

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079a.1
(139 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
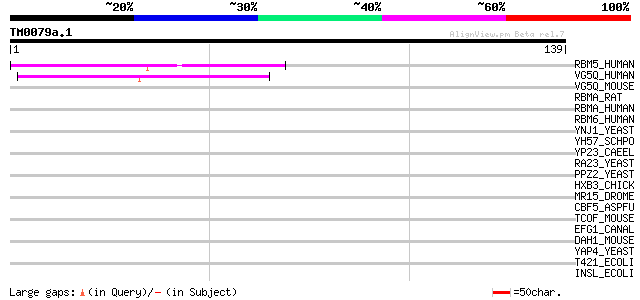
Score E
Sequences producing significant alignments: (bits) Value
RBM5_HUMAN (P52756) RNA-binding protein 5 (RNA binding motif pro... 47 1e-05
VG5Q_HUMAN (Q8N302) Angiogenic factor VG5Q (Vasculogenesis gene ... 42 3e-04
VG5Q_MOUSE (Q7TN31) Angiogenic factor VG5Q (mVG5Q) 40 0.002
RBMA_RAT (P70501) RNA-binding protein 10 (RNA binding motif prot... 39 0.003
RBMA_HUMAN (P98175) RNA-binding protein 10 (RNA binding motif pr... 39 0.003
RBM6_HUMAN (P78332) RNA-binding protein 6 (RNA binding motif pro... 34 0.11
YNJ1_YEAST (P53935) Hypothetical 141.5 kDa protein in YPT53-RHO2... 33 0.19
YH57_SCHPO (O94312) PWWP domain protein C215.07c 33 0.25
YP23_CAEEL (Q09202) Hypothetical protein AH6.3 in chromosome II 32 0.32
RA23_YEAST (P32628) UV excision repair protein RAD23 32 0.42
PPZ2_YEAST (P33329) Serine/threonine protein phosphatase PP-Z2 (... 32 0.42
HXB3_CHICK (P23682) Homeobox protein Hox-B3 (Chox-2.7) 32 0.42
MR15_DROME (Q9Y0I1) MRG15 protein 32 0.55
CBF5_ASPFU (O43102) Centromere/microtubule binding protein CBF5 ... 32 0.55
TCOF_MOUSE (O08784) Treacle protein (Treacher Collins syndrome p... 31 0.72
EFG1_CANAL (P43064) Enhanced filamentous growth protein 31 0.93
DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (Dach1) 31 0.93
YAP4_YEAST (P40917) AP-1 like transcription factor YAP4 (Transcr... 30 1.2
T421_ECOLI (P11901) Transposase for insertion sequence element I... 30 1.2
INSL_ECOLI (P08409) Putative transposase insL for insertion sequ... 30 1.2
>RBM5_HUMAN (P52756) RNA-binding protein 5 (RNA binding motif
protein 5) (Putative tumor suppressor LUCA15) (G15
protein)
Length = 815
Score = 47.0 bits (110), Expect = 1e-05
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 4/72 (5%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y P S++ + S
Sbjct: 462 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYV-PAAESSSHQQSGLP 520
Query: 58 QSNKGNKPHNGP 69
+ +G + P
Sbjct: 521 PAKEGKEKKEKP 532
Score = 36.2 bits (82), Expect = 0.022
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 3/63 (4%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 459 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSHQQSG 518
Query: 63 NKP 65
P
Sbjct: 519 LPP 521
>VG5Q_HUMAN (Q8N302) Angiogenic factor VG5Q (Vasculogenesis gene on
5q) (hVG5Q)
Length = 714
Score = 42.4 bits (98), Expect = 3e-04
