

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.13
(267 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
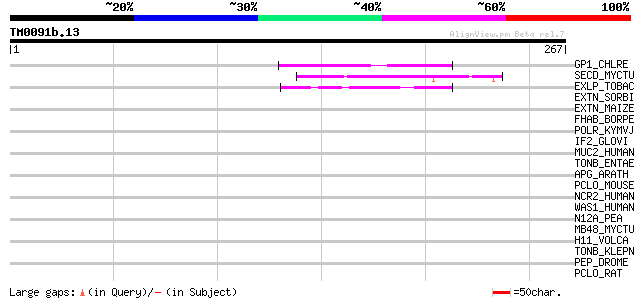
Score E
Sequences producing significant alignments: (bits) Value
GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor (H... 47 5e-05
SECD_MYCTU (Q50634) Protein-export membrane protein secD 44 4e-04
EXLP_TOBAC (Q03211) Pistil-specific extensin-like protein precur... 44 4e-04
EXTN_SORBI (P24152) Extensin precursor (Proline-rich glycoprotein) 43 0.001
EXTN_MAIZE (P14918) Extensin precursor (Proline-rich glycoprotein) 43 0.001
FHAB_BORPE (P12255) Filamentous hemagglutinin 42 0.001
POLR_KYMVJ (P36304) RNA replicase polyprotein (EC 2.7.7.48) 42 0.002
IF2_GLOVI (Q7NH85) Translation initiation factor IF-2 42 0.002
MUC2_HUMAN (Q02817) Mucin 2 precursor (Intestinal mucin 2) 41 0.003
TONB_ENTAE (P46383) TonB protein 40 0.005
APG_ARATH (P40602) Anter-specific proline-rich protein APG precu... 40 0.005
PCLO_MOUSE (Q9QYX7) Piccolo protein (Presynaptic cytomatrix prot... 40 0.008
NCR2_HUMAN (Q9Y618) Nuclear receptor corepressor 2 (N-CoR2) (Sil... 40 0.008
WAS1_HUMAN (Q92558) Wiskott-Aldrich syndrome protein family memb... 39 0.010
N12A_PEA (P20799) Early nodulin 12A precursor (N-12A) 39 0.010
MB48_MYCTU (Q933K8) Antigen MTB48 39 0.010
H11_VOLCA (Q08864) Histone H1-I 39 0.013
TONB_KLEPN (P45610) TonB protein 39 0.017
PEP_DROME (P41073) Zinc finger protein on ecdysone puffs 39 0.017
PCLO_RAT (Q9JKS6) Piccolo protein (Multidomain presynaptic cytom... 39 0.017
>GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor
(Hydroxyproline-rich glycoprotein 1)
Length = 555
Score = 47.0 bits (110), Expect = 5e-05
Identities = 28/84 (33%), Positives = 35/84 (41%), Gaps = 7/84 (8%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+PP PSP+ P P SP PP P V P PP +PS PP+ AP
Sbjct: 289 SPPSPPPSPAPPTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPS-------PPPSPAPP 341
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
+ S S +PS P+ P
Sbjct: 342 TPSPSPSPSPSPSPSPSPSPSPSP 365
Score = 42.4 bits (98), Expect = 0.001
Identities = 27/84 (32%), Positives = 31/84 (36%), Gaps = 10/84 (11%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+P P P PA P SP PP P P PP +P + A APP+ AP
Sbjct: 62 SPAPPSPGPPSPAPPSPPSPAPPS--PAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAP- 118
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
S P PS P P
Sbjct: 119 -------PSPPSPAPPSPSPPAPP 135
Score = 41.6 bits (96), Expect = 0.002
Identities = 25/80 (31%), Positives = 34/80 (42%), Gaps = 3/80 (3%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+P PSP +P P SP PP P + P P + PS + A +PP+ AP
Sbjct: 175 SPTPPSPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAPPSPPSPAPP 234
Query: 190 KKALSQHVSAPKKVTPSKRP 209
+ + P V PS P
Sbjct: 235 SPS---PPAPPSPVPPSPAP 251
Score = 39.7 bits (91), Expect = 0.008
Identities = 25/92 (27%), Positives = 36/92 (38%), Gaps = 6/92 (6%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
+P + +P P+P +P P SP PP P + A P SP+ + +
Sbjct: 180 SPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAPPSPPSPAPPSPSPP 239
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
APP+ P A P PS +P P
Sbjct: 240 APPSPVPPSPA------PPSPAPPSPKPPAPP 265
Score = 38.9 bits (89), Expect = 0.013
Identities = 27/87 (31%), Positives = 36/87 (41%), Gaps = 6/87 (6%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAI---RAPPNT 186
+PP P+P +P P SP PP P V P PP +P + + + APP+
Sbjct: 138 SPP--SPAPPLPPSPAPPSPSPPVPPSPSPP-VPPSPAPPSPTPPSPSPPVPPSPAPPSP 194
Query: 187 APMKKALSQHVSAPKKVTPSKRPATQP 213
AP S V PS P + P
Sbjct: 195 APPVPPSPAPPSPAPPVPPSPAPPSPP 221
Score = 38.9 bits (89), Expect = 0.013
Identities = 28/84 (33%), Positives = 31/84 (36%), Gaps = 10/84 (11%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+P PSP P P SP PP PK P A PP SP PP P
Sbjct: 230 SPAPPSPSPPAPPSPVPPSPAPPSPAPPSPK---PPAPPPPPSPP-------PPPPPRPP 279
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
A + +P PS P T P
Sbjct: 280 FPANTPMPPSPPSPPPSPAPPTPP 303
Score = 38.1 bits (87), Expect = 0.022
Identities = 26/79 (32%), Positives = 29/79 (35%), Gaps = 3/79 (3%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P P P SP PP P + P A P S PS A PP+ AP
Sbjct: 98 PPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSPPSPAPP---LPPSPAPPS 154
Query: 191 KALSQHVSAPKKVTPSKRP 209
+ S V PS P
Sbjct: 155 PSPPVPPSPSPPVPPSPAP 173
Score = 37.7 bits (86), Expect = 0.029
Identities = 27/84 (32%), Positives = 33/84 (39%), Gaps = 5/84 (5%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+PP PSP+ P+ P SP PP P + P P + PS A PP+ AP
Sbjct: 164 SPPVP-PSPAPPS-PTPPSPSPPVPPSPAPPSPAPPVPPSPAPPSPAPP---VPPSPAPP 218
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
S P PS P P
Sbjct: 219 SPPSPAPPSPPSPAPPSPSPPAPP 242
Score = 37.4 bits (85), Expect = 0.038
