

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.14
(487 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
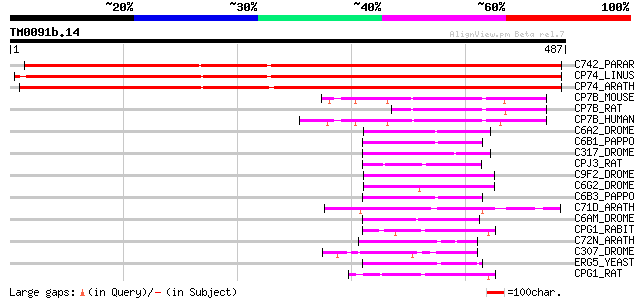
Sequences producing significant alignments: (bits) Value
C742_PARAR (Q40778) Allene oxide synthase (EC 4.2.1.92) (Rubber ... 485 e-137
CP74_LINUS (P48417) Allene oxide synthase, chloroplast precursor... 479 e-135
CP74_ARATH (Q96242) Allene oxide synthase, chloroplast precursor... 473 e-133
CP7B_MOUSE (Q60991) Cytochrome P450 7B1 (Oxysterol 7-alpha-hydro... 60 1e-08
CP7B_RAT (Q63688) Cytochrome P450 7B1 (Oxysterol 7-alpha-hydroxy... 58 5e-08
CP7B_HUMAN (O75881) Cytochrome P450 7B1 (Oxysterol 7-alpha-hydro... 56 2e-07
C6A2_DROME (P33270) Cytochrome P450 6a2 (EC 1.14.-.-) (CYPVIA2) ... 56 2e-07
C6B1_PAPPO (Q04552) Cytochrome P450 6B1 (EC 1.14.14.1) (CYPVIB1)... 54 9e-07
C317_DROME (Q9V776) Probable cytochrome P450 317a1 (EC 1.14.-.-)... 54 9e-07
CPJ3_RAT (P51590) Cytochrome P450 2J3 (EC 1.14.14.1) (CYPIIJ3) 52 3e-06
C9F2_DROME (Q9VG82) Probable cytochrome P450 9f2 (EC 1.14.-.-) (... 51 6e-06
C6G2_DROME (Q9V675) Probable cytochrome P450 6g2 (EC 1.14.-.-) (... 51 6e-06
C6B3_PAPPO (Q27756) Cytochrome P450 6B3 (EC 1.14.14.1) (CYPVIB4)... 51 6e-06
C71D_ARATH (O49342) Cytochrome P450 71A13 (EC 1.14.-.-) 51 8e-06
C6AM_DROME (Q9V769) Cytochrome P450 6a22 (EC 1.14.-.-) (CYPVIA22) 51 8e-06
CPG1_RABIT (P24461) Cytochrome P450 2G1 (EC 1.14.14.1) (CYPIIG1)... 50 1e-05
C72N_ARATH (Q9LTM0) Cytochrome P450 71B23 (EC 1.14.-.-) 50 2e-05
C307_DROME (Q9VRM7) Probable cytochrome P450 307a1 (EC 1.14.-.-)... 50 2e-05
ERG5_YEAST (P54781) Cytochrome P450 61 (EC 1.14.14.-) (C-22 ster... 49 2e-05
CPG1_RAT (P10610) Cytochrome P450 2G1 (EC 1.14.14.1) (CYPIIG1) (... 49 2e-05
>C742_PARAR (Q40778) Allene oxide synthase (EC 4.2.1.92) (Rubber
particle protein) (RPP)
Length = 473
Score = 485 bits (1249), Expect = e-137
Identities = 244/473 (51%), Positives = 327/473 (68%), Gaps = 6/473 (1%)
Query: 14 SNLPLKPIPGSYGLPFFGAMHDRHDYFYNQG-RDKFFATRVEKYNSTVIRTNMPPGPGIS 72
S+ PL+ IPGSYG+PFF + DR +YFY G RD++F +R++KY STV R NMPPGP +S
Sbjct: 4 SSKPLREIPGSYGIPFFQPIKDRLEYFYGTGGRDEYFRSRMQKYQSTVFRANMPPGPFVS 63
Query: 73 SDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPSHKLLK 132
S+ KVI LLD+ SFPILFD SKVEK+D+ GT+MPST TG YR ++ D +EP H LK
Sbjct: 64 SNPKVIVLLDAKSFPILFDVSKVEKKDLFTGTYMPSTKLTGAYRVLSYLDPSEPRHAQLK 123
Query: 133 TFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNFLFLLLTD 192
