

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
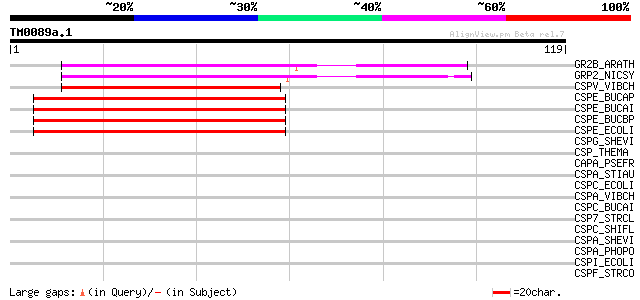
GR2B_ARATH (Q38896) Glycine-rich protein 2b (AtGRP2b) 54 5e-08
GRP2_NICSY (P27484) Glycine-rich protein 2 53 1e-07
CSPV_VIBCH (Q9KL16) Cold shock protein cspV 44 8e-05
CSPE_BUCAP (P63238) Cold shock-like protein cspE (CSP-E) 44 8e-05
CSPE_BUCAI (P63237) Cold shock-like protein cspE (CSP-E) 44 8e-05
CSPE_BUCBP (Q89A90) Cold shock-like protein cspE (CSP-E) 42 2e-04
CSPE_ECOLI (P36997) Cold shock-like protein cspE (CSP-E) 42 2e-04
CSPG_SHEVI (Q9S170) Cold shock-like protein cspG 40 0.001
CSP_THEMA (O54310) Cold shock-like protein 39 0.002
CAPA_PSEFR (P72188) Cold shock protein capA (Cold acclimation pr... 39 0.003
CSPA_STIAU (P72366) Cold shock-like protein cspA 38 0.004
CSPC_ECOLI (P36996) Cold shock-like protein cspC (CSP-C) 38 0.005
CSPA_VIBCH (Q9KN00) Cold shock-like protein cspA 38 0.005
CSPC_BUCAI (P57407) Cold shock-like protein cspC (CSP-C) 37 0.006
CSP7_STRCL (Q01761) Cold shock-like protein 7.0 36 0.014
CSPC_SHIFL (Q83RI9) Cold shock-like protein cspC (CSP-C) 35 0.023
CSPA_SHEVI (Q9S1B7) Cold shock-like protein cspA 35 0.023
CSPA_PHOPO (Q52287) Major cold-shock protein (Fragment) 35 0.030
CSPI_ECOLI (P77605) Cold shock-like protein cspI (CPS-I) 34 0.051
CSPF_STRCO (P48859) Cold shock protein scoF 34 0.051
>GR2B_ARATH (Q38896) Glycine-rich protein 2b (AtGRP2b)
Length = 201
Score = 54.3 bits (129), Expect = 5e-08
Identities = 37/94 (39%), Positives = 48/94 (50%), Gaps = 15/94 (15%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DGG+DLF+ S I+S+GF LA E V+ + + I +
Sbjct: 19 VKWFDTQKGFGFITPSDGGDDLFVHQSSIRSEGFRSLAAEESVEFDVEVDNSGRPKAIEV 78
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGG 98
+ PD AP+ G GG S G G GG
Sbjct: 79 SG--------PDGAPVQGNSGGGGSSGGRGGFGG 104
>GRP2_NICSY (P27484) Glycine-rich protein 2
Length = 214
Score = 52.8 bits (125), Expect = 1e-07
Identities = 39/95 (41%), Positives = 48/95 (50%), Gaps = 16/95 (16%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL-------TILIPM 64
+K F DQKGF IT DDGG DLF+ S I+S+GF LAE E V+ + T + +
Sbjct: 13 VKWFSDQKGFGFITPDDGGEDLFVHQSGIRSEGFRSLAEGETVEFEVESGGDGRTKAVDV 72
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGGY 99
PD A + G GG G G GGGY
Sbjct: 73 TG--------PDGAAVQGGRGGGGGGGGRG-GGGY 98
>CSPV_VIBCH (Q9KL16) Cold shock protein cspV
Length = 70
Score = 43.5 bits (101), Expect = 8e-05
Identities = 20/47 (42%), Positives = 33/47 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL 58
+K F++ KGF +T D+GGND+F+ + I+S+GF LAE + V ++
Sbjct: 9 VKWFNETKGFGFLTQDNGGNDVFVHFNSIQSEGFKTLAEGQRVSFIV 55
>CSPE_BUCAP (P63238) Cold shock-like protein cspE (CSP-E)
Length = 68
Score = 43.5 bits (101), Expect = 8e-05
