

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
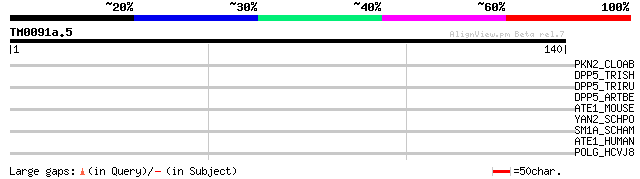
Query= TM0091a.5
(140 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
PKN2_CLOAB (Q97IC2) Probable serine/threonine protein kinase CAC... 31 0.96
DPP5_TRISH (Q8J1L4) Dipeptidyl-peptidase V precursor (EC 3.4.14.... 30 1.3
DPP5_TRIRU (Q9UW98) Dipeptidyl-peptidase V precursor (EC 3.4.14.... 30 1.3
DPP5_ARTBE (Q8J1M3) Dipeptidyl-peptidase V precursor (EC 3.4.14.... 30 1.3
ATE1_MOUSE (Q9Z2A5) Arginyl-tRNA--protein transferase 1 (EC 2.3.... 29 2.8
YAN2_SCHPO (Q10068) Hypothetical protein C3H1.02c in chromosome I 29 3.6
SM1A_SCHAM (Q26473) Semaphorin 1A precursor (Semaphorin-I) (Sema... 28 4.8
ATE1_HUMAN (O95260) Arginyl-tRNA--protein transferase 1 (EC 2.3.... 28 4.8
POLG_HCVJ8 (P26661) Genome polyprotein [Contains: Capsid protein... 28 8.1
>PKN2_CLOAB (Q97IC2) Probable serine/threonine protein kinase
CAC1728 (EC 2.7.1.37)
Length = 657
Score = 30.8 bits (68), Expect = 0.96
Identities = 29/108 (26%), Positives = 50/108 (45%), Gaps = 16/108 (14%)
Query: 23 NIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQ-------- 74
N V+ D + ++ +P + + NPN+ L + ++ Y G P P EE Q
Sbjct: 272 NYVNDDEEFTRVMDPAQIQN--ESNPNNKLDNDDTYYNGEPYNKEQPQEEPQEENEEPKN 329
Query: 75 -LNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS--SSVFEPSSS 119
+ L ++K L S + + I A +ALA S S +F+P+S+
Sbjct: 330 KIKGNNMLSGKAKKALVAS---IIVVLILAGSALAFSMGSGIFKPNSN 374
>DPP5_TRISH (Q8J1L4) Dipeptidyl-peptidase V precursor (EC 3.4.14.-)
(DPP V) (DppV) (Allergen Tri s 4)
Length = 726
Score = 30.4 bits (67), Expect = 1.3
Identities = 27/94 (28%), Positives = 38/94 (39%), Gaps = 5/94 (5%)
Query: 27 FDGKLQQLKEPVK----AWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLV 82
FDGKL+ L +PVK + + ++ L S P +P L ++I FLV
Sbjct: 203 FDGKLKNLVQPVKYAESPYPPFGGSGDYDLSSDGKTVAFMSKAPELPKANLTTSYI-FLV 261
Query: 83 PRSKSRLPLSLEDLCALAIKANAALAHSSSVFEP 116
P SR+ + A SS VF P
Sbjct: 262 PHDGSRVAEPINKRNGPRTPQGIEGASSSPVFSP 295
>DPP5_TRIRU (Q9UW98) Dipeptidyl-peptidase V precursor (EC 3.4.14.-)
(DPP V) (DppV) (Allergen Tri r 4)
Length = 726
Score = 30.4 bits (67), Expect = 1.3
Identities = 27/94 (28%), Positives = 38/94 (39%), Gaps = 5/94 (5%)
Query: 27 FDGKLQQLKEPVK----AWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLV 82
FDGKL+ L +PVK + + ++ L S P +P L ++I FLV
Sbjct: 203 FDGKLKNLVQPVKYAESPYPPFGGSGDYDLSSDGKTVAFMSKAPELPKANLTTSYI-FLV 261
Query: 83 PRSKSRLPLSLEDLCALAIKANAALAHSSSVFEP 116
P SR+ + A SS VF P
Sbjct: 262 PHDGSRVAEPINKRNGPRTPQGIEGASSSPVFSP 295
>DPP5_ARTBE (Q8J1M3) Dipeptidyl-peptidase V precursor (EC 3.4.14.-)
(DPP V) (DppV) (Allergen Tri m 4)
Length = 726
Score = 30.4 bits (67), Expect = 1.3
Identities = 27/94 (28%), Positives = 38/94 (39%), Gaps = 5/94 (5%)
Query: 27 FDGKLQQLKEPVK----AWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLV 82
FDGKL+ L +PVK + + ++ L S P +P L ++I FLV
Sbjct: 203 FDGKLKNLVQPVKYAESPYPPFGGSGDYDLSSDGKTVAFMSKAPELPKANLTTSYI-FLV 261
Query: 83 PRSKSRLPLSLEDLCALAIKANAALAHSSSVFEP 116
P SR+ + A SS VF P