Identities = 29/83 (34%), Positives = 34/83 (40%), Gaps = 9/83 (10%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P+P PA P SP PP P + P PP SPS A PP+ AP
Sbjct: 199 PP--SPAPPSPAPPVPPSPAPPSPPSPAPPSP-PSPAPP--SPSPPAPPSPVPPSPAPPS 253
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
A +PK P P+ P
Sbjct: 254 PA----PPSPKPPAPPPPPSPPP 272
Score = 36.2 bits (82), Expect = 0.085
Identities = 27/87 (31%), Positives = 29/87 (33%), Gaps = 4/87 (4%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKS----SPSNAAEAIRAPPNT 186
PP P P P SP PP P P P S SPS A +PP+
Sbjct: 83 PPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSPPSP 142
Query: 187 APMKKALSQHVSAPKKVTPSKRPATQP 213
AP S V PS P P
Sbjct: 143 APPLPPSPAPPSPSPPVPPSPSPPVPP 169
Score = 35.8 bits (81), Expect = 0.11
Identities = 23/84 (27%), Positives = 31/84 (36%), Gaps = 1/84 (1%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+P P+P PA P SP PP P + P + P PS A + +PP
Sbjct: 105 SPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSPPSPAPPLPPSPAPPS-PSPPVPPSP 163
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
+ + P PS P P
Sbjct: 164 SPPVPPSPAPPSPTPPSPSPPVPP 187
Score = 35.0 bits (79), Expect = 0.19
Identities = 29/87 (33%), Positives = 34/87 (38%), Gaps = 5/87 (5%)
Query: 131 PPFKKPSPSMPAQP-KTDSPKPPQIKGCQPKNVLP-KADPPKSSPSNAAEAIRA--PPNT 186
PP P P P P ++P PP P P PP SP + A PP+
Sbjct: 266 PPPSPPPPPPPRPPFPANTPMPPSPPSPPPSPAPPTPPTPPSPSPPSPVPPSPAPVPPSP 325
Query: 187 APMKKALSQHVSAPKKVTPSKRPATQP 213
AP A S S P TPS P+ P
Sbjct: 326 APPSPAPSPPPS-PAPPTPSPSPSPSP 351
Score = 34.3 bits (77), Expect = 0.32
Identities = 25/78 (32%), Positives = 31/78 (39%), Gaps = 4/78 (5%)
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
PSPS PA P SP PP P + P + P PS + +P +P + S
Sbjct: 127 PSPSPPAPP---SPSPPSPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPSP 183
Query: 196 HVSAPKKVTPSKRPATQP 213
V P PS P P
Sbjct: 184 PV-PPSPAPPSPAPPVPP 200
Score = 33.5 bits (75), Expect = 0.55
Identities = 27/95 (28%), Positives = 42/95 (43%), Gaps = 20/95 (21%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP-PNTAPM 189
PP P P PA P + +P PP P PP SPS + +P P+ +P
Sbjct: 315 PPSPAPVPPSPA-PPSPAPSPP-----------PSPAPPTPSPSPSPSPSPSPSPSPSP- 361
Query: 190 KKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSN 224
S S +PS +P+ P+ + +L+W++
Sbjct: 362 ----SPSPSPSPIPSPSPKPSPSPV--AVKLVWAD 390
Score = 32.7 bits (73), Expect = 0.94
Identities = 22/79 (27%), Positives = 31/79 (38%), Gaps = 1/79 (1%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P P+P P+ P P PP P +P + P PS A + P + P+
Sbjct: 127 PSPSPPAPPSPSPPSPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPSPPVP 186
Query: 191 KALSQHVSAPKKVTPSKRP 209
+ + AP V PS P
Sbjct: 187 PSPAPPSPAP-PVPPSPAP 204
Score = 30.8 bits (68), Expect = 3.6
Identities = 19/59 (32%), Positives = 22/59 (37%), Gaps = 3/59 (5%)
Query: 151 PPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRP 209
P I C P P PP +P + A APP+ P A S P PS P
Sbjct: 33 PGGIFNCPPSPAPPSPAPPSPAPPSPAPPSPAPPSPGPPSPA---PPSPPSPAPPSPAP 88
>SECD_MYCTU (Q50634) Protein-export membrane protein secD
Length = 573
Score = 43.9 bits (102), Expect = 4e-04
Identities = 36/107 (33%), Positives = 49/107 (45%), Gaps = 10/107 (9%)
Query: 139 SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVS 198
SMPAQP + P+P QP P A PP S + A+ A P P A S + +
Sbjct: 124 SMPAQPAAEEPQPAPSAEPQPPG-QPAAPPPAQSGAPASPQPGAQPRPYPQDPAPSPNPT 182
Query: 199 APKK---VTPSKRPATQPLRRSNRLIWSNRKNLEQS-----QVVAVQ 237
+P P++ PAT P + I + K L QS Q+VA+Q
Sbjct: 183 SPASPPPAPPAEAPATDPRKDLAERI-AQEKKLRQSTNQYMQMVALQ 228
Score = 36.6 bits (83), Expect = 0.065
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 11/83 (13%)
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP--------PKSSPSNAAEAIRAPPNT 186
+P+PS QP PP + P + P A P P +P++ A APP
Sbjct: 135 QPAPSAEPQPPGQPAAPPPAQSGAPASPQPGAQPRPYPQDPAPSPNPTSPASPPPAPPAE 194
Query: 187 APM---KKALSQHVSAPKKVTPS 206
AP +K L++ ++ KK+ S
Sbjct: 195 APATDPRKDLAERIAQEKKLRQS 217
>EXLP_TOBAC (Q03211) Pistil-specific extensin-like protein precursor
(PELP)
Length = 426
Score = 43.9 bits (102), Expect = 4e-04
Identities = 29/83 (34%), Positives = 35/83 (41%), Gaps = 12/83 (14%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP K PSPS QP P PP +K P P PP P A ++P P
Sbjct: 184 PPVKAPSPSPAKQPP---PPPPPVKAPSPS---PATQPPTKQPPPPPRAKKSPLLPPP-- 235
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
V+ P +TPS PA +P
Sbjct: 236 ----PPVAYPPVMTPSPSPAAEP 254
Score = 40.8 bits (94), Expect = 0.003
Identities = 30/86 (34%), Positives = 36/86 (40%), Gaps = 7/86 (8%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQP--KNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
PP K PSPS QP T P PP P P A PP +PS + A PP AP
Sbjct: 202 PPVKAPSPSPATQPPTKQPPPPPRAKKSPLLPPPPPVAYPPVMTPSPSPAA--EPPIIAP 259
Query: 189 MKKALSQHVSAPKKVTPSKRPATQPL 214
+ P++ P P +PL
Sbjct: 260 FPSPPANPPLIPRRPAP---PVVKPL 282
