 932
V D + +N ++++ GLA+
Sbjct: 827 WSVVGDEVVLINRSLVERGLAQ 848
>YHCR_BACSU (P54602) Hypothetical protein yhcR precursor
Length = 1217
Score = 38.1 bits (87), Expect = 0.12
Identities = 26/78 (33%), Positives = 39/78 (49%), Gaps = 6/78 (7%)
Query: 291 VRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVENGYA-KYVEWSANMMEEEAKRRLKT 349
V++ E D + L+G V ES ++ LELV+NGYA Y W + EE ++ +
Sbjct: 448 VKVGSEAKDSYGRLLGQVIT---ESGSNVNLELVKNGYAPTYFIWPVD--NEEDYQQFQA 502
Query: 350 AELEAKKSRLRMWTNYVP 367
A AKK + +W P
Sbjct: 503 AVAAAKKDQKGIWNENDP 520
>PAB4_ECOLI (P22997) Protein parB
Length = 281
Score = 37.7 bits (86), Expect = 0.15
Identities = 27/128 (21%), Positives = 56/128 (43%), Gaps = 14/128 (10%)
Query: 596 VLSGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDR 655
++ G +L+ K+ + L + P + + + E A + + +R V ++ + DR
Sbjct: 139 IIDGDTIDVLVDKQPVRVR--LVDIDAPEKRQAFGERARQALAGMVFRRHVLVDEKDTDR 196
Query: 656 NGTFLGSLWESR---------TNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKK 706
G LG++W + NV ++ G+A G PE + R EQ A+
Sbjct: 197 YGRTLGTVWVNMELASRPPQPRNVNAAMVHQGMAWAYRFHGRAADPE---MLRLEQEARG 253
Query: 707 QKLKIWEN 714
+++ +W +
Sbjct: 254 KRVGLWSD 261
>TDR5_HUMAN (Q8NAT2) Tudor domain containing protein 5
Length = 682
Score = 37.0 bits (84), Expect = 0.26
Identities = 30/132 (22%), Positives = 53/132 (39%), Gaps = 13/132 (9%)
Query: 734 VLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALN-------LKEAPVLGAFSPKKGDI 786
++ V V ++ +FY++ ++ + + + + V+ + G +
Sbjct: 119 LIGVFVEYIISPSQFYIRIYSRDSSELLEDMMIEMRRCYSNQLVSDRYVMPECFIQPGHL 178
Query: 787 VLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAP 846
DK WYR ++ K EVFY D+GN V S LR L + P
Sbjct: 179 CCVRISEDKWWYRVIIHRVLE------KQEVEVFYPDFGNIGIVQKSSLRFLKCCYTKLP 232
Query: 847 GLAQLCSLAYIK 858
A CSLA+++
Sbjct: 233 AQAIPCSLAWVR 244
>ST31_MOUSE (Q99MW1) Serine/threonine-protein kinase 31 (EC
2.7.1.37)
Length = 1018
Score = 37.0 bits (84), Expect = 0.26
Identities = 44/166 (26%), Positives = 69/166 (41%), Gaps = 15/166 (9%)
Query: 728 ESKQQEVLKVIVTEVLGGGKFYVQTVGDQK-IASIQQQLAALNLKEAPVLGAFSPKKGDI 786
++ +V V+ + V F+ Q V K I I L+ + V G PKK I
Sbjct: 27 DTHYNKVEDVVGSHVEDAVTFWAQNVSKNKDIMKIGCSLSEVCPLANSVFGNLDPKK--I 84
Query: 787 VLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAP 846
F DK WYR V+ T + K + V YIDYGN E + S + + + +
Sbjct: 85 YGGLFSEDKCWYRCKVLKT----ISDDKCL--VRYIDYGNTEILNRSDIVEIPPELQFS- 137
Query: 847 GLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEER 892
+A+ L ++ PS GQE ++ T F +++ R
Sbjct: 138 SIAKKYRLWGLQIPS-----GQEVTQFDQGRTFLGSLIFEKEIKMR 178
>NUC_STAHY (P43270) Thermonuclease precursor (EC 3.1.31.1) (TNase)
(Micrococcal nuclease) (Staphylococcal nuclease)
Length = 169
Score = 35.0 bits (79), Expect = 0.99
Identities = 21/79 (26%), Positives = 37/79 (46%), Gaps = 9/79 (11%)
Query: 381 TGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEF 440
T KV+ V+ GD IIV D ++ + + + P+ P +P Y +EA +F
Sbjct: 41 TYKVIRVIDGDTIIVDKDG-------KQQNLRMIGVDTPETVKPNTPVQP--YGKEASDF 91
Query: 441 LRTRLLGRQVSVEMEYSRK 459
+ L ++V +E + K
Sbjct: 92 TKRHLTNQKVRLEYDKQEK 110
Score = 32.3 bits (72), Expect = 6.4