+L + N +P TT ++ F L+++LA K+GKA+FN A F FL +
Sbjct: 124 NLLFFMLKNSSNRVIPQFETTYTELFEGLEAELA-KNGKAAFNDVGEQAAFRFLGRAYFN 182
Query: 193 KHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAWTVKWSYKK 252
+P ET +G P L+++W+ LAP LGLP + ++ L+ T PA+ +K +Y K
Sbjct: 183 SNPEETKLGTSAPTLISSWVLFNLAPTLDLGLPW---FLQEPLLHTFRLPAFLIKSTYNK 239
Query: 253 LCEGFASAGASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLAGPEL 312
L + F S ++++AE++G+ +DEA HN++F + FN FGG+ FP +KWIG+AG L
Sbjct: 240 LYDYFQSVATPVMEQAEKLGVPKDEAVHNILFAVCFNTFGGVKILFPNTLKWIGVAGENL 299
Query: 313 HAKLAEEIRTVVRE-SNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVES 371
H +LAEEIR ++ +G V+L A+++M LTKSVVYE LRIEP VP QY KA+ + +ES
Sbjct: 300 HTQLAEEIRGAIKSYGDGNVTLEAIEQMPLTKSVVYESLRIEPPVPPQYGKAKSNFTIES 359
Query: 372 HDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWSNGRETEEP 431
HDA FE+KKGEM+FGYQPFATKDPKVF+ PEEFV RFVG+GE LLKYV+WSNG ETE P
Sbjct: 360 HDATFEVKKGEMLFGYQPFATKDPKVFDRPEEFVPDRFVGDGEALLKYVWWSNGPETESP 419
Query: 432 TVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKAT 484
TV+NKQC K+ VVL+ RL+V+E F RYD+F E LG VT+ L +A+
Sbjct: 420 TVENKQCAGKDFVVLITRLFVIELFRRYDSFEIELGESPLGAAVTLTFLKRAS 472
>CP74_LINUS (P48417) Allene oxide synthase, chloroplast precursor
(EC 4.2.1.92) (Hydroperoxide dehydrase) (Cytochrome P450
74A)
Length = 536
Score = 479 bits (1234), Expect = e-135
Identities = 241/481 (50%), Positives = 332/481 (68%), Gaps = 10/481 (2%)
Query: 5 SSDDTKNSSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRTN 64
SSD+T LP++ IPG YGLP G + DR DYFYNQGR++FF +R++KY STV R N
Sbjct: 64 SSDET-----TLPIRQIPGDYGLPGIGPIQDRLDYFYNQGREEFFKSRLQKYKSTVYRAN 118
Query: 65 MPPGPGISSDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTT 124
MPPGP I+S+ +VI LLD+ SFP+LFD SKVEK+D+ GT+MPST TGGYR ++ D +
Sbjct: 119 MPPGPFIASNPRVIVLLDAKSFPVLFDMSKVEKKDLFTGTYMPSTELTGGYRILSYLDPS 178
Query: 125 EPSHKLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFN 184
EP+H LK ++ ++ + +P ++ +D ++ LA K GKA+FN A FN
Sbjct: 179 EPNHTKLKQLLFNLIKNRRDYVIPEFSSSFTDLCEVVEYDLATK-GKAAFNDPAEQAAFN 237
Query: 185 FLFLLLTDKHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAW 244
FL P +T +G P L++ W+ LAP+ ++GLPK E+ + ++ P
Sbjct: 238 FLSRAFFGVKPIDTPLGKDAPSLISKWVLFNLAPILSVGLPK---EVEEATLHSVRLPPL 294
Query: 245 TVKWSYKKLCEGFASAGASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKW 304
V+ Y +L E F SA S+LDEAE+ GI RDEA HN++F + FN++GG FP L+KW