Score = 39.7 bits (91), Expect = 0.008
Identities = 29/92 (31%), Positives = 42/92 (45%), Gaps = 8/92 (8%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKS-SPSNAAEAIRAPPNTAPM 189
PP PSP P+ P + +PPQ + + P P K+ SPS A + PP P
Sbjct: 148 PPPPPPSPCKPSPPDQSAKQPPQPPPAKQPSPPPPPPPVKAPSPSPA----KQPPPPPPP 203
Query: 190 KKALSQHVSAPKKVTPSKRPATQPLRRSNRLI 221
KA S +P P+K+P P + + L+
Sbjct: 204 VKAPS---PSPATQPPTKQPPPPPRAKKSPLL 232
Score = 38.5 bits (88), Expect = 0.017
Identities = 26/84 (30%), Positives = 30/84 (34%), Gaps = 1/84 (1%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP +P PS P+ P PP P + K P + PP AP
Sbjct: 132 PPVNQPKPSSPSPLVKPPPPPPSPCKPSPPDQSAKQPPQPPPAKQPSPPPPPPPVKAPSP 191
Query: 191 KALSQHVSAPKKV-TPSKRPATQP 213
Q P V PS PATQP
Sbjct: 192 SPAKQPPPPPPPVKAPSPSPATQP 215
>EXTN_SORBI (P24152) Extensin precursor (Proline-rich glycoprotein)
Length = 283
Score = 42.7 bits (99), Expect = 0.001
Identities = 27/92 (29%), Positives = 39/92 (42%), Gaps = 5/92 (5%)
Query: 123 PITMIRYNPPFKKPSPSMPA-QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
P+ PP KP P+ +P T P PP P PK PP +P+ A +
Sbjct: 166 PVYTPNPKPPVTKPPTHTPSPKPPTSKPTPPVY---TPSPKPPKPSPPTYTPTPKPPATK 222
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP + P + H P+ P+ +PA +P
Sbjct: 223 -PPTSTPTHPKPTPHTPYPQAHPPTYKPAPKP 253
Score = 40.0 bits (92), Expect = 0.006
Identities = 29/83 (34%), Positives = 33/83 (38%), Gaps = 8/83 (9%)
Query: 131 PPFKKPSPSMPAQ--PKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
PP PSP PA P +PKPP K P P PP + P PP P
Sbjct: 114 PPTYTPSPKPPATKPPTYPTPKPPATKPPTPPVYTPSPKPPVTKPPTPKP---TPPVYTP 170
Query: 189 MKKALSQHVSAPKKVTPSKRPAT 211
K V+ P TPS +P T
Sbjct: 171 NPK---PPVTKPPTHTPSPKPPT 190
Score = 38.1 bits (87), Expect = 0.022
Identities = 32/90 (35%), Positives = 39/90 (42%), Gaps = 9/90 (10%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPP-QIKGCQPKNVL----PKAD-PPKSSPSNAAEAIRAPP 184
PP PSP P T P PP +PK+ + PKA PP +PS A + P
Sbjct: 71 PPTYTPSPKPTPPPATPKPTPPTYTPSPKPKSPVYPPPPKASTPPTYTPSPKPPATKPP- 129
Query: 185 NTAPMKKALSQHVSAPKKVTPS-KRPATQP 213
T P K + P TPS K P T+P
Sbjct: 130 -TYPTPKPPATKPPTPPVYTPSPKPPVTKP 158
Score = 37.7 bits (86), Expect = 0.029
Identities = 28/91 (30%), Positives = 36/91 (38%), Gaps = 5/91 (5%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPK-TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
P+T + P KP S P P T SPKPP+ P P PP + P +
Sbjct: 175 PVTKPPTHTPSPKPPTSKPTPPVYTPSPKPPKPS---PPTYTPTPKPPATKPPTSTPTHP 231
Query: 182 APPNTAPMKKALSQ-HVSAPKKVTPSKRPAT 211
P P +A + APK P+ P T
Sbjct: 232 KPTPHTPYPQAHPPTYKPAPKPSPPAPTPPT 262
Score = 37.4 bits (85), Expect = 0.038
Identities = 26/88 (29%), Positives = 34/88 (38%), Gaps = 7/88 (7%)
Query: 132 PFKKPSPSMPAQPK--TDSPKPPQIKGCQPKNV----LPKADPPKSSPSNAAEAIRAPPN 185
P KP + P P T SPKPP K PK P PP + P + + PP
Sbjct: 132 PTPKPPATKPPTPPVYTPSPKPPVTKPPTPKPTPPVYTPNPKPPVTKPPTHTPSPK-PPT 190
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ P + PK P+ P +P
Sbjct: 191 SKPTPPVYTPSPKPPKPSPPTYTPTPKP 218
Score = 33.1 bits (74), Expect = 0.72
Identities = 26/81 (32%), Positives = 31/81 (38%), Gaps = 9/81 (11%)
Query: 129 YNPPFKKPSPSMPAQ---PKTDSPKPPQIKGCQPK----NVLPKADPP--KSSPSNAAEA 179
Y P K P PS P PK + KPP PK P+A PP K +P + A
Sbjct: 198 YTPSPKPPKPSPPTYTPTPKPPATKPPTSTPTHPKPTPHTPYPQAHPPTYKPAPKPSPPA 257
Query: 180 IRAPPNTAPMKKALSQHVSAP 200
P T P+ S P
Sbjct: 258 PTPPTYTPPVSHTPSSPPPPP 278
Score = 32.7 bits (73), Expect = 0.94
Identities = 32/105 (30%), Positives = 40/105 (37%), Gaps = 26/105 (24%)
Query: 129 YNPPFKKPSPSMPAQ-PKTDSP----------KPPQIKGCQPKNVL--PKADPPKSSPSN 175
Y + P+P PA+ PK + P KPP+ P PK PP ++P
Sbjct: 30 YGGGYPTPTPKPPAKGPKPEKPPTKGHGHKPEKPPKEHKPTPPTYTPSPKPTPPPATPK- 88
Query: 176 AAEAIRAPPNTAPMKKALS------QHVSAPKKVTPS-KRPATQP 213
PP P K S S P TPS K PAT+P
Sbjct: 89 -----PTPPTYTPSPKPKSPVYPPPPKASTPPTYTPSPKPPATKP 128
>EXTN_MAIZE (P14918) Extensin precursor (Proline-rich glycoprotein)
Length = 267
Score = 42.7 bits (99), Expect = 0.001
Identities = 30/97 (30%), Positives = 36/97 (36%), Gaps = 12/97 (12%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNV-----------LPKADPPKSSPSNAA 177
Y P K P+P T SPKPP K PK PK PP +PS
Sbjct: 58 YTPSPKPPTPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKP 117
Query: 178 EAIRAP-PNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
A + P P P S PK P+ P+ +P
Sbjct: 118 PATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKP 154
Score = 42.7 bits (99), Expect = 0.001
Identities = 31/102 (30%), Positives = 38/102 (36%), Gaps = 12/102 (11%)
Query: 124 ITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNV-----------LPKADPPKSS 172
+T Y P K P+P T SPKPP K PK PK PP +
Sbjct: 16 LTPPTYTPSPKPPTPKPTPPTYTPSPKPPASKPPTPKPTPPTYTPSPKPPTPKPTPPTYT 75
Query: 173 PSNAAEAIRAP-PNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PS A + P P P S PK P+ P+ +P
Sbjct: 76 PSPKPPATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKP 117