Identities = 22/54 (40%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQSVEFEIT 54
>CSPE_BUCAI (P63237) Cold shock-like protein cspE (CSP-E)
Length = 68
Score = 43.5 bits (101), Expect = 8e-05
Identities = 22/54 (40%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQSVEFEIT 54
>CSPE_BUCBP (Q89A90) Cold shock-like protein cspE (CSP-E)
Length = 68
Score = 42.4 bits (98), Expect = 2e-04
Identities = 21/54 (38%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I+S GF L+E + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLSEGQSVEFEIT 54
>CSPE_ECOLI (P36997) Cold shock-like protein cspE (CSP-E)
Length = 68
Score = 42.0 bits (97), Expect = 2e-04
Identities = 21/54 (38%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 54
>CSPG_SHEVI (Q9S170) Cold shock-like protein cspG
Length = 70
Score = 40.0 bits (92), Expect = 0.001
Identities = 20/43 (46%), Positives = 29/43 (66%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F+++KGF IT D+GG+D+F+ I S GF LAE + V
Sbjct: 9 VKWFNEEKGFGFITQDNGGDDVFVHFRSITSDGFKTLAEGQKV 51
>CSP_THEMA (O54310) Cold shock-like protein
Length = 66
Score = 38.9 bits (89), Expect = 0.002
Identities = 21/48 (43%), Positives = 32/48 (65%), Gaps = 1/48 (2%)
Query: 8 VREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
+R K+K FD +KG+ IT D+GG D+F+ S I+ +GF L E + V+
Sbjct: 1 MRGKVKWFDSKKGYGFITKDEGG-DVFVHWSAIEMEGFKTLKEGQVVE 47
>CAPA_PSEFR (P72188) Cold shock protein capA (Cold acclimation
protein A) (C7.0) (Fragment)
Length = 64
Score = 38.5 bits (88), Expect = 0.003
Identities = 20/46 (43%), Positives = 28/46 (60%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLL 57
+K F+D+KGF IT GG+DLF+ I+S GF L E + V +
Sbjct: 9 VKWFNDEKGFGFITPQGGGDDLFVHFKAIESDGFKSLKEGQTVSFV 54
>CSPA_STIAU (P72366) Cold shock-like protein cspA
Length = 68
Score = 38.1 bits (87), Expect = 0.004
Identities = 22/53 (41%), Positives = 28/53 (52%)
Query: 7 MVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
M +K F+D KGF IT D GG D+F S I GF L E + V+ +T
Sbjct: 1 MATGTVKWFNDAKGFGFITQDGGGEDVFCHHSAINMDGFRTLQEGQKVEFEVT 53
>CSPC_ECOLI (P36996) Cold shock-like protein cspC (CSP-C)
Length = 68
Score = 37.7 bits (86), Expect = 0.005
Identities = 19/48 (39%), Positives = 30/48 (61%)
Query: 8 VREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
++ ++K F++ KGF IT DG D+F+ S I+ GF LAE + V+
Sbjct: 3 IKGQVKWFNESKGFGFITPADGSKDVFVHFSAIQGNGFKTLAEGQNVE 50
>CSPA_VIBCH (Q9KN00) Cold shock-like protein cspA
Length = 70
Score = 37.7 bits (86), Expect = 0.005
Identities = 19/43 (44%), Positives = 28/43 (64%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F++ KGF I+ D+GG D+F+ I S+GF LAE + V
Sbjct: 9 VKWFNETKGFGFISQDNGGQDVFVHFKSIVSEGFKTLAEGQRV 51
>CSPC_BUCAI (P57407) Cold shock-like protein cspC (CSP-C)