Sbjct: 295 LVQNDYHRLYEFFTSAAGSVLDEAEQSGISRDEACHNILFAVCFNSWGGFKILFPSLMKW 354
Query: 305 IGLAGPELHAKLAEEIRTVVRESNG-EVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKA 363
IG AG ELH KLA+EIR+ ++ + G +V+++A+++M L KSVVYE LRIEP V QY KA
Sbjct: 355 IGRAGLELHTKLAQEIRSAIQSTGGGKVTMAAMEQMPLMKSVVYETLRIEPPVALQYGKA 414
Query: 364 REDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWS 423
++D ++ESH+AA+++K+GEM+FGYQPFATKDPK+F+ PEEFV RFVGEG KL++YV WS
Sbjct: 415 KKDFILESHEAAYQVKEGEMLFGYQPFATKDPKIFDRPEEFVADRFVGEGVKLMEYVMWS 474
Query: 424 NGRETEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKA 483
NG ETE P+V NKQC K+ VV+ RL+VVE F RYD+F E LG ++T+ SL ++
Sbjct: 475 NGPETETPSVANKQCAGKDFVVMAARLFVVELFKRYDSFDIEVGTSSLGASITLTSLKRS 534
Query: 484 T 484
T
Sbjct: 535 T 535
>CP74_ARATH (Q96242) Allene oxide synthase, chloroplast precursor
(EC 4.2.1.92) (Hydroperoxide dehydrase) (Cytochrome P450
74A)
Length = 518
Score = 473 bits (1217), Expect = e-133
Identities = 236/477 (49%), Positives = 323/477 (67%), Gaps = 5/477 (1%)
Query: 9 TKNSSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRTNMPPG 68
T+ S +LP++ IPG+YGLP G + DR DYFY+QG ++FF +R+ KYNSTV R NMPPG
Sbjct: 45 TRTGSKDLPIRNIPGNYGLPIVGPIKDRWDYFYDQGAEEFFKSRIRKYNSTVYRVNMPPG 104
Query: 69 PGISSDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPSH 128
I+ + +V+ALLD SFP+LFD KVEK+D+ GT+MPST TGGYR ++ D +EP H
Sbjct: 105 AFIAENPQVVALLDGKSFPVLFDVDKVEKKDLFTGTYMPSTELTGGYRILSYLDPSEPKH 164
Query: 129 KLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNFLFL 188
+ LK +L S N P + T S+ F L+ +L+ K GKA F S FNFL
Sbjct: 165 EKLKNLLFFLLKSSRNRIFPEFQATYSELFDSLEKELSLK-GKADFGGSSDGTAFNFLAR 223
Query: 189 LLTDKHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAWTVKW 248
+P++T + PGL+T W+ L PL ++GLP++ E+ LI T P VK
Sbjct: 224 AFYGTNPADTKLKADAPGLITKWVLFNLHPLLSIGLPRVI---EEPLIHTFSLPPALVKS 280
Query: 249 SYKKLCEGFASAGASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLA 308
Y++L E F + +L EA+++GI R+EA HNL+F FN +GG+ FP ++K IG A
Sbjct: 281 DYQRLYEFFLESAGEILVEADKLGISREEATHNLLFATCFNTWGGMKILFPNMVKRIGRA 340
Query: 309 GPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVV 368
G ++H +LAEEIR+V++ + GE+++ A++KM LTKSVVYE LR EP V QY +A++D+V
Sbjct: 341 GHQVHNRLAEEIRSVIKSNGGELTMGAIEKMELTKSVVYECLRFEPPVTAQYGRAKKDLV 400
Query: 369 VESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVG-EGEKLLKYVYWSNGRE 427
+ESHDAAF++K GEM++GYQP AT+DPK+F+ +EFV RFVG EGEKLL++V WSNG E