Identities = 20/65 (30%), Positives = 31/65 (46%), Gaps = 2/65 (3%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAI--GKWVTQEEAYASPHFASTTKKASSTSQSN 60
F+ S+G+YY N YYDP +G YY + G++ P+ S+TK++
Sbjct: 217 FDHSTGFYYDSENQLYYDPSTGIYYYCDVESGRYQFHSRVDLQPYPTSSTKQSKDKKLKK 276
Query: 61 KGNKP 65
K P
Sbjct: 277 KRKDP 281
Score = 30.0 bits (66), Expect = 1.6
Identities = 10/23 (43%), Positives = 15/23 (64%)
Query: 5 SSSGYYYHKTNGFYYDPKSGFYY 27
S +G+ Y + G Y+D +GFYY
Sbjct: 203 SQTGFSYDENTGLYFDHSTGFYY 225
>VG5Q_MOUSE (Q7TN31) Angiogenic factor VG5Q (mVG5Q)
Length = 711
Score = 39.7 bits (91), Expect = 0.002
Identities = 18/51 (35%), Positives = 27/51 (52%), Gaps = 2/51 (3%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAI--GKWVTQEEAYASPHFASTTK 51
F+ S+G+YY N YYDP +G YY + G++ P+ S+TK
Sbjct: 217 FDHSTGFYYDSENQLYYDPSTGIYYYCDVESGRYQFHSRVDLQPYQTSSTK 267
Score = 33.5 bits (75), Expect = 0.14
Identities = 16/62 (25%), Positives = 28/62 (44%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
++ S+G Y+ + GFYYD ++ YY + G + + F S + S K
Sbjct: 209 YDESTGLYFDHSTGFYYDSENQLYYDPSTGIYYYCDVESGRYQFHSRVDLQPYQTSSTKP 268
Query: 63 NK 64
N+
Sbjct: 269 NR 270
Score = 30.4 bits (67), Expect = 1.2
Identities = 10/23 (43%), Positives = 16/23 (69%)
Query: 5 SSSGYYYHKTNGFYYDPKSGFYY 27
S +G+ Y ++ G Y+D +GFYY
Sbjct: 203 SQTGFTYDESTGLYFDHSTGFYY 225
>RBMA_RAT (P70501) RNA-binding protein 10 (RNA binding motif protein
10) (S1-1 protein)
Length = 852
Score = 39.3 bits (90), Expect = 0.003
Identities = 22/69 (31%), Positives = 31/69 (44%), Gaps = 5/69 (7%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAY--ASPHFASTTKKASS 55
+ ++ +SGYYY G YYDP S +YY S W + Y A A K +
Sbjct: 494 YQYDETSGYYYDPQTGLYYDPNSQYYYNAQSQQYLYWDGERRTYIPALEQSADGHKDTGA 553
Query: 56 TSQSNKGNK 64
+S+ K K
Sbjct: 554 SSKEGKEKK 562
Score = 35.0 bits (79), Expect = 0.050
Identities = 12/21 (57%), Positives = 17/21 (80%)
Query: 7 SGYYYHKTNGFYYDPKSGFYY 27
S Y Y +T+G+YYDP++G YY
Sbjct: 492 STYQYDETSGYYYDPQTGLYY 512
>RBMA_HUMAN (P98175) RNA-binding protein 10 (RNA binding motif
protein 10) (DXS8237E)
Length = 929
Score = 39.3 bits (90), Expect = 0.003
Identities = 22/69 (31%), Positives = 31/69 (44%), Gaps = 5/69 (7%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAY--ASPHFASTTKKASS 55
+ ++ +SGYYY G YYDP S +YY S W + Y A A K+ +
Sbjct: 571 YQYDETSGYYYDPQTGLYYDPNSQYYYNAQSQQYLYWDGERRTYVPALEQSADGHKETGA 630
Query: 56 TSQSNKGNK 64
S+ K K
Sbjct: 631 PSKEGKEKK 639
Score = 35.0 bits (79), Expect = 0.050
Identities = 12/21 (57%), Positives = 17/21 (80%)
Query: 7 SGYYYHKTNGFYYDPKSGFYY 27
S Y Y +T+G+YYDP++G YY
Sbjct: 569 STYQYDETSGYYYDPQTGLYY 589
