

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
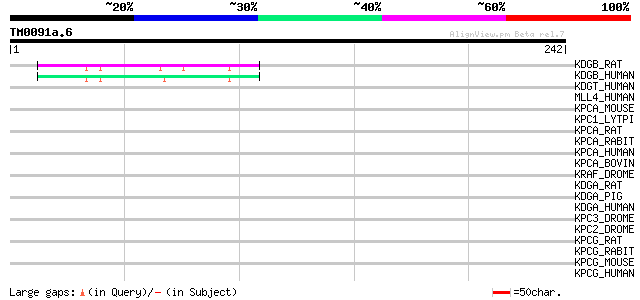
Query= TM0091a.6
(242 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
KDGB_RAT (P49621) Diacylglycerol kinase, beta (EC 2.7.1.107) (Di... 47 4e-05
KDGB_HUMAN (Q9Y6T7) Diacylglycerol kinase, beta (EC 2.7.1.107) (... 45 1e-04
KDGT_HUMAN (P52824) Diacylglycerol kinase, theta (EC 2.7.1.107) ... 40 0.007
MLL4_HUMAN (Q9UMN6) Myeloid/lymphoid or mixed-lineage leukemia p... 37 0.033
KPCA_MOUSE (P20444) Protein kinase C, alpha type (EC 2.7.1.37) (... 36 0.073
KPC1_LYTPI (Q25378) Protein kinase C (EC 2.7.1.-) 36 0.073
KPCA_RAT (P05696) Protein kinase C, alpha type (EC 2.7.1.37) (PK... 35 0.12
KPCA_RABIT (P10102) Protein kinase C, alpha type (EC 2.7.1.37) (... 35 0.12
KPCA_HUMAN (P17252) Protein kinase C, alpha type (EC 2.7.1.37) (... 35 0.12
KPCA_BOVIN (P04409) Protein kinase C, alpha type (EC 2.7.1.37) (... 35 0.12
KRAF_DROME (P11346) Raf homolog serine/threonine-protein kinase ... 35 0.21
KDGA_RAT (P51556) Diacylglycerol kinase, alpha (EC 2.7.1.107) (D... 35 0.21
KDGA_PIG (P20192) Diacylglycerol kinase, alpha (EC 2.7.1.107) (D... 35 0.21
KDGA_HUMAN (P23743) Diacylglycerol kinase, alpha (EC 2.7.1.107) ... 35 0.21
KPC3_DROME (P13678) Protein kinase C (EC 2.7.1.-) (PKC) (dPKC98F) 34 0.28
KPC2_DROME (P13677) Protein kinase C, eye isozyme (EC 2.7.1.-) (... 34 0.36
KPCG_RAT (P63319) Protein kinase C, gamma type (EC 2.7.1.37) (PK... 33 0.47
KPCG_RABIT (P10829) Protein kinase C, gamma type (EC 2.7.1.37) (... 33 0.47
KPCG_MOUSE (P63318) Protein kinase C, gamma type (EC 2.7.1.37) (... 33 0.47
KPCG_HUMAN (P05129) Protein kinase C, gamma type (EC 2.7.1.37) (... 33 0.47
>KDGB_RAT (P49621) Diacylglycerol kinase, beta (EC 2.7.1.107)
(Diglyceride kinase) (DGK-beta) (DAG kinase beta) (90
kDa diacylglycerol kinase)
Length = 801
Score = 47.0 bits (110), Expect = 4e-05
Identities = 34/107 (31%), Positives = 47/107 (43%), Gaps = 10/107 (9%)
Query: 13 QHKLRFEHNAFPFKCDGCKE--IGIGSR-YKCSICDYDLHMQCAIPSPTLVHPFY-TKCS 68
QH R +H P C+ C IG+G + CS C Y +H +CA P+ + + +K +
Sbjct: 243 QHVWRLKHFNKPAYCNLCLNMLIGVGKQGLCCSFCKYTVHERCARAPPSCIKTYVKSKKN 302
Query: 69 FQFMSS--PPGNIPRYCNACEKDVNGFV----YHCKSCGFDLHPCCA 109
M GN P C+ C K V + HC C LH CA
Sbjct: 303 TDVMHHYWVEGNCPTKCDKCHKTVKCYQGLTGLHCVWCQTTLHNKCA 349
>KDGB_HUMAN (Q9Y6T7) Diacylglycerol kinase, beta (EC 2.7.1.107)
(Diglyceride kinase) (DGK-beta) (DAG kinase beta) (90
kDa diacylglycerol kinase)
Length = 804
Score = 45.4 bits (106), Expect = 1e-04
Identities = 33/108 (30%), Positives = 42/108 (38%), Gaps = 11/108 (10%)
Query: 13 QHKLRFEHNAFPFKCDGCKE--IGIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTK--- 66
QH R +H P C+ C IG+G + CS C Y +H +C +P Y K
Sbjct: 244 QHVWRLKHFNKPAYCNLCLNMLIGVGKQGLCCSFCKYTVHERCVARAPPSCIKTYVKSKR 303
Query: 67 -CSFQFMSSPPGNIPRYCNACEKDVNGFV----YHCKSCGFDLHPCCA 109
GN P C+ C K V + HC C LH CA