Sbjct: 401 IESHDAAFKVKAGEMLYGYQPLATRDPKIFDRADEFVPERFVGEEGEKLLRHVLWSNGPE 460
Query: 428 TEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKAT 484
TE PTV NKQC K+ VVL+ RL+V+E F RYD+F E LG +V SL KA+
Sbjct: 461 TETPTVGNKQCAGKDFVVLVARLFVIEIFRRYDSFDIEVGTSPLGSSVNFSSLRKAS 517
>CP7B_MOUSE (Q60991) Cytochrome P450 7B1 (Oxysterol
7-alpha-hydroxylase) (EC 1.14.13.-) (HCT-1)
Length = 507
Score = 60.1 bits (144), Expect = 1e-08
Identities = 48/215 (22%), Positives = 98/215 (45%), Gaps = 25/215 (11%)
Query: 274 KRDEAE---HNLVFMLGFNAFGGLTNQFPILI--KWIGLAGPELHAKLAEEIRTVVRESN 328
+ D++E H+L F+ + L N P + + L PE L +EI + ++ +
Sbjct: 274 RHDDSEIGAHHLGFL-----WASLANTIPAMFWAMYYILRHPEAMEALRDEIDSFLQSTG 328
Query: 329 GE--------VSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKK 380
+ + LD + +S + EVLR+ + + +ED+ + +F ++K
Sbjct: 329 QKKGPGISVHFTREQLDSLVCLESTILEVLRL-CSYSSIIREVQEDMNLSLESKSFSLRK 387
Query: 381 GEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWSNGRETEEPT----VDNK 436
G+ + + P DP++F+ P+EF RF+ +G+K K ++ G+ + +
Sbjct: 388 GDFVALFPPLIHNDPEIFDAPKEFRFDRFIEDGKK--KSTFFKGGKRLKTYVMPFGLGTS 445
Query: 437 QCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVL 471
+CP + V +L ++E +D + KP+ L
Sbjct: 446 KCPGRYFAVNEMKLLLIELLTYFDLEIIDRKPIGL 480
>CP7B_RAT (Q63688) Cytochrome P450 7B1 (Oxysterol
7-alpha-hydroxylase) (EC 1.14.13.-) (HCT-1) (Fragment)
Length = 414
Score = 58.2 bits (139), Expect = 5e-08
Identities = 34/140 (24%), Positives = 69/140 (49%), Gaps = 7/140 (5%)
Query: 336 LDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQPFATKDP 395
LD + +S + EVLR+ + + +ED+ S ++ ++KG+ + + P DP
Sbjct: 251 LDSLVCLESAILEVLRL-CSYSSIIREVQEDMDFSSESRSYRLRKGDFVAVFPPMIHNDP 309
Query: 396 KVFENPEEFVGHRFVGEGEKLLKYVYWSNGRETEEPTV----DNKQCPAKNLVVLMCRLY 451
+VF+ P++F RFV +G+K K ++ G++ + + +CP + + +L
Sbjct: 310 EVFDAPKDFRFDRFVEDGKK--KTTFFKGGKKLKSYIIPFGLGTSKCPGRYFAINEMKLL 367
Query: 452 VVEFFLRYDTFTFEFKPVVL 471
V+ +D + KP+ L
Sbjct: 368 VIILLTYFDLEVIDTKPIGL 387
>CP7B_HUMAN (O75881) Cytochrome P450 7B1 (Oxysterol
7-alpha-hydroxylase) (EC 1.14.13.-)
Length = 506
Score = 56.2 bits (134), Expect = 2e-07
Identities = 49/233 (21%), Positives = 103/233 (44%), Gaps = 24/233 (10%)
Query: 255 EGFASAGASLLDEAERVGIKRDE--AEHNLVFMLGFNAFGGLTNQFPILI--KWIGLAGP 310
+G++ S D E+ + D H+L F+ + + N P + + L P
Sbjct: 258 QGWSEVFQSRQDVLEKYYVHEDLEIGAHHLGFL-----WASVANTIPTMFWAMYYLLRHP 312
Query: 311 ELHAKLAEEIRTVVRESNGE--------VSLSALDKMTLTKSVVYEVLRIEPAVPYQYAK 362