Score = 41.2 bits (95), Expect = 0.003
Identities = 30/85 (35%), Positives = 35/85 (40%), Gaps = 5/85 (5%)
Query: 131 PPFKKPSPSMPA-QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR-APPNTAP 188
PP PSP PA +P T P PP PK PK PP +PS + PP P
Sbjct: 108 PPTYTPSPKPPATKPPTPKPTPPTYTP-SPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTP 166
Query: 189 MKKALSQHV--SAPKKVTPSKRPAT 211
K + P TPS +P T
Sbjct: 167 SPKPPTHPTPKPTPPTYTPSPKPPT 191
Score = 41.2 bits (95), Expect = 0.003
Identities = 28/84 (33%), Positives = 32/84 (37%), Gaps = 5/84 (5%)
Query: 131 PPFKKPSPSMPAQPK-TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP KP P P T SPKPP K P P PP + P PP P
Sbjct: 43 PPASKPPTPKPTPPTYTPSPKPPTPKP-TPPTYTPSPKPPATKPPTPKP---TPPTYTPS 98
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
K + + P K PAT+P
Sbjct: 99 PKPPTPKPTPPTYTPSPKPPATKP 122
Score = 40.4 bits (93), Expect = 0.005
Identities = 29/95 (30%), Positives = 36/95 (37%), Gaps = 10/95 (10%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGC---------QPKNVLPKADPPKSSPSNAAEA 179
Y P K P+P T SPKPP K P + PK PP +PS
Sbjct: 132 YTPSPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPTHPTPKPTPPTYTPSPKPPT 191
Query: 180 IR-APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ PP P K + + P K PAT+P
Sbjct: 192 PKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPATKP 226
Score = 40.4 bits (93), Expect = 0.005
Identities = 32/96 (33%), Positives = 36/96 (37%), Gaps = 14/96 (14%)
Query: 131 PPFKKPSPSMPAQPK-TDSPKPPQIKGCQP-----------KNVLPKADPPKSSPSNAAE 178
PP KP P P T SPKPP K P K PK PP +PS
Sbjct: 80 PPATKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPSPKPP 139
Query: 179 AIR-APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ PP P K + P TPS +P T P
Sbjct: 140 TPKPTPPTYTPSPKPPTPK-PTPPTYTPSPKPPTHP 174
Score = 40.0 bits (92), Expect = 0.006
Identities = 28/91 (30%), Positives = 32/91 (34%), Gaps = 17/91 (18%)
Query: 131 PPFKKPSPSMPAQPK--------TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
PP PSP P P T SPKPP K P P PP P+
Sbjct: 161 PPTYTPSPKPPTHPTPKPTPPTYTPSPKPPTPKP-TPPTYTPSPKPPTPKPT-------- 211
Query: 183 PPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP P K + PK P+ P +P
Sbjct: 212 PPTYTPSPKPPATKPPTPKPTPPTYTPTPKP 242
Score = 37.4 bits (85), Expect = 0.038
Identities = 31/94 (32%), Positives = 38/94 (39%), Gaps = 13/94 (13%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKP-PQIKGCQPKNVLPKADPPKSSPSNAAEAI 180
TP +P K P+ PK +PKP P PK PK PP +PS A
Sbjct: 165 TPSPKPPTHPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPAT 224
Query: 181 RAPPNTAPMKKALSQHVSAPKKVTPS-KRPATQP 213
+ PP P P TP+ K PAT+P
Sbjct: 225 K-PPTPKP----------TPPTYTPTPKPPATKP 247
Score = 37.4 bits (85), Expect = 0.038
Identities = 29/90 (32%), Positives = 37/90 (40%), Gaps = 9/90 (10%)
Query: 129 YNPPFKKPSPSMPAQPK------TDSPKPPQIKGCQPK-NVLPKADPPKSSPSNAAEAIR 181
Y P K P+ + P PK T SPKPP K P PK PK +P + +
Sbjct: 111 YTPSPKPPA-TKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPKPPTPKPTPPTYTPSPK 169
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPAT 211
P + P K + +PK TP P T
Sbjct: 170 PPTHPTP-KPTPPTYTPSPKPPTPKPTPPT 198
Score = 33.9 bits (76), Expect = 0.42
Identities = 25/81 (30%), Positives = 28/81 (33%), Gaps = 15/81 (18%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
Y P K P+P T SPKPP K PK P P P+ PP P
Sbjct: 199 YTPSPKPPTPKPTPPTYTPSPKPPATKPPTPKPTPPTYTPTPKPPAT------KPPTYTP 252
Query: 189 MKKALSQHVSAPKKVTPSKRP 209
+ P TPS P
Sbjct: 253 ---------TPPVSHTPSPPP 264
>FHAB_BORPE (P12255) Filamentous hemagglutinin
Length = 3590
Score = 42.4 bits (98), Expect = 0.001
Identities = 36/145 (24%), Positives = 55/145 (37%), Gaps = 18/145 (12%)
Query: 116 PFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSP-------------KPPQIKGCQPKNV 162
P V+ +T P K P +P+T P PP ++ QP
Sbjct: 3371 PRPVVAEKVTTPAVQPQLAKVETVQPVKPETTKPLPKPLPVAKVTKAPPPVVETAQP--- 3427
Query: 163 LPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIW 222
LP P K++P AE +A T ++ A + K+ P+ +P +P ++ R
Sbjct: 3428 LPPVKPQKATPGPVAEVGKATVTTVQVQSAPPKPAPVAKQPAPAPKPKPKPKPKAERPKP 3487
Query: 223 SNRKNLEQSQVV--AVQKIQANAGD 245
L VV VQ +Q A D
Sbjct: 3488 GKTTPLSGRHVVQQQVQVLQRQASD 3512
>POLR_KYMVJ (P36304) RNA replicase polyprotein (EC 2.7.7.48)
Length = 1874
Score = 42.0 bits (97), Expect = 0.002
Identities = 31/106 (29%), Positives = 41/106 (38%), Gaps = 8/106 (7%)
Query: 115 LPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPP----- 169
LP+ P + +P F P P P+QP Q QP P PP
Sbjct: 606 LPYAHSPLPPLPVNSSPLFPPPPPLPPSQPPLSQGPATQAPSAQPTPGEPLLAPPTTELK 665
Query: 170 --KSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
S+P+N + A N P K + S + AP K TP+ T P
Sbjct: 666 PESSNPNNPNPSSSAGSNPPP-KSSSSDNPPAPNKPTPTSSSTTPP 710
Score = 32.7 bits (73), Expect = 0.94
Identities = 27/95 (28%), Positives = 34/95 (35%), Gaps = 9/95 (9%)