Sbjct: 304 NTDVMHHYWVEGNCPTKCDKCHKTVKCYQGLTGLHCVWCQITLHNKCA 351
>KDGT_HUMAN (P52824) Diacylglycerol kinase, theta (EC 2.7.1.107)
(Diglyceride kinase) (DGK-theta) (DAG kinase theta)
Length = 942
Score = 39.7 bits (91), Expect = 0.007
Identities = 29/103 (28%), Positives = 40/103 (38%), Gaps = 7/103 (6%)
Query: 12 PQHKLRFEHNAFPFKCDGCKEIGIG-SRYKCSICDYDLHMQCA----IPSPTLVHPFYTK 66
P H R P C C + G + + C +C++ H +C IP T V P +
Sbjct: 59 PGHSFRKVTLTKPTFCHLCSDFIWGLAGFLCDVCNFMSHEKCLKHVRIPC-TSVAPSLVR 117
Query: 67 CSFQFMSSPPGNIPR-YCNACEKDVNGFVYHCKSCGFDLHPCC 108
P G R +C C K + HC+ C LHP C
Sbjct: 118 VPVAHCFGPRGLHKRKFCAVCRKVLEAPALHCEVCELHLHPDC 160
>MLL4_HUMAN (Q9UMN6) Myeloid/lymphoid or mixed-lineage leukemia
protein 4 (Trithorax homolog 2)
Length = 2715
Score = 37.4 bits (85), Expect = 0.033
Identities = 28/95 (29%), Positives = 42/95 (43%), Gaps = 15/95 (15%)
Query: 72 MSSPPGNIPRYCNAC-EKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKLYLYRKV-- 128
++S PG P C C K ++ V+ C+ C HP C L++ E L +
Sbjct: 1193 LTSVPGGPPMVCLLCASKGLHELVF-CQVCCDPFHPFC------LEEAERPLPQHHDTWC 1245
Query: 129 ---SSPCHRCGRKGRSWSYRSKCK--SYNLHVACV 158
CH CGRKGR + +C+ + H AC+
Sbjct: 1246 CRRCKFCHVCGRKGRGSKHLLECERCRHAYHPACL 1280
>KPCA_MOUSE (P20444) Protein kinase C, alpha type (EC 2.7.1.37)
(PKC-alpha) (PKC-A)
Length = 671
Score = 36.2 bits (82), Expect = 0.073
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK + P CD C + G + KC CD ++H QC I P+L
Sbjct: 100 KHKFKIHTYGSPTFCDHCGSLLYGLIHQGMKCDTCDMNVHKQCVINDPSL 149
>KPC1_LYTPI (Q25378) Protein kinase C (EC 2.7.1.-)
Length = 658
Score = 36.2 bits (82), Expect = 0.073
Identities = 29/111 (26%), Positives = 44/111 (39%), Gaps = 36/111 (32%)
Query: 24 PFKCDGCKEI--GIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPPG--- 77
P C CK+ G G + ++C +C + +H +C H F T FQ PG
Sbjct: 38 PTFCSHCKDFIWGFGKQGFQCKVCSFVVHKRC--------HEFVT---FQCPGLDPGVDS 86
Query: 78 --------------NIPRYCNACEKDVNGFVYH----CKSCGFDLHPCCAK 110
N P +C+ C + G +YH C +C ++H C K
Sbjct: 87 DDPRNKHKFKVHSYNSPTFCDHCGSLLYG-LYHQGMKCGACDMNVHKRCQK 136
Score = 31.2 bits (69), Expect = 2.3
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 5/60 (8%)
Query: 12 PQHKLRFEHNAF--PFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTLVHPFYTK 66
P++K +F+ +++ P CD C + G + KC CD ++H +C P L +T+
Sbjct: 89 PRNKHKFKVHSYNSPTFCDHCGSLLYGLYHQGMKCGACDMNVHKRCQKSVPNLCGADHTE 148
>KPCA_RAT (P05696) Protein kinase C, alpha type (EC 2.7.1.37)
(PKC-alpha) (PKC-A)
Length = 671
Score = 35.4 bits (80), Expect = 0.12
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK + P CD C + G + KC CD ++H QC I P+L
Sbjct: 100 KHKFKIHTYGSPTFCDHCGSLLYGLIHQGMKCDTCDMNVHKQCVINVPSL 149
>KPCA_RABIT (P10102) Protein kinase C, alpha type (EC 2.7.1.37)
(PKC-alpha) (PKC-A)
Length = 671
Score = 35.4 bits (80), Expect = 0.12
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK + P CD C + G + KC CD ++H QC I P+L