E A + +EI +++ + + ++ LD + +S ++E LR+ +
Sbjct: 313 EAMAAVRDEIDRLLQSTGQKKGSGFPIHLTREQLDSLICLESSIFEALRLS-SYSTTIRF 371
Query: 363 AREDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYW 422
ED+ + S + ++KG+++ + P DP++FE PEEF RF+ +G+K K ++
Sbjct: 372 VEEDLTLSSETGDYCVRKGDLVAIFPPVLHGDPEIFEAPEEFRYDRFIEDGKK--KTTFF 429
Query: 423 SNGRETE----EPTVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVL 471
G++ + +CP + ++ + +V +D + KP+ L
Sbjct: 430 KRGKKLKCYLMPFGTGTSKCPGRFFALMEIKQLLVILLTYFDLEIIDDKPIGL 482
>C6A2_DROME (P33270) Cytochrome P450 6a2 (EC 1.14.-.-) (CYPVIA2)
(P450-B1)
Length = 506
Score = 55.8 bits (133), Expect = 2e-07
Identities = 34/112 (30%), Positives = 54/112 (47%), Gaps = 1/112 (0%)
Query: 311 ELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVE 370
++ +L EI+TV+ E G+++ ++ MT V+ E LR+ VP+ KA D VV
Sbjct: 331 DIQDRLRNEIQTVLEEQEGQLTYESIKAMTYLNQVISETLRLYTLVPHLERKALNDYVVP 390
Query: 371 SHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYW 422
H+ I+KG + +D ++ NPE F RF E + V W
Sbjct: 391 GHE-KLVIEKGTQVIIPACAYHRDEDLYPNPETFDPERFSPEKVAARESVEW 441
>C6B1_PAPPO (Q04552) Cytochrome P450 6B1 (EC 1.14.14.1) (CYPVIB1)
(CYP6B1V1/CYP6B1V2/ CYP6B1V3)
Length = 498
Score = 53.9 bits (128), Expect = 9e-07
Identities = 35/106 (33%), Positives = 47/106 (44%), Gaps = 2/106 (1%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
P++ KL EI V+ +G ++ L +MT V E LR P + A+ D V
Sbjct: 323 PDIQDKLIAEIDEVLSRHDGNITYECLSEMTYLSKVFDETLRKYPVADFTQRNAKTDYVF 382
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEK 415
D IKKG+ I DPK + NPE+F RF E K
Sbjct: 383 PGTD--ITIKKGQTIIVSTWGIQNDPKYYPNPEKFDPERFNPENVK 426
>C317_DROME (Q9V776) Probable cytochrome P450 317a1 (EC 1.14.-.-)
(CYPCCCXVIIA1)
Length = 518
Score = 53.9 bits (128), Expect = 9e-07
Identities = 28/113 (24%), Positives = 51/113 (44%), Gaps = 1/113 (0%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
PEL ++ EE++ + +G ++ + ++ V+ E LR+ P PY + D V
Sbjct: 340 PELQVRVREEVKKAIERHDGHITHEGIKSLSFMGQVINETLRMHPITPYILRRTLNDYAV 399
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYW 422
H +K+ +I DP ++ +PEEF R+ G + L + W
Sbjct: 400 PDHPKYILVKELFLIIPTHAI-HHDPDIYPDPEEFKPDRWSGPRDSLQEQGTW 451
>CPJ3_RAT (P51590) Cytochrome P450 2J3 (EC 1.14.14.1) (CYPIIJ3)
Length = 502
Score = 52.0 bits (123), Expect = 3e-06
Identities = 31/105 (29%), Positives = 54/105 (50%), Gaps = 4/105 (3%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