Sbjct: 262 PHDGSRVAEPINKRNGPRTPQGIEGASSSPVFSP 295
>ATE1_MOUSE (Q9Z2A5) Arginyl-tRNA--protein transferase 1 (EC
2.3.2.8) (R-transferase 1) (Arginyltransferase 1)
(Arginine-tRNA--protein transferase 1)
Length = 516
Score = 29.3 bits (64), Expect = 2.8
Identities = 18/46 (39%), Positives = 25/46 (54%), Gaps = 2/46 (4%)
Query: 81 LVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFK 126
L+P++KS P SLEDL ++ NA +H V SS P F+
Sbjct: 240 LLPKAKSNQPKSLEDLIFQSLPENA--SHKLEVRVVRSSPPSPQFR 283
>YAN2_SCHPO (Q10068) Hypothetical protein C3H1.02c in chromosome I
Length = 1036
Score = 28.9 bits (63), Expect = 3.6
Identities = 24/98 (24%), Positives = 42/98 (42%), Gaps = 12/98 (12%)
Query: 15 NHGMQSTVNIVHFDGKLQQLKEPVKAWHVL-----SQNPN-HFLCSSESMYV------GS 62
N+ +Q +H G + +L +P+ W+ L +NPN FL + Y+ S
Sbjct: 746 NYMLQHQAIRIHVIGDILKLPQPISTWNKLLDSECHRNPNMEFLSTFSKNYLKPQEFGSS 805
Query: 63 PMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALA 100
V+P + + + F +P S L SL + +A
Sbjct: 806 SSLTVIPMPSNESSDLVFSIPGITSWLDPSLPVIVLIA 843
>SM1A_SCHAM (Q26473) Semaphorin 1A precursor (Semaphorin-I) (Sema I)
(Fasciclin IV)
Length = 730
Score = 28.5 bits (62), Expect = 4.8
Identities = 16/66 (24%), Positives = 30/66 (45%), Gaps = 4/66 (6%)
Query: 36 EPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLED 95
+PV + H P + +G + V+PN ++NH +P S+ +LP+ +
Sbjct: 572 DPVHSIHQAEFEPE----IDNEIVIGVDDSNVIPNTLAEINHAGSKLPSSQEKLPIYTAE 627
Query: 96 LCALAI 101
+AI
Sbjct: 628 TLTIAI 633
>ATE1_HUMAN (O95260) Arginyl-tRNA--protein transferase 1 (EC
2.3.2.8) (R-transferase 1) (Arginyltransferase 1)
(Arginine-tRNA--protein transferase 1)
Length = 518
Score = 28.5 bits (62), Expect = 4.8
Identities = 18/44 (40%), Positives = 23/44 (51%), Gaps = 2/44 (4%)
Query: 83 PRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFK 126
P++KS P SLEDL ++ NA +H V SS P FK
Sbjct: 244 PKAKSNQPKSLEDLIFESLPENA--SHKLEVRVVRSSPPSSQFK 285
>POLG_HCVJ8 (P26661) Genome polyprotein [Contains: Capsid protein C
(Core protein) (P22); Envelope glycoprotein E1 (GP32)
(GP35); Envelope glycoprotein E2 (GP68) (GP70) (NS1);
Protein P7; Nonstructural protein NS2 (P21) (EC
3.4.22.-); Protease/helicase NS
Length = 3033
Score = 27.7 bits (60), Expect = 8.1
Identities = 13/46 (28%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Query: 28 DGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNEE 72
+G+ + + K+W +S + +C S S + G+ +TP P EE
Sbjct: 2416 EGECEVIDSDSKSWSTVSDQEDSVICCSMSYSWTGALITPCGPEEE 2461
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.130 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,930,874
Number of Sequences: 164201
Number of extensions: 644561
Number of successful extensions: 1397
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1393
Number of HSP's gapped (non-prelim): 10
length of query: 140
length of database: 59,974,054
effective HSP length: 99
effective length of query: 41
effective length of database: 43,718,155
effective search space: 1792444355
effective search space used: 1792444355
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0091a.5