Sbjct: 100 KHKFKIHTYGSPTFCDHCGSLLYGLIHQGMKCDTCDMNVHKQCVINVPSL 149
>KPCA_HUMAN (P17252) Protein kinase C, alpha type (EC 2.7.1.37)
(PKC-alpha) (PKC-A)
Length = 671
Score = 35.4 bits (80), Expect = 0.12
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK + P CD C + G + KC CD ++H QC I P+L
Sbjct: 100 KHKFKIHTYGSPTFCDHCGSLLYGLIHQGMKCDTCDMNVHKQCVINVPSL 149
>KPCA_BOVIN (P04409) Protein kinase C, alpha type (EC 2.7.1.37)
(PKC-alpha) (PKC-A)
Length = 671
Score = 35.4 bits (80), Expect = 0.12
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK + P CD C + G + KC CD ++H QC I P+L
Sbjct: 100 KHKFKIHTYGSPTFCDHCGSLLYGLIHQGMKCDTCDMNVHKQCVINVPSL 149
>KRAF_DROME (P11346) Raf homolog serine/threonine-protein kinase
dRAF-1 (EC 2.7.1.37) (Pole-hole protein)
Length = 781
Score = 34.7 bits (78), Expect = 0.21
Identities = 19/71 (26%), Positives = 33/71 (45%), Gaps = 4/71 (5%)
Query: 2 KYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVH 61
K+ +H H + F F C+GC+ + + + + CS C++ H +CA P L
Sbjct: 256 KFPIRTHIKHQIIRKTFFSLVF---CEGCRRL-LFTGFYCSQCNFRFHQRCANRVPMLCQ 311
Query: 62 PFYTKCSFQFM 72
PF +Q +
Sbjct: 312 PFPMDSYYQLL 322
>KDGA_RAT (P51556) Diacylglycerol kinase, alpha (EC 2.7.1.107)
(Diglyceride kinase) (DGK-alpha) (DAG kinase alpha) (80
kDa diacylglycerol kinase)
Length = 727
Score = 34.7 bits (78), Expect = 0.21
Identities = 24/96 (25%), Positives = 39/96 (40%), Gaps = 11/96 (11%)
Query: 24 PFKCDGCK-EIGIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSP-----P 76
P C+ C+ IG+G + C++C Y +H CA+ + Y K P
Sbjct: 214 PVYCNLCELSIGLGKQGLSCNLCKYIVHDHCAMKAQPCEVSTYAKSRKDIGVQPHVWVRG 273
Query: 77 GNIPRYCNACEKDVNGF----VYHCKSCGFDLHPCC 108
G C+ C+K + + HC C ++H C
Sbjct: 274 GCHSGRCDRCQKKIRTYHSLTGLHCVWCHLEIHDDC 309
>KDGA_PIG (P20192) Diacylglycerol kinase, alpha (EC 2.7.1.107)
(Diglyceride kinase) (DGK-alpha) (DAG kinase alpha) (80
kDa diacylglycerol kinase)
Length = 734
Score = 34.7 bits (78), Expect = 0.21
Identities = 27/107 (25%), Positives = 44/107 (40%), Gaps = 11/107 (10%)
Query: 13 QHKLRFEHNAFPFKCDGCKE-IGIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQ 70
QH R + P C+ C+ IG+G + C++C Y +H QCA+ + Y K
Sbjct: 204 QHMWRPKRFPRPVYCNLCESSIGLGKQGLSCNLCKYTVHDQCAMKALPCEVSTYAKSRKD 263
Query: 71 F-----MSSPPGNIPRYCNACEKDVNGF----VYHCKSCGFDLHPCC 108
+ G C+ C+K + + HC C ++H C
Sbjct: 264 IGVQTHVWVRGGCESGRCDRCQKKIRIYHSLVGLHCVWCHLEIHDDC 310
>KDGA_HUMAN (P23743) Diacylglycerol kinase, alpha (EC 2.7.1.107)
(Diglyceride kinase) (DGK-alpha) (DAG kinase alpha) (80
kDa diacylglycerol kinase)
Length = 735
Score = 34.7 bits (78), Expect = 0.21
Identities = 27/107 (25%), Positives = 44/107 (40%), Gaps = 11/107 (10%)
Query: 13 QHKLRFEHNAFPFKCDGCKE-IGIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQ 70
QH R + P C+ C+ IG+G + C++C Y +H QCA+ + Y K
Sbjct: 205 QHMWRPKRFPRPVYCNLCESSIGLGKQGLSCNLCKYTVHDQCAMKALPCEVSTYAKSRKD 264