Query: 132 PFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP-------- 183
P+ +PS P P SP PP P P PP P + A +AP
Sbjct: 596 PYLALAPS-PFLPYAHSPLPPLPVNSSPLFPPPPPLPPSQPPLSQGPATQAPSAQPTPGE 654
Query: 184 PNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSN 218
P AP L S P PS + P +S+
Sbjct: 655 PLLAPPTTELKPESSNPNNPNPSSSAGSNPPPKSS 689
>IF2_GLOVI (Q7NH85) Translation initiation factor IF-2
Length = 925
Score = 42.0 bits (97), Expect = 0.002
Identities = 30/107 (28%), Positives = 50/107 (46%), Gaps = 21/107 (19%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP +P+P +PA P ++P+P V P A P + P+ P AP
Sbjct: 199 PPEPEPTP-LPA-PAAEAPQP----------VRPPAPPAPAKPT---------PAPAPAP 237
Query: 191 KALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLEQSQVVAVQ 237
+ ++ S P++ P +P ++P +R LI NR L+ ++ V Q
Sbjct: 238 RPTAEQPSEPRRPQPPAQPPSRPEKRGGPLIAPNRGGLQPTRPVPAQ 284
Score = 32.0 bits (71), Expect = 1.6
Identities = 27/89 (30%), Positives = 39/89 (43%), Gaps = 15/89 (16%)
Query: 132 PFKKPSPSMPAQPK---TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
P + P+P PA+P +P+P + +P+ P A PP S P R P AP
Sbjct: 217 PVRPPAPPAPAKPTPAPAPAPRPTAEQPSEPRRPQPPAQPP-SRPEK-----RGGPLIAP 270
Query: 189 MKKALSQHVSAPKKVTPSKRPATQPLRRS 217
+ L P + P+ +PA Q RS
Sbjct: 271 NRGGLQ-----PTRPVPA-QPAPQTPTRS 293
>MUC2_HUMAN (Q02817) Mucin 2 precursor (Intestinal mucin 2)
Length = 5179
Score = 41.2 bits (95), Expect = 0.003
Identities = 26/98 (26%), Positives = 36/98 (36%), Gaps = 6/98 (6%)
Query: 122 TPITMIRYNPPFKKPSP---SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE 178
TP T PP PSP + P T +P PP P + PP ++PS
Sbjct: 1548 TPPTSTTTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTTPSPPTITTTTPPPTTTPSPPTT 1607
Query: 179 AIRAPPNTAPMKKALSQHVSAPKKVT---PSKRPATQP 213
PP T + ++ P T P+ P+ P
Sbjct: 1608 TTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPPP 1645
Score = 40.0 bits (92), Expect = 0.006
Identities = 26/95 (27%), Positives = 36/95 (37%), Gaps = 3/95 (3%)
Query: 122 TPITMIRYNPPFKKPSP---SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE 178
TP T PP PSP + P T +P PP P PP ++PS+
Sbjct: 1627 TPPTSTTTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTTPSPPITTTTTPPPTTTPSSPIT 1686
Query: 179 AIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP T + + S+P T + T P
Sbjct: 1687 TTPSPPTTTMTTPSPTTTPSSPITTTTTPSSTTTP 1721
Score = 37.4 bits (85), Expect = 0.038
Identities = 28/101 (27%), Positives = 37/101 (35%), Gaps = 3/101 (2%)
Query: 115 LPFGVLCTPITMIRYNPPFKKPSP-SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSP 173
LP PI+ PP PSP + P T +P PP P + P +
Sbjct: 1449 LPTTTPSPPISTTTTPPPTTTPSPPTTTPSPPTTTPSPPTTTTTTPPPTTTPSPPMTTPI 1508
Query: 174 SNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
+ A PP T P + + P TPS P T P+
Sbjct: 1509 TPPASTTTLPPTTTPSPPTTTT-TTPPPTTTPSP-PTTTPI 1547
Score = 35.0 bits (79), Expect = 0.19
Identities = 31/99 (31%), Positives = 36/99 (36%), Gaps = 27/99 (27%)
Query: 123 PITMIRYNPPFKKPSPSM------PAQ----PKTDSPKPPQIKGCQPKNVLPKADPPKSS 172
P T PP PSP M PA P T +P PP P PP ++
Sbjct: 1487 PTTTTTTPPPTTTPSPPMTTPITPPASTTTLPPTTTPSPPTTTTTTP--------PPTTT 1538
Query: 173 PSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPAT 211
PS PP T P+ S + P TPS P T
Sbjct: 1539 PS--------PPTTTPITPPTST-TTLPPTTTPSPPPTT 1568
Score = 35.0 bits (79), Expect = 0.19
Identities = 25/91 (27%), Positives = 32/91 (34%), Gaps = 2/91 (2%)
Query: 123 PITMIRYNPPFKKPSP--SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAI 180
P T PP PSP + P P T + P P PP ++PS
Sbjct: 1605 PTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTT 1664
Query: 181 RAPPNTAPMKKALSQHVSAPKKVTPSKRPAT 211
+PP T + S+P TPS T
Sbjct: 1665 PSPPITTTTTPPPTTTPSSPITTTPSPPTTT 1695
Score = 35.0 bits (79), Expect = 0.19
Identities = 24/93 (25%), Positives = 31/93 (32%), Gaps = 2/93 (2%)
Query: 123 PITMIRYNPPFKKPSP--SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAI 180
P T PP PSP + P P T + P P PP ++PS
Sbjct: 1526 PTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPPPTTTTTPPPTTTPSPPTTTT 1585
Query: 181 RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP + S P T + P T P
Sbjct: 1586 PSPPTITTTTPPPTTTPSPPTTTTTTPPPTTTP 1618
Score = 32.3 bits (72), Expect = 1.2
Identities = 24/94 (25%), Positives = 33/94 (34%), Gaps = 5/94 (5%)
Query: 123 PITMIRYNPPFKKPSP---SMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEA 179
P T PP PSP + P T +P PP P + P ++ +
Sbjct: 1409 PPTTTTTLPPTTTPSPPTTTTTTPPPTTTPSPPITTTTTPLPTTTPSPPISTTTTPPPTT 1468
Query: 180 IRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP T P + S P T + P T P
Sbjct: 1469 TPSPPTTTPSPPTTTP--SPPTTTTTTPPPTTTP 1500
Score = 31.6 bits (70), Expect = 2.1
Identities = 20/79 (25%), Positives = 30/79 (37%), Gaps = 2/79 (2%)
Query: 136 PSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALS 194
P P+ P T +P PP P P + PP ++ + +PP T ++
Sbjct: 1612 PPPTTTPSPPTTTPITPPTSTTTLPPTTTP-SPPPTTTTTPPPTTTPSPPTTTTPSPPIT 1670