>RBM6_HUMAN (P78332) RNA-binding protein 6 (RNA binding motif
protein 6) (RNA-binding protein DEF-3) (Lung cancer
antigen NY-LU-12) (Protein G16)
Length = 1123
Score = 33.9 bits (76), Expect = 0.11
Identities = 22/73 (30%), Positives = 28/73 (38%), Gaps = 9/73 (12%)
Query: 5 SSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPH----FASTTKKASSTSQSN 60
SS Y Y G+YYDP +G YY TQ+E Y K+ TSQ
Sbjct: 786 SSDCYIYDSATGYYYDPLAGTYYDPN-----TQQEVYVPQDPGLPEEEEIKEKKPTSQGK 840
Query: 61 KGNKPHNGPLPGR 73
+K G+
Sbjct: 841 SSSKKEMSKRDGK 853
Score = 30.4 bits (67), Expect = 1.2
Identities = 17/62 (27%), Positives = 26/62 (41%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
++S++GYYY G YYDP + + +EE S K +S S +
Sbjct: 792 YDSATGYYYDPLAGTYYDPNTQQEVYVPQDPGLPEEEEIKEKKPTSQGKSSSKKEMSKRD 851
Query: 63 NK 64
K
Sbjct: 852 GK 853
>YNJ1_YEAST (P53935) Hypothetical 141.5 kDa protein in YPT53-RHO2
intergenic region
Length = 1240
Score = 33.1 bits (74), Expect = 0.19
Identities = 15/35 (42%), Positives = 24/35 (67%)
Query: 96 KRKRPGEKPKVVSEEEKSALKAREAARKRVQEREK 130
KRKR EK ++ E E+ ++ REA RK+V+E ++
Sbjct: 667 KRKREEEKERLKKELEEREMRRREAQRKKVEEAKR 701
>YH57_SCHPO (O94312) PWWP domain protein C215.07c
Length = 568
Score = 32.7 bits (73), Expect = 0.25
Identities = 26/86 (30%), Positives = 34/86 (39%), Gaps = 12/86 (13%)
Query: 53 ASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEK 112
ASS SN K T ++P R+ A+ KR E K EEE
Sbjct: 283 ASSNVSSNPAKK------------TRVSPRRSTAASKKKSPSSKRASSDEIEKDKEEEEG 330
Query: 113 SALKAREAARKRVQEREKSLLGLYSK 138
S + A++ RE+SLL L K
Sbjct: 331 SVANEEDVAKRTFHSREQSLLFLRHK 356
>YP23_CAEEL (Q09202) Hypothetical protein AH6.3 in chromosome II
Length = 230
Score = 32.3 bits (72), Expect = 0.32
Identities = 21/83 (25%), Positives = 36/83 (43%), Gaps = 5/83 (6%)
Query: 54 SSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSL-----SLGKRKRPGEKPKVVS 108
+++ +SNK KP G + S K L + ++P K K
Sbjct: 85 ANSKKSNKSKKPEKGSKKSKKSEKSKKKKTEEKVMSEDKPERPDKLTEEEKPDAKEKKEI 144
Query: 109 EEEKSALKAREAARKRVQEREKS 131
E+EK+ K + +++ QE+EKS
Sbjct: 145 EKEKTKEKEKTKEKEKTQEKEKS 167
>RA23_YEAST (P32628) UV excision repair protein RAD23
Length = 398
Score = 32.0 bits (71), Expect = 0.42
Identities = 18/69 (26%), Positives = 35/69 (50%), Gaps = 1/69 (1%)
Query: 46 FASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPK 105
F + KK++ T + P + PGR +T +P+ + AAP++ + + +P E+
Sbjct: 70 FMVSQKKSTKTKVTEPPIAPESATTPGRENSTEASPSTDASAAPAA-TAPEGSQPQEEQT 128
Query: 106 VVSEEEKSA 114
+E +SA
Sbjct: 129 ATTERTESA 137
>PPZ2_YEAST (P33329) Serine/threonine protein phosphatase PP-Z2 (EC
3.1.3.16)