Query: 71 F-----MSSPPGNIPRYCNACEKDVNGF----VYHCKSCGFDLHPCC 108
+ G C+ C+K + + HC C ++H C
Sbjct: 265 IGVQSHVWVRGGCESGRCDRCQKKIRIYHSLTGLHCVWCHLEIHDDC 311
>KPC3_DROME (P13678) Protein kinase C (EC 2.7.1.-) (PKC) (dPKC98F)
Length = 634
Score = 34.3 bits (77), Expect = 0.28
Identities = 27/119 (22%), Positives = 46/119 (37%), Gaps = 22/119 (18%)
Query: 14 HKLRFEHNAFPFKCDGCKEI--GIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQ 70
HK P C C+E GIG + Y+C +C +H +C + + + +
Sbjct: 72 HKFMATFLRQPTFCSHCREFIWGIGKQGYQCQVCTLVVHKKCHLSVVSKCPGMRDEQPAK 131
Query: 71 FMSSPPG-----NIPR-----------YCNACEKDVNGFV---YHCKSCGFDLHPCCAK 110
P G N+P +C+ C + G + C++CG ++H C K
Sbjct: 132 VEMVPAGQRFNVNLPHRFVVHSYKRFTFCDHCGSLLYGLIKQGLQCETCGMNVHKRCQK 190
>KPC2_DROME (P13677) Protein kinase C, eye isozyme (EC 2.7.1.-)
(PKC) (dPKC53E(EY)) (Protein INAC) (Inactivation no
after-potential C protein) (Photoreceptor-specific PKC)
(Eye-PKC)
Length = 700
Score = 33.9 bits (76), Expect = 0.36
Identities = 34/127 (26%), Positives = 54/127 (41%), Gaps = 19/127 (14%)
Query: 16 LRFEHNAFPFKCDGCKEI--GIGSR-YKCSICDYDLHMQC----AIPSPTLVHPFYTKCS 68
+RF N P C CK+ G G + ++C C +++H +C P F C+
Sbjct: 76 VRFFKN--PTYCGHCKDFIWGFGKQGFQCEECRFNIHQKCCKFVVFKCPGKDTDFDADCA 133
Query: 69 --FQFMSSPPGNIPRYCNACEKDVNGFVYH---CKSCGFDLHPCCAKL--PMV-LDDGEV 120
S P +C+ C ++G + C++C ++H C + PM D EV
Sbjct: 134 KVKHGWISTTYTTPTFCDECGLLLHGVAHQGVKCENCNLNVHHACQETVPPMCGADISEV 193
Query: 121 --KLYLY 125
KL LY
Sbjct: 194 RGKLLLY 200
Score = 31.2 bits (69), Expect = 2.3
Identities = 24/93 (25%), Positives = 42/93 (44%), Gaps = 17/93 (18%)
Query: 80 PRYCNACEKDVNGFV---YHCKSCGFDLHPCCAKLPMV----------LDDGEVKL-YLY 125
P YC C+ + GF + C+ C F++H C K + D +VK ++
Sbjct: 82 PTYCGHCKDFIWGFGKQGFQCEECRFNIHQKCCKFVVFKCPGKDTDFDADCAKVKHGWIS 141
Query: 126 RKVSSP--CHRCGRKGRSWSYRS-KCKSYNLHV 155
++P C CG +++ KC++ NL+V
Sbjct: 142 TTYTTPTFCDECGLLLHGVAHQGVKCENCNLNV 174
>KPCG_RAT (P63319) Protein kinase C, gamma type (EC 2.7.1.37)
(PKC-gamma)
Length = 697
Score = 33.5 bits (75), Expect = 0.47
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK R + P CD C + G + KCS C+ ++H +C P+L
Sbjct: 100 KHKFRLHSYSSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVRSVPSL 149
Score = 30.0 bits (66), Expect = 5.2
Identities = 27/113 (23%), Positives = 43/113 (37%), Gaps = 20/113 (17%)
Query: 14 HKLRFEHNAFPFKCDGCKEI--GIGSR-YKCSICDYDLHMQC----------AIPSPTLV 60
HK P C C + GIG + +C +C + +H +C A P
Sbjct: 36 HKFTARFFKQPTFCSHCTDFIWGIGKQGLQCQVCSFVVHRRCHEFVTFECPGAGKGPQTD 95
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYH---CKSCGFDLHPCCAK 110
P K F+ S + P +C+ C + G V+ C C ++H C +
Sbjct: 96 DP-RNKHKFRLHSY---SSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVR 144
>KPCG_RABIT (P10829) Protein kinase C, gamma type (EC 2.7.1.37)
(PKC-gamma) (PKC-delta)
Length = 697
Score = 33.5 bits (75), Expect = 0.47
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK R + P CD C + G + KCS C+ ++H +C P+L