Query: 195 QHVSAPKKVTPSKRPATQP 213
+ P TPS T P
Sbjct: 1671 TTTTPPPTTTPSSPITTTP 1689
Score = 31.2 bits (69), Expect = 2.7
Identities = 21/85 (24%), Positives = 28/85 (32%), Gaps = 7/85 (8%)
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKAD-------PPKSSPSNAAEAIRAPPNTAP 188
P + P+ P T + PP P P PP ++PS PP T
Sbjct: 1597 PPTTTPSPPTTTTTTPPPTTTPSPPTTTPITPPTSTTTLPPTTTPSPPPTTTTTPPPTTT 1656
Query: 189 MKKALSQHVSAPKKVTPSKRPATQP 213
+ S P T + P T P
Sbjct: 1657 PSPPTTTTPSPPITTTTTPPPTTTP 1681
Score = 31.2 bits (69), Expect = 2.7
Identities = 20/78 (25%), Positives = 27/78 (33%), Gaps = 1/78 (1%)
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P + P+ P T + P P PP ++PS P T +S
Sbjct: 1401 PPTTTPSPPPTTTTTLPPTTTPSPPTTTTTTPPPTTTPSPPITTTTTPLPTTTPSPPIST 1460
Query: 196 HVSAPKKVTPSKRPATQP 213
+ P TPS P T P
Sbjct: 1461 TTTPPPTTTPSP-PTTTP 1477
>TONB_ENTAE (P46383) TonB protein
Length = 243
Score = 40.4 bits (93), Expect = 0.005
Identities = 31/87 (35%), Positives = 39/87 (44%), Gaps = 9/87 (10%)
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSP---SNAAEAIRAPPNTAPMKK 191
KP P +PK PKP K +PK + A+P SP +N A A AP TA K
Sbjct: 93 KPEPKPKPKPK---PKPKPEKKVEPKREVKPAEPRPVSPFENNNTAPARTAPSTTAATAK 149
Query: 192 ALSQHVSAPKKV---TPSKRPATQPLR 215
++ S PK + PS Q LR
Sbjct: 150 PMTTAPSVPKALKRGDPSYPQRAQALR 176
>APG_ARATH (P40602) Anter-specific proline-rich protein APG
precursor
Length = 534
Score = 40.4 bits (93), Expect = 0.005
Identities = 31/103 (30%), Positives = 41/103 (39%), Gaps = 16/103 (15%)
Query: 132 PFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP-----PKSSPSNAAEAI---RAP 183
P P PS P +P+ PKP C P P+ P PK +P A + + P
Sbjct: 103 PSPSPCPSPPPKPQ---PKPVPPPACPPTPPKPQPKPAPPPEPKPAPPPAPKPVPCPSPP 159
Query: 184 PNTAPMKKALSQHVSAPKKV-----TPSKRPATQPLRRSNRLI 221
AP K + H PK PS +PA P + N+ I
Sbjct: 160 KPPAPTPKPVPPHGPPPKPAPAPTPAPSPKPAPSPPKPENKTI 202
Score = 33.1 bits (74), Expect = 0.72
Identities = 25/86 (29%), Positives = 31/86 (35%), Gaps = 6/86 (6%)
Query: 130 NPPFKKPSPS--MPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
NPP PSP P P + PP G P P PK P+ + +PP
Sbjct: 59 NPPTPDPSPKPVAPPGPSSKPVAPP---GPSPCPSPPPKPQPKPPPAPSPSPCPSPP-PK 114
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQP 213
P K + P P +PA P
Sbjct: 115 PQPKPVPPPACPPTPPKPQPKPAPPP 140
Score = 31.6 bits (70), Expect = 2.1
Identities = 24/86 (27%), Positives = 30/86 (33%), Gaps = 8/86 (9%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLP-KADPPKSSPSNAAEAIRAPPNTA 187
Y P+ P+ PK +P P K P P + PPK P + PP +
Sbjct: 52 YPQPWPMNPPTPDPSPKPVAPPGPSSKPVAPPGPSPCPSPPPKPQP-------KPPPAPS 104
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQP 213
P PK V P P T P
Sbjct: 105 PSPCPSPPPKPQPKPVPPPACPPTPP 130
Score = 30.0 bits (66), Expect = 6.1
Identities = 23/76 (30%), Positives = 28/76 (36%), Gaps = 18/76 (23%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP KP P P+ PK +P PK V P PPK +P+ P AP
Sbjct: 147 PPAPKPVPC-PSPPKPPAP--------TPKPVPPHGPPPKPAPA---------PTPAPSP 188
Query: 191 KALSQHVSAPKKVTPS 206
K K P+
Sbjct: 189 KPAPSPPKPENKTIPA 204
>PCLO_MOUSE (Q9QYX7) Piccolo protein (Presynaptic cytomatrix
protein) (Aczonin) (Brain-derived HLMN protein)
Length = 5038
Score = 39.7 bits (91), Expect = 0.008
Identities = 37/144 (25%), Positives = 57/144 (38%), Gaps = 5/144 (3%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P T + P ++ SP K DSPK P+ K P P P + + A
Sbjct: 581 PATASKSPVPSQQASPKKELPSKQDSPKAPESKKPPPLVKQPTLHGPTPATAPQPPVAEA 640
Query: 183 PPNTAPMKK---ALSQHVSAP-KKVTPSKRPATQPLRRS-NRLIWSNRKNLEQSQVVAVQ 237
P AP KK AL + AP V P + T+ L S + +++ + SQV A
Sbjct: 641 LPKPAPPKKPSAALPEQAKAPVADVEPKQPKTTETLTDSPSSAAATSKPAILSSQVQAQA 700
Query: 238 KIQANAGDNAELVEEATQVPATGE 261
++ + + + P TG+
Sbjct: 701 QVTTAPPLKTDSAKTSQSFPPTGD 724
Score = 33.1 bits (74), Expect = 0.72
Identities = 37/138 (26%), Positives = 55/138 (39%), Gaps = 19/138 (13%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP--PKSSPSNAAEAIRAPPNTAP 188
PP P P P P SPKPP PK L A P P + + A+AI TA
Sbjct: 2315 PPPPPPPPPPPPLPPATSPKPP----TYPKRKLAAAAPVAPTAIVTAHADAIPTVEATAA 2370
Query: 189 MKK----ALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLEQSQVV---AVQKIQA 241
+ A +AP V P +P++ P L++++R + + AV +I
Sbjct: 2371 RRSNGLPATKICAAAPPPVPP--KPSSIP----TGLVFTHRPEASKPPIAPKPAVPEIPV 2424
Query: 242 NAGDNAELVEEATQVPAT 259
+ + T +P T
Sbjct: 2425 TTQKTTDTCPKPTGLPLT 2442
Score = 33.1 bits (74), Expect = 0.72
Identities = 29/83 (34%), Positives = 35/83 (41%), Gaps = 12/83 (14%)
Query: 141 PAQPKTDSPKPPQIKGCQ--PKNVLPKADPP-------KSSPSNAAEAIRAPPNTAP-MK 190
PA P +P P Q Q PK+ P K + AAE + A P AP +K
Sbjct: 868 PAAPSKQAPPPSQTLAAQGPPKSTGQHPSAPAKTTAVKKETKGPAAENLEAKPAQAPTVK 927
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
KA P KV SK P T+P
Sbjct: 928 KAEKDKKHPPGKV--SKPPPTEP 948
Score = 31.6 bits (70), Expect = 2.1
Identities = 22/93 (23%), Positives = 35/93 (36%), Gaps = 2/93 (2%)