PE+ K+ EI V+ + + +L+ D M T +V++EV RI +P+ + +V V
Sbjct: 331 PEVQEKMQAEIDRVIGQGR-QPNLADRDSMPYTNAVIHEVQRIGNIIPFNVPR---EVAV 386
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGE 414
+++ A F + KG MI +DPK + P+ F F+ G+
Sbjct: 387 DTYLAGFNLPKGTMILTNLTALHRDPKEWATPDTFNPEHFLENGQ 431
>C9F2_DROME (Q9VG82) Probable cytochrome P450 9f2 (EC 1.14.-.-)
(CYPIXF2)
Length = 516
Score = 51.2 bits (121), Expect = 6e-06
Identities = 36/117 (30%), Positives = 59/117 (49%), Gaps = 2/117 (1%)
Query: 311 ELHAKLAEEIRTVVRESNG-EVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
++ +L EE++ V ++ G E++ A+ M VV EVLR PA + +D+
Sbjct: 338 DVQQRLYEEVQQVDQDLEGKELTYEAIMGMKYLDQVVNEVLRKWPAAIAVDRECNKDITF 397
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEG-EKLLKYVYWSNG 425
+ E+KKG++I+ +DPK FENP +F RF E E + + Y+ G
Sbjct: 398 DVDGQKVEVKKGDVIWLPTCGFHRDPKYFENPMKFDPERFSDENKESIQPFTYFPFG 454
>C6G2_DROME (Q9V675) Probable cytochrome P450 6g2 (EC 1.14.-.-)
(CYPVIG2)
Length = 519
Score = 51.2 bits (121), Expect = 6e-06
Identities = 32/118 (27%), Positives = 57/118 (48%), Gaps = 3/118 (2%)
Query: 311 ELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPY--QYAKAREDVV 368
++ +L EEI+ + ES G+V+L ++ + + ++ EVLR+ P +P+ + + D
Sbjct: 337 DVQQRLREEIKDALVESGGQVTLKMIESLEFMQMILLEVLRMYPPLPFLDRECTSGRDYS 396
Query: 369 VESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKL-LKYVYWSNG 425
+ F + KG ++ DP+ F P +F+ RF E KL Y Y G
Sbjct: 397 LAPFHKKFVVPKGMPVYIPCYALHMDPQYFPQPRKFLPERFSPENRKLHTPYTYMPFG 454
>C6B3_PAPPO (Q27756) Cytochrome P450 6B3 (EC 1.14.14.1) (CYPVIB4)
(CYP6B3V1/CYP6B3V2)
Length = 498
Score = 51.2 bits (121), Expect = 6e-06
Identities = 35/106 (33%), Positives = 47/106 (44%), Gaps = 2/106 (1%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
P++ KL EI V+ +G ++ L +MT V E LR P + A+ D V
Sbjct: 323 PDIQDKLIAEIDEVLSRHDGNITYECLGEMTFLGRVFDETLRKYPVGDFTQRNAKTDYVF 382
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEK 415
D IKKG+ I DPK + NPE+F RF E K
Sbjct: 383 PGTD--ITIKKGQTIIVSTWGIQNDPKYYPNPEKFDPERFNPENIK 426
>C71D_ARATH (O49342) Cytochrome P450 71A13 (EC 1.14.-.-)
Length = 497
Score = 50.8 bits (120), Expect = 8e-06
Identities = 53/216 (24%), Positives = 92/216 (42%), Gaps = 28/216 (12%)
Query: 277 EAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIRTVVRESNGEVSL 333
+ + N + + + F G T+ L++W + P+ KL +EIR+ +R +
Sbjct: 282 QVQRNDIKFMILDMFIGGTSTTSTLLEWTMTELIRSPKSMKKLQDEIRSTIRPHGSYIKE 341