Sbjct: 100 KHKFRLHSYSSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVRTVPSL 149
Score = 30.0 bits (66), Expect = 5.2
Identities = 27/113 (23%), Positives = 43/113 (37%), Gaps = 20/113 (17%)
Query: 14 HKLRFEHNAFPFKCDGCKEI--GIGSR-YKCSICDYDLHMQC----------AIPSPTLV 60
HK P C C + GIG + +C +C + +H +C A P
Sbjct: 36 HKFTARFFKQPTFCSHCTDFIWGIGKQGLQCQVCSFVVHRRCHEFVTFECPGAGKGPQTD 95
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYH---CKSCGFDLHPCCAK 110
P K F+ S + P +C+ C + G V+ C C ++H C +
Sbjct: 96 DP-RNKHKFRLHSY---SSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVR 144
>KPCG_MOUSE (P63318) Protein kinase C, gamma type (EC 2.7.1.37)
(PKC-gamma)
Length = 697
Score = 33.5 bits (75), Expect = 0.47
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK R + P CD C + G + KCS C+ ++H +C P+L
Sbjct: 100 KHKFRLHSYSSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVRSVPSL 149
Score = 30.0 bits (66), Expect = 5.2
Identities = 27/113 (23%), Positives = 43/113 (37%), Gaps = 20/113 (17%)
Query: 14 HKLRFEHNAFPFKCDGCKEI--GIGSR-YKCSICDYDLHMQC----------AIPSPTLV 60
HK P C C + GIG + +C +C + +H +C A P
Sbjct: 36 HKFTARFFKQPTFCSHCTDFIWGIGKQGLQCQVCSFVVHRRCHEFVTFECPGAGKGPQTD 95
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYH---CKSCGFDLHPCCAK 110
P K F+ S + P +C+ C + G V+ C C ++H C +
Sbjct: 96 DP-RNKHKFRLHSY---SSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVR 144
>KPCG_HUMAN (P05129) Protein kinase C, gamma type (EC 2.7.1.37)
(PKC-gamma)
Length = 697
Score = 33.5 bits (75), Expect = 0.47
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Query: 13 QHKLRFEHNAFPFKCDGCKEIGIGSRY---KCSICDYDLHMQCAIPSPTL 59
+HK R + P CD C + G + KCS C+ ++H +C P+L
Sbjct: 100 KHKFRLHSYSSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVRSVPSL 149
Score = 30.0 bits (66), Expect = 5.2
Identities = 27/113 (23%), Positives = 43/113 (37%), Gaps = 20/113 (17%)
Query: 14 HKLRFEHNAFPFKCDGCKEI--GIGSR-YKCSICDYDLHMQC----------AIPSPTLV 60
HK P C C + GIG + +C +C + +H +C A P
Sbjct: 36 HKFTARFFKQPTFCSHCTDFIWGIGKQGLQCQVCSFVVHRRCHEFVTFECPGAGKGPQTD 95
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYH---CKSCGFDLHPCCAK 110
P K F+ S + P +C+ C + G V+ C C ++H C +
Sbjct: 96 DP-RNKHKFRLHSY---SSPTFCDHCGSLLYGLVHQGMKCSCCEMNVHRRCVR 144
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.138 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,318,065
Number of Sequences: 164201
Number of extensions: 1377571
Number of successful extensions: 4023
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 3943
Number of HSP's gapped (non-prelim): 105
length of query: 242
length of database: 59,974,054
effective HSP length: 107
effective length of query: 135
effective length of database: 42,404,547
effective search space: 5724613845
effective search space used: 5724613845
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0091a.6


