

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.10
(353 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
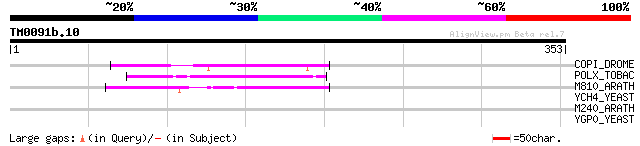
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 72 2e-12
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 58 3e-08
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 48 4e-05
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 43 0.001
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 40 0.007
YGP0_YEAST (P53115) Hypothetical 171.5 kDa helicase in NUT1-ARO2... 31 5.4
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 72.0 bits (175), Expect = 2e-12
Identities = 49/146 (33%), Positives = 74/146 (50%), Gaps = 20/146 (13%)
Query: 65 LALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEEL 124
+ + IE +D++++ Q Y+ K+L F M+ + T L P + N EL
Sbjct: 1124 IGIRIEMQEDKIYLSQSAYVKKILSKFNMENCNAVSTPL-------------PSKINYEL 1170
Query: 125 L----DPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHW-NGKQVFRYLKG 179
L D P S I L+Y+ TR D++ A+N+L+RYSS W N K+V RYLKG
Sbjct: 1171 LNSDEDCNTPCRSLIGCLMYIMLCTRPDLTTAVNILSRYSSKNNSELWQNLKRVLRYLKG 1230
Query: 180 ALDMSLFFP--FMSKLNLIGYADAGY 203
+DM L F + +IGY D+ +
Sbjct: 1231 TIDMKLIFKKNLAFENKIIGYVDSDW 1256
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 58.2 bits (139), Expect = 3e-08
Identities = 42/128 (32%), Positives = 68/128 (52%), Gaps = 4/128 (3%)
Query: 75 EVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPYLSA 134
++++ Q YI +VL+ F M + P+ T L L ++K P E+ +VPY SA
Sbjct: 1057 KLWLSQEKYIERVLERFNMKNAKPVSTPLA-GHLKLSKK-MCPTTVEEKGNMAKVPYSSA 1114
Query: 135 IKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKL 193
+ +L+Y TR DI+ A+ +++R+ +P + HW K + RYL+G L F +
Sbjct: 1115 VGSLMYAMVCTRPDIAHAVGVVSRFLENPGKEHWEAVKWILRYLRGTTGDCLCFGGSDPI 1174
Query: 194 NLIGYADA 201
L GY DA
Sbjct: 1175 -LKGYTDA 1181
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 47.8 bits (112), Expect = 4e-05
Identities = 42/146 (28%), Positives = 71/146 (47%), Gaps = 17/146 (11%)
Query: 62 NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLR---SLNVNKDPFRP* 118
++ L + I+ +F+ Q Y ++L M P+ T L L+ S++ K P
Sbjct: 40 HYFLGIQIKTHPSGLFLSQTKYAEQILNNAGMLDCKPMSTPLPLKLNSSVSTAKYP---- 95
Query: 119 EKNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYL 177
DP + S + AL YL TR DIS+A+N++ + PT ++ K+V RY+
Sbjct: 96 -------DPS-DFRSIVGALQYLTL-TRPDISYAVNIVCQRMHEPTLADFDLLKRVLRYV 146
Query: 178 KGALDMSLFFPFMSKLNLIGYADAGY 203
KG + L+ SKLN+ + D+ +
Sbjct: 147 KGTIFHGLYIHKNSKLNVQAFCDSDW 172
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 43.1 bits (100), Expect = 0.001
Identities = 26/75 (34%), Positives = 41/75 (54%), Gaps = 1/75 (1%)
Query: 130 PYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHW-NGKQVFRYLKGALDMSLFFP 188
PY S + LL+ A+ R DIS+ ++LL+R+ P H + ++V RYL M L +
Sbjct: 182 PYQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTTRSMCLKYR 241
Query: 189 FMSKLNLIGYADAGY 203
S++ L Y DA +
Sbjct: 242 SGSQVALTVYCDASH 256
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 40.4 bits (93), Expect = 0.007
Identities = 23/62 (37%), Positives = 37/62 (59%), Gaps = 1/62 (1%)
Query: 145 TRLDISFAINLLARYSS-SPTQRHWNGKQVFRYLKGALDMSLFFPFMSKLNLIGYADAGY 203
TR D++FA+N L+++SS S T + +V Y+KG + LF+ S L L +AD+ +
Sbjct: 6 TRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAFADSDW 65
Query: 204 LS 205
S
Sbjct: 66 AS 67
>YGP0_YEAST (P53115) Hypothetical 171.5 kDa helicase in NUT1-ARO2
intergenic region
Length = 1489
Score = 30.8 bits (68), Expect = 5.4
Identities = 24/79 (30%), Positives = 36/79 (45%), Gaps = 7/79 (8%)
Query: 72 LKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPY 131
L E Y+R Y+ + +G++ + S P T+ VL S +V N EL DP +
Sbjct: 1191 LNVEKHAYEREYLNCIQRGYHPNVSAPPVTIEVLGSSHVTN------SINNELFDPLISQ 1244
Query: 132 -LSAIKALLYLASHTRLDI 149
LS I A+ H + I
Sbjct: 1245 ALSDIPAITQYNMHVKKGI 1263
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.357 0.160 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,457,339
Number of Sequences: 164201
Number of extensions: 1240247
Number of successful extensions: 5215
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 5208
Number of HSP's gapped (non-prelim): 6
length of query: 353
length of database: 59,974,054
effective HSP length: 111
effective length of query: 242
effective length of database: 41,747,743
effective search space: 10102953806
effective search space used: 10102953806
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0091b.10