Query: 131 PPFKKPSPSMPA--QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
P KP P PA +P+ P P + + QP P+ P + P+
Sbjct: 409 PVATKPQPQQPAPAKPQPQHPTPAKPQPQQPTPAKPQPQQPTPAKPQPQHPGLGKPSAQQ 468
Query: 189 MKKALSQHVSAPKKVTPSKRPATQPLRRSNRLI 221
K++SQ V+ P A P + ++ I
Sbjct: 469 PSKSISQTVTGRPLQAPPTSAAQAPAQGLSKTI 501
Score = 31.2 bits (69), Expect = 2.7
Identities = 29/110 (26%), Positives = 38/110 (34%), Gaps = 15/110 (13%)
Query: 116 PFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSN 175
P G L + +P F+ S + AQP P PP P PP P
Sbjct: 2277 PSGGLPVSTHPSKSHPFFRSSSLDISAQPPPPPPPPPP----------PPPPPPPPPPPP 2326
Query: 176 AAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPL-----RRSNRL 220
A P T P +K + AP + + A + RRSN L
Sbjct: 2327 LPPATSPKPPTYPKRKLAAAAPVAPTAIVTAHADAIPTVEATAARRSNGL 2376
Score = 30.8 bits (68), Expect = 3.6
Identities = 23/84 (27%), Positives = 34/84 (40%), Gaps = 11/84 (13%)
Query: 132 PFKKPSPSMPA--QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
P ++P P+ P+ QP P+P Q +P+ P P K P + A P P
Sbjct: 385 PAQQPGPTKPSPQQPIPAKPQPQQPVATKPQPQQPA--PAKPQPQHPTPAKPQPQQPTPA 442
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
K P++ TP+K P
Sbjct: 443 K-------PQPQQPTPAKPQPQHP 459
>NCR2_HUMAN (Q9Y618) Nuclear receptor corepressor 2 (N-CoR2)
(Silencing mediator of retinoic acid and thyroid hormone
receptor) (SMRT) (SMRTe) (Thyroid-,
retinoic-acid-receptor-associated corepressor) (T3
receptor-associating factor) (TRAC) (CTG repeat pr
Length = 2517
Score = 39.7 bits (91), Expect = 0.008
Identities = 43/189 (22%), Positives = 64/189 (33%), Gaps = 18/189 (9%)
Query: 83 PLLNDVDAMHLGAYAEYKKQNVEVFVEH-DDEELPFGVLCTPITMIRYNPPFKKPSPSMP 141
P++ D + G ++ E H E+P G P T+ + PSP
Sbjct: 698 PVVEDEEMEASGVSGNEEEMVEEAEALHASGNEVPRGECSGPATVNNSSDTESIPSPHTE 757
Query: 142 AQPKT--DSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSA 199
A T + PKPP G PP P+ RAP P +A
Sbjct: 758 AAKDTGQNGPKPPATLGAD--------GPPPGPPTPPRRTSRAPIEPTPASEATGAPTPP 809
Query: 200 PKKVTPSKRPATQPLRRSNR-------LIWSNRKNLEQSQVVAVQKIQANAGDNAELVEE 252
P +PS P P + + ++ +AV +A +E EE
Sbjct: 810 PAPPSPSAPPPVVPKEEKEEETAAAPPVEEGEEQKPPAAEELAVDTGKAEEPVKSECTEE 869
Query: 253 ATQVPATGE 261
A + PA G+
Sbjct: 870 AEEGPAKGK 878
Score = 33.1 bits (74), Expect = 0.72
Identities = 27/106 (25%), Positives = 41/106 (38%), Gaps = 23/106 (21%)
Query: 129 YNPPFKKPSPSMPA----------QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE 178
+ PP + +P+ PA QP++D+P+ P G P+ PP + AAE
Sbjct: 982 HEPPREDAAPTKPAPPAPPPPQNLQPESDAPQQP---GSSPRGKSRSPAPPADKEAFAAE 1038
Query: 179 AIRAP----------PNTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
A + P P P ++ + AP S P PL
Sbjct: 1039 AQKLPGDPPCWTSGLPFPVPPREVIKASPHAPDPSAFSYAPPGHPL 1084
>WAS1_HUMAN (Q92558) Wiskott-Aldrich syndrome protein family member
1 (WASP-family protein member 1) (Verprolin homology
domain-containing protein 1)
Length = 559
Score = 39.3 bits (90), Expect = 0.010
Identities = 22/87 (25%), Positives = 34/87 (38%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P F P+P P P + ++ P PP P+ A +A PP AP++
Sbjct: 317 PVFVSPTPPPPPPPLPSALSTSSLRASMTSTPPPPVPPPPPPPATALQAPAVPPPPAPLQ 376
Query: 191 KALSQHVSAPKKVTPSKRPATQPLRRS 217
A AP + P + P+ R+
Sbjct: 377 IAPGVLHPAPPPIAPPLVQPSPPVARA 403
Score = 32.0 bits (71), Expect = 1.6
Identities = 20/96 (20%), Positives = 38/96 (38%), Gaps = 8/96 (8%)
Query: 126 MIRYNPPFKKPSP---SMPAQPKTDS-----PKPPQIKGCQPKNVLPKADPPKSSPSNAA 177
+++ +PP + +P ++P P P PP P + P + ++ ++
Sbjct: 393 LVQPSPPVARAAPVCETVPVHPLPQGEVQGLPPPPPPPPLPPPGIRPSSPVTVTALAHPP 452
Query: 178 EAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ P+TAP S P +V P+ P P
Sbjct: 453 SGLHPTPSTAPGPHVPLMPPSPPSQVIPASEPKRHP 488
>N12A_PEA (P20799) Early nodulin 12A precursor (N-12A)
Length = 110
Score = 39.3 bits (90), Expect = 0.010
Identities = 23/72 (31%), Positives = 32/72 (43%), Gaps = 7/72 (9%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
Y PP P + P Q +T KPPQ + P + P+ +PP+ P + PP P
Sbjct: 32 YEPPVNGPPVNKPPQKETPVHKPPQKE--TPVHKPPQKEPPRHKPPQ-----KEPPRHKP 84
Query: 189 MKKALSQHVSAP 200
K HV+ P
Sbjct: 85 PHKKSHLHVTKP 96
>MB48_MYCTU (Q933K8) Antigen MTB48
Length = 460
Score = 39.3 bits (90), Expect = 0.010
Identities = 29/97 (29%), Positives = 47/97 (47%), Gaps = 17/97 (17%)
Query: 93 LGAYAEYKKQNVEVFVEHDDEELPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPP 152
L YAEY++++ +V E++++ P+ NPP KP P++ K D P PP
Sbjct: 247 LPVYAEYQQRSEKVLTEYNNK-----AALEPV-----NPP--KPPPAI----KIDPPPPP 290
Query: 153 QIKGCQPKNVLPKADPPKSSPSNAAEAI-RAPPNTAP 188
Q +G P ++P +D +P A PP +P
Sbjct: 291 QEQGLIPGFLMPPSDGSGVTPGTGMPAAPMVPPTGSP 327
>H11_VOLCA (Q08864) Histone H1-I
Length = 260
Score = 38.9 bits (89), Expect = 0.013
Identities = 26/78 (33%), Positives = 42/78 (53%), Gaps = 7/78 (8%)
Query: 145 KTDSPKPPQIKGCQPKNVLPKADPPK--SSPSNAAEAIRAP-PNTAPMKKALSQHVSAPK 201