Identities = 24/99 (24%), Positives = 49/99 (49%), Gaps = 11/99 (11%)
Query: 269 TAADPFGPDAKFFTEMRVLNRDVRIVL--EGVDKFSNLIGSVYYPDGESAKDLALE-LVE 325
T P+G +A FT+ + N+ VR+ + D++ + V+ K++ E L +
Sbjct: 79 TPVQPYGKEASDFTKRHLTNQKVRLEYDKQEKDRYGRTLAYVWL-----GKEMFNEKLAK 133
Query: 326 NGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTN 364
G A+ + N +E R++ A+ +A+K + +W+N
Sbjct: 134 EGLARAKFYRPNYKYQE---RIEQAQKQAQKLKKNIWSN 169
>E4L1_HUMAN (Q9H4G0) Band 4.1-like protein 1 (Neuronal protein 4.1)
(4.1N)
Length = 881
Score = 34.7 bits (78), Expect = 1.3
Identities = 27/103 (26%), Positives = 46/103 (44%), Gaps = 7/103 (6%)
Query: 154 GAAEASIRNLPPSAIGDASNFDAMGLLAA---NKGSPME----AIVEQVRDGSTLRVYLL 206
G+A P D S+ D GLL + NKG+P + +E D +
Sbjct: 630 GSAFEDFSRSLPELDRDKSDSDTEGLLFSRDLNKGAPSQDDESGGIEDSPDRGACSTPDM 689
Query: 207 PEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPG 249
P+F+ V+ + S + ++ PE V++T ++A +N V G
Sbjct: 690 PQFEPVKTETMTVSSLAIRKKIEPEAVLQTRVSAMDNTQQVDG 732
>PYRG_THETN (Q8R720) CTP synthase (EC 6.3.4.2) (UTP--ammonia ligase)
(CTP synthetase)
Length = 537
Score = 34.3 bits (77), Expect = 1.7
Identities = 60/259 (23%), Positives = 95/259 (36%), Gaps = 50/259 (19%)
Query: 324 VENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGK 383
VE+ Y +E ++E +RL L +S L+ W YV N + GK
Sbjct: 242 VESIYQVPLELERQKVDEYVIKRLN---LPLGQSDLKEWREYVEKEKNPEHQVEVALVGK 298
Query: 384 VVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDE--------------- 428
V++ D I +++ + + VN+ + KV + DE
Sbjct: 299 YVDL--HDAYISVVEALKHAGVYHKTAVNIRWVNAEKVNDKTVDELLKGADGILVPGGFG 356
Query: 429 --------KPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDG--SAVPSPAADSRVM 478
+ YARE K LG Q +V +E++R + G S P V+
Sbjct: 357 DRGIEGKIRAIQYARENKIPYLGLCLGMQCAV-IEFARNVAGLKGANSTEFDPNTPHPVI 415
Query: 479 DFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVI-------RHRDFE 531
D L + D D+ G+ GV +++ G V RHR
Sbjct: 416 D------LMPEQKDIDE------KGGTMRLGVYPCKVIEGTKAYEVYKDELVYERHRHRY 463
Query: 532 ERSNYYDALLTAESRALSG 550
E +N Y LLT++ +SG
Sbjct: 464 EFNNQYRELLTSKGLVISG 482
>CO8G_RABIT (Q28679) Complement component C8 gamma chain precursor
Length = 202
Score = 34.3 bits (77), Expect = 1.7
Identities = 21/63 (33%), Positives = 31/63 (48%), Gaps = 1/63 (1%)
Query: 146 LGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAANKGSPMEAIVEQVRDGSTLRVYL 205
LGRW++ P A ++I + P A DA F LLAA GS + EQ +++
Sbjct: 19 LGRWAQKPRGAPSAISAIQPKANFDAQQFAGTWLLAA-VGSACHFLQEQGHRAEATALHV 77
Query: 206 LPE 208
P+
Sbjct: 78 APQ 80
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.133 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 115,873,569
Number of Sequences: 164201
Number of extensions: 5081428
Number of successful extensions: 10509
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 10450
Number of HSP's gapped (non-prelim): 81
length of query: 990
length of database: 59,974,054
effective HSP length: 120
effective length of query: 870
effective length of database: 40,269,934
effective search space: 35034842580
effective search space used: 35034842580
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0094a.6