Length = 68
Score = 37.4 bits (85), Expect = 0.006
Identities = 18/48 (37%), Positives = 29/48 (59%)
Query: 8 VREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
++ ++K F++ KGF IT DG D+F+ S I+ GF L E + V+
Sbjct: 3 IKGQVKWFNESKGFGFITPSDGSKDVFVHFSSIQGNGFKTLTEGQNVE 50
>CSP7_STRCL (Q01761) Cold shock-like protein 7.0
Length = 66
Score = 36.2 bits (82), Expect = 0.014
Identities = 20/53 (37%), Positives = 29/53 (53%)
Query: 7 MVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
M +K F+ +KGF I D GG D+F+ S I + GF L E + V+ +T
Sbjct: 1 MATGTVKWFNAEKGFGFIAQDGGGPDVFVHYSAINATGFRSLEENQVVNFDVT 53
>CSPC_SHIFL (Q83RI9) Cold shock-like protein cspC (CSP-C)
Length = 68
Score = 35.4 bits (80), Expect = 0.023
Identities = 18/48 (37%), Positives = 29/48 (59%)
Query: 8 VREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
++ ++K F++ KGF IT DG D+F+ S I+ G LAE + V+
Sbjct: 3 IKGQVKWFNESKGFGFITPADGSKDVFVHFSAIQGNGLKTLAEGQNVE 50
>CSPA_SHEVI (Q9S1B7) Cold shock-like protein cspA
Length = 70
Score = 35.4 bits (80), Expect = 0.023
Identities = 18/43 (41%), Positives = 27/43 (61%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F++ KGF +T D+GG D+F+ I S+GF L E + V
Sbjct: 9 VKWFNEDKGFGFLTQDNGGADVFVHFRAIASEGFKTLDEGQKV 51
>CSPA_PHOPO (Q52287) Major cold-shock protein (Fragment)
Length = 46
Score = 35.0 bits (79), Expect = 0.030
Identities = 18/37 (48%), Positives = 24/37 (64%)
Query: 18 QKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+KGF IT D+GG D+F+ I S+GF LAE + V
Sbjct: 1 EKGFGFITQDNGGADVFVHFRAIASEGFKTLAEGQKV 37
>CSPI_ECOLI (P77605) Cold shock-like protein cspI (CPS-I)
Length = 70
Score = 34.3 bits (77), Expect = 0.051
Identities = 18/44 (40%), Positives = 26/44 (58%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
+K F+ +KGF IT DG D+F+ S I+S F L E + V+
Sbjct: 9 VKWFNPEKGFGFITPKDGSKDVFVHFSAIQSNDFKTLTENQEVE 52
>CSPF_STRCO (P48859) Cold shock protein scoF
Length = 67
Score = 34.3 bits (77), Expect = 0.051
Identities = 20/53 (37%), Positives = 28/53 (52%)
Query: 7 MVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
M +K F+ +KGF I D GG D+F S I +QG+ L E + V +T
Sbjct: 1 MASGTVKWFNSEKGFGFIAQDGGGPDVFAHYSNINAQGYRELQEGQAVTFDIT 53
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,801,228
Number of Sequences: 164201
Number of extensions: 429868
Number of successful extensions: 3133
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 3061
Number of HSP's gapped (non-prelim): 80
length of query: 119
length of database: 59,974,054
effective HSP length: 95
effective length of query: 24
effective length of database: 44,374,959
effective search space: 1064999016
effective search space used: 1064999016
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0089a.1