Length = 709
Score = 32.0 bits (71), Expect = 0.42
Identities = 30/115 (26%), Positives = 49/115 (42%), Gaps = 16/115 (13%)
Query: 5 SSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNK 64
+ S Y+ +++N KS F T AY++P + +K SS + G+
Sbjct: 240 NGSDYFSNRSNSHASSRKSSF--------GSTGNTAYSTPLHSPALRKMSSRDNDDSGDN 291
Query: 65 PHNGPLPGRVVTTSLNPTRNV-KAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAR 118
+ GR TS P N+ K +PS+ S KR+ P + + S+L R
Sbjct: 292 VN-----GR--GTSPIPNLNIDKPSPSASSASKREYLSAYPTLAHRDSSSSLSPR 339
>HXB3_CHICK (P23682) Homeobox protein Hox-B3 (Chox-2.7)
Length = 399
Score = 32.0 bits (71), Expect = 0.42
Identities = 31/110 (28%), Positives = 46/110 (41%), Gaps = 6/110 (5%)
Query: 3 FESSS---GYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQS 59
+ESS+ GY Y TNGF Y+ + + + Q A + +TT+ A S +
Sbjct: 7 YESSTLFGGYSYGSTNGFGYEGPQQPFQPGSHVENDFQRSACSLQSLGNTTQHAKSKDLN 66
Query: 60 NKGNKPH--NGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVV 107
+P P V+ NPT N + +S+ G K P KP V
Sbjct: 67 GSCMRPSLPQEHHPPPTVSPPPNPTTN-STSSNSIPSGSAKVPRVKPTSV 115
>MR15_DROME (Q9Y0I1) MRG15 protein
Length = 424
Score = 31.6 bits (70), Expect = 0.55
Identities = 30/104 (28%), Positives = 41/104 (38%), Gaps = 18/104 (17%)
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLP------GRVVTTSLNPTRN----------VKAAPS 90
AST K S+TSQS + P P G TT+ N T N ++ PS
Sbjct: 118 ASTPSKDSNTSQSTASSTPTTSAGPGSKSEAGSTGTTTTNSTANSTTSRAHRKSTQSTPS 177
Query: 91 SLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLG 134
+ G E P E+ A + +KR+ E+ SL G
Sbjct: 178 TARPGTPSDKKEDPAAAETTEEEGPVAPK--KKRMSEQRPSLTG 219
>CBF5_ASPFU (O43102) Centromere/microtubule binding protein CBF5
(Centromere-binding factor 5) (Small nucleolar RNP
protein CBF5) (H/ACA snoRNP protein CBF5)
Length = 487
Score = 31.6 bits (70), Expect = 0.55
Identities = 23/110 (20%), Positives = 48/110 (42%), Gaps = 10/110 (9%)
Query: 22 KSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNP 81
K G D KW T+ + Y++P AS + ST++ + +TT +
Sbjct: 377 KYGRANEDTPAKWKTEYKDYSAPEEASAHAASESTAKEDVA----------AALTTEQDE 426
Query: 82 TRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKS 131
+ + + K ++ ++ + + EE++ K ++ +K +ER KS
Sbjct: 427 APSSPQSKMDVDETKEEKKRKRHEGETAEERAERKRKKKEKKEKKERRKS 476
>TCOF_MOUSE (O08784) Treacle protein (Treacher Collins syndrome
protein homolog)
Length = 1320
Score = 31.2 bits (69), Expect = 0.72
Identities = 27/93 (29%), Positives = 41/93 (44%), Gaps = 14/93 (15%)
Query: 43 SPHFASTTKKASST-SQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG 101
SP A T KA S + + G K P P + K P LG + G