K + PK + + + K PKA+ PK ++P +A + P P AP K+ ++ + PK
Sbjct: 172 KVERPKKEKKEKVEKKKATPKAEKPKKAATPKSAGKKKATPKPKAAP--KSPAKKDAKPK 229
Query: 202 KVTPSKR--PATQPLRRS 217
K TPSK+ P P ++S
Sbjct: 230 KATPSKKAAPKKAPAKKS 247
Score = 34.7 bits (78), Expect = 0.25
Identities = 26/71 (36%), Positives = 34/71 (47%), Gaps = 7/71 (9%)
Query: 142 AQPKTDSPKPPQIKGCQPKNV-LPKADP-PKSSPSNAAEAIRAPPNTAPMKKALSQHVSA 199
A PK + PK K PK+ KA P PK++P + A+ P P KKA + A
Sbjct: 189 ATPKAEKPK----KAATPKSAGKKKATPKPKAAPKSPAKKDAKPKKATPSKKAAPKKAPA 244
Query: 200 PKKVTPSKRPA 210
KK TP + A
Sbjct: 245 -KKSTPKAKEA 254
>TONB_KLEPN (P45610) TonB protein
Length = 243
Score = 38.5 bits (88), Expect = 0.017
Identities = 39/119 (32%), Positives = 50/119 (41%), Gaps = 13/119 (10%)
Query: 104 VEVFVEHDDEELPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPK-NV 162
VE VE + E P V P + + P KP P +PK P+ K QPK V
Sbjct: 64 VEPVVEPEPEPEPEVVPEPPKEAVVIHKPEPKPKPKPKPKPK------PEKKVEQPKREV 117
Query: 163 LPKADPPKSSP---SNAAEAIRAPPNTAPMKKALSQHVSAPK---KVTPSKRPATQPLR 215
P A+P +SP +N A A AP + K S P+ +V PS Q LR
Sbjct: 118 KPAAEPRPASPFENNNTAPARTAPSTSTAAAKPTVTAPSGPRAISRVQPSYPARAQALR 176
>PEP_DROME (P41073) Zinc finger protein on ecdysone puffs
Length = 716
Score = 38.5 bits (88), Expect = 0.017
Identities = 25/77 (32%), Positives = 35/77 (44%), Gaps = 1/77 (1%)
Query: 134 KKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKAL 193
++P P PA +T +P P P KA P + S AA A A + +P KKA
Sbjct: 634 QEPEPE-PAPVQTPAPAEPAPPAKTPAKTPTKAAAPAAVASPAAAATSADASPSPAKKAT 692
Query: 194 SQHVSAPKKVTPSKRPA 210
+A K TP ++ A
Sbjct: 693 PARAAAGAKATPQRQRA 709
>PCLO_RAT (Q9JKS6) Piccolo protein (Multidomain presynaptic
cytomatrix protein)
Length = 5085
Score = 38.5 bits (88), Expect = 0.017
Identities = 31/132 (23%), Positives = 52/132 (38%), Gaps = 7/132 (5%)
Query: 134 KKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKA- 192
K P P P +PK KPP+ K P P P + + A P AP K+
Sbjct: 636 KPPEPKKPPEPK----KPPEPKKPPPLVKQPTLHGPTPATAPQLPVAEALPEPAPPKEPS 691
Query: 193 --LSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLEQSQVVAVQKIQANAGDNAELV 250
L + AP K+P R + + + ++ SQV + +++ + +
Sbjct: 692 GPLPEQAKAPVGDVEPKQPKMTETRADIQSSSTTKPDILSSQVQSQAQVKTASPLKTDSA 751
Query: 251 EEATQVPATGER 262
+ + P TGE+
Sbjct: 752 KPSQSFPPTGEK 763
Score = 34.3 bits (77), Expect = 0.32
Identities = 21/91 (23%), Positives = 38/91 (41%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P +P PA+P+ P P + + P P+ PP ++ + + A P+
Sbjct: 432 PAKPQPQQPTPAKPQPQPPTPAKPQPQPPTATKPQPQPPTATKPHHQQPGLAKPSAQQPT 491
Query: 191 KALSQHVSAPKKVTPSKRPATQPLRRSNRLI 221
K++SQ V+ P A P + ++ I
Sbjct: 492 KSISQTVTGRPLQPPPTSAAQTPAQGLSKTI 522
Score = 33.9 bits (76), Expect = 0.42
Identities = 20/83 (24%), Positives = 32/83 (38%), Gaps = 1/83 (1%)
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALS 194
+P S PA+P+ P P + + QP P+ PP + P P A
Sbjct: 416 QPQQSAPAKPQPQQPAPAKPQPQQPTPAKPQPQPPTPAKPQPQPPTATKPQPQP-PTATK 474
Query: 195 QHVSAPKKVTPSKRPATQPLRRS 217
H P PS + T+ + ++
Sbjct: 475 PHHQQPGLAKPSAQQPTKSISQT 497
Score = 31.6 bits (70), Expect = 2.1
Identities = 25/85 (29%), Positives = 40/85 (46%), Gaps = 15/85 (17%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR--APPNTAP 188
PP ++PSP KP + Q K V+ + PKS S AE ++ AP P
Sbjct: 205 PPLQQPSP-----------KPIPKQQGQVKEVIQQDSSPKSVSSQQAEKVKPQAPGTGKP 253
Query: 189 MKKALSQHVSAPKKVTPSKRPATQP 213
+++ +Q + ++ +P K A QP
Sbjct: 254 SQQSPAQ--TPAQQASPGKPVAQQP 276
Score = 31.2 bits (69), Expect = 2.7
Identities = 24/79 (30%), Positives = 29/79 (36%), Gaps = 7/79 (8%)
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALS 194
KPSP P P P+ P QP+ ++ P K P A A P P K
Sbjct: 394 KPSPQQPI-PAKPQPQQPVATKTQPQ----QSAPAKPQPQQPAPAKPQPQQPTPAKP--Q 446
Query: 195 QHVSAPKKVTPSKRPATQP 213
P K P AT+P
Sbjct: 447 PQPPTPAKPQPQPPTATKP 465
Score = 30.4 bits (67), Expect = 4.7
Identities = 25/82 (30%), Positives = 39/82 (47%), Gaps = 8/82 (9%)
Query: 132 PFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKK 191
P ++P P+ P+ P+ P PQ + QP V K P +S+P+ AP P +
Sbjct: 386 PAQQPGPTKPS-PQQPIPAKPQPQ--QP--VATKTQPQQSAPAKPQPQQPAPAKPQPQQP 440
Query: 192 ALSQHVSAPKKVTPSKRPATQP 213
++ P+ TP+K P QP
Sbjct: 441 TPAK--PQPQPPTPAK-PQPQP 459
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.133 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,137,579
Number of Sequences: 164201
Number of extensions: 1668730
Number of successful extensions: 9626
Number of sequences better than 10.0: 394
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 333
Number of HSP's that attempted gapping in prelim test: 7510
Number of HSP's gapped (non-prelim): 1480
length of query: 267
length of database: 59,974,054
effective HSP length: 108
effective length of query: 159
effective length of database: 42,240,346
effective search space: 6716215014
effective search space used: 6716215014
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0091b.13