Query: 334 SALDKMTLTKSVVYEVLRIEPAVPYQYAK-AREDVVVESHDAAFEIKKGEMIFGYQPFAT 392
++ M K+V+ EVLR+ P++P + EDV V+ + I G +
Sbjct: 342 KEVENMKYLKAVIKEVLRLHPSLPMILPRLLSEDVKVK----GYNIAAGTEVIINAWAIQ 397
Query: 393 KDPKVF-ENPEEFVGHRFVGEG----EKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLM 447
+D ++ + EEF R + G K L Y+ + +GR + CP NL + +
Sbjct: 398 RDTAIWGPDAEEFKPERHLDSGLDYHGKNLNYIPFGSGR---------RICPGINLALGL 448
Query: 448 CRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKA 483
+ V R+D V GPN LT+A
Sbjct: 449 AEVTVANLVGRFDW------RVEAGPNGDQPDLTEA 478
>C6AM_DROME (Q9V769) Cytochrome P450 6a22 (EC 1.14.-.-) (CYPVIA22)
Length = 499
Score = 50.8 bits (120), Expect = 8e-06
Identities = 30/103 (29%), Positives = 50/103 (48%), Gaps = 1/103 (0%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
P L AKL EEI +R NGE + ++ ++ + V+ E LR P +P Q + +
Sbjct: 318 PALQAKLREEIDQALRLHNGEFTYDSMQELRYMELVIAETLRKYPILP-QLTRISRHLYA 376
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE 412
D F I+ G+M+ DP ++ P +F+ RF+ +
Sbjct: 377 AKGDRHFYIEPGQMLLIPVYGIHHDPALYPEPHKFIPERFLAD 419
>CPG1_RABIT (P24461) Cytochrome P450 2G1 (EC 1.14.14.1) (CYPIIG1)
(P450-NMB) (Olfactive)
Length = 494
Score = 50.1 bits (118), Expect = 1e-05
Identities = 35/123 (28%), Positives = 62/123 (49%), Gaps = 13/123 (10%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALD---KMTLTKSVVYEVLRIEPAVPYQYAKARED 366
PE+ K+ EEI V+ G + ++D KM T +V++E+ R+ VP +
Sbjct: 321 PEVQTKIYEEINQVI----GPHRIPSVDDRVKMPFTDAVIHEIQRLTDIVPMGVP---HN 373
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLK---YVYWS 423
V+ ++H + + KG +F KDPK F +P++F F+ E + K +V +S
Sbjct: 374 VIRDTHFRGYLLPKGTDVFPLLGSVLKDPKYFCHPDDFYPQHFLDEQGRFKKNEAFVPFS 433
Query: 424 NGR 426
+G+
Sbjct: 434 SGK 436
>C72N_ARATH (Q9LTM0) Cytochrome P450 71B23 (EC 1.14.-.-)
Length = 501
Score = 49.7 bits (117), Expect = 2e-05
Identities = 29/105 (27%), Positives = 54/105 (50%), Gaps = 5/105 (4%)
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAK-ARE 365
+ P + K+ +E+RTV+ E ++ L+++ K V+ E R+ PA P + A
Sbjct: 320 IRNPRVMKKVQDEVRTVLGEKRDRITEQDLNQLNYFKLVIKETFRLHPAAPLLLPREAMA 379
Query: 366 DVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFV 410
+ ++ +D +K +++ +DP ++ENPEEF RFV
Sbjct: 380 KIKIQGYDIP---EKTQIMVNVYAIG-RDPDLWENPEEFKPERFV 420
>C307_DROME (Q9VRM7) Probable cytochrome P450 307a1 (EC 1.14.-.-)
(CYPCCCVIIA1)
Length = 543
Score = 49.7 bits (117), Expect = 2e-05
Identities = 38/143 (26%), Positives = 71/143 (49%), Gaps = 18/143 (12%)
Query: 275 RDEAEHNLVFML-----GFNAFGGLTNQFPILIKWIGLAGPELHAKLAEEIRTVVRESNG 329
+D + + ++FML G +A G L +++ +I ++ ++ EEI ++ E N
Sbjct: 310 KDVSRNTIIFMLEDFIGGHSAVGNLVM---LVLAYIA-KNVDIGRRIQEEIDAIIEEENR 365
Query: 330 EVSLSALDKMTLTKSVVYEVLRI--EPAVPYQYAKAREDVVVESHDAAFEIKKGEMIFGY 387
++L ++ M T + ++EVLR P VP+ A ED V+ + + + KG ++F
Sbjct: 366 SINLLDMNAMPYTMATIFEVLRYSSSPIVPH---VATEDTVI----SGYGVTKGTIVFIN 418
Query: 388 QPFATKDPKVFENPEEFVGHRFV 410
K + NP+EF RF+
Sbjct: 419 NYVLNTSEKFWVNPKEFNPLRFL 441
>ERG5_YEAST (P54781) Cytochrome P450 61 (EC 1.14.14.-) (C-22 sterol
desaturase)
Length = 538
Score = 49.3 bits (116), Expect = 2e-05
Identities = 36/107 (33%), Positives = 56/107 (51%), Gaps = 5/107 (4%)
Query: 310 PELHAKLAEEIRTVVR-ESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVV 368
P++ AK+ EE V + + E++L ++KM T V+ E LR P V +++
Sbjct: 356 PDVLAKIREEQLAVRNNDMSTELNLDLIEKMKYTNMVIKETLRYRPPVLMVPYVVKKNFP 415
Query: 369 VESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEK 415
V + A KG M+ A DP+V+ENP+EF+ R+V EG K
Sbjct: 416 VSPNYTA---PKGAMLIPTLYPALHDPEVYENPDEFIPERWV-EGSK 458
>CPG1_RAT (P10610) Cytochrome P450 2G1 (EC 1.14.14.1) (CYPIIG1)
(P450-OLF1) (Olfactive)
Length = 494
Score = 49.3 bits (116), Expect = 2e-05
Identities = 37/132 (28%), Positives = 65/132 (49%), Gaps = 12/132 (9%)
Query: 298 FPILIKWIGLAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVP 357
F +L+K+ PE+ AK+ EEI V+ ++ + KM T +V++E+ R+ VP
Sbjct: 314 FLLLMKY-----PEVEAKIHEEINQVIG-THRTPRVDDRAKMPYTDAVIHEIQRLTDIVP 367
Query: 358 YQYAKAREDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLL 417
+V+ ++H + + KG ++ KDPK F PE F F+ E +
Sbjct: 368 LGVP---HNVIRDTHFRGYFLPKGTDVYPLIGSVLKDPKYFRYPEAFYPQHFLDEQGRFK 424
Query: 418 K---YVYWSNGR 426
K +V +S+G+
Sbjct: 425 KNDAFVAFSSGK 436
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 59,445,646
Number of Sequences: 164201
Number of extensions: 2604410
Number of successful extensions: 5977
Number of sequences better than 10.0: 348
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 290
Number of HSP's that attempted gapping in prelim test: 5774
Number of HSP's gapped (non-prelim): 390
length of query: 487
length of database: 59,974,054
effective HSP length: 114
effective length of query: 373
effective length of database: 41,255,140
effective search space: 15388167220
effective search space used: 15388167220
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0091b.14