Sbjct: 1216 SPEKAPMTSKAKSKLDKGSAGGKGKGSPGP-----------QGAKEKPDGELLGIKLESG 1264
Query: 102 EK--PKVVSEEEKSALKAREAARKRVQEREKSL 132
E+ PK S+++KS K ++ +K ++ +KSL
Sbjct: 1265 EQSDPKSKSKKKKSLKKKKDKEKKEKKKGKKSL 1297
>EFG1_CANAL (P43064) Enhanced filamentous growth protein
Length = 552
Score = 30.8 bits (68), Expect = 0.93
Identities = 32/123 (26%), Positives = 56/123 (45%), Gaps = 19/123 (15%)
Query: 9 YYYHKTNGFYYDPKS--GFYYSDAIGKWVTQEEAY----ASPHFASTTKKASSTSQSNKG 62
Y Y+ N + Y + G YY I ++Q + ++ ++ S A S S G
Sbjct: 94 YDYNTYNRYQYPAATSQGNYYQQTIPNQLSQPQPQHYNGSNRNYTSAPSGAPIPSNSTSG 153
Query: 63 NKPHNGPLPGRV-------VTTSLNPT--RNVKAAPSSLSLGKRKRPGEKPKVVS---EE 110
PLPG+ V+T PT ++ A S+ ++G+ + PG +P+V + E+
Sbjct: 154 PS-QQPPLPGQQAVPIPPHVSTMQQPTPVQDTLNASSTSTVGQFQPPGIRPRVTTTMWED 212
Query: 111 EKS 113
EK+
Sbjct: 213 EKT 215
>DAH1_MOUSE (Q9QYB2) Dachshund homolog 1 (Dach1)
Length = 751
Score = 30.8 bits (68), Expect = 0.93
Identities = 14/43 (32%), Positives = 26/43 (59%)
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRN 84
+S +S++ +SS+S S+ + GPLPG+ V ++ +P N
Sbjct: 132 SSSSSSSSSSSSSSSSSSSSSSSSSCGPLPGKPVYSTPSPVEN 174
>YAP4_YEAST (P40917) AP-1 like transcription factor YAP4
(Transcription activator CIN5) (Chromosome instability
protein 5)
Length = 295
Score = 30.4 bits (67), Expect = 1.2
Identities = 37/140 (26%), Positives = 64/140 (45%), Gaps = 16/140 (11%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQS--- 59
FESS Y + F P + ++ S++ T++ + A+ + S ++ S S S
Sbjct: 165 FESS----YTTASTFTSQPAASYFPSNSTP--ATRKNS-ATTNLPSEERRRVSVSLSEQV 217
Query: 60 -NKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAR 118
N+G + +N G+++ + P RN K A + S K R + + + EEKS K
Sbjct: 218 FNEGERYNND---GQLIGKTGKPLRNTKRAAQNRSAQKAFRQRREKYIKNLEEKS--KLF 272
Query: 119 EAARKRVQEREKSLLGLYSK 138
+ K E +K + L SK
Sbjct: 273 DGLMKENSELKKMIESLKSK 292
>T421_ECOLI (P11901) Transposase for insertion sequence element
IS421
Length = 371
Score = 30.4 bits (67), Expect = 1.2
Identities = 27/93 (29%), Positives = 40/93 (42%), Gaps = 17/93 (18%)
Query: 60 NKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVS----------- 108
N GNK P P R++ SL P + + + LS +RK + + +
Sbjct: 233 NSGNKKAGAPFPARLIAVSLPPEKALISKTRLLSENRRKGRVVQAETLEAAGHVLLLTSL 292
Query: 109 -EEEKSALKAREAARKRVQ-----EREKSLLGL 135
E+E SA + + R R Q +R KSLL L
Sbjct: 293 PEDEYSAEQVADCYRLRWQIELAFKRLKSLLHL 325
>INSL_ECOLI (P08409) Putative transposase insL for insertion
sequence element IS186A/B/C
Length = 370
Score = 30.4 bits (67), Expect = 1.2
Identities = 27/93 (29%), Positives = 40/93 (42%), Gaps = 17/93 (18%)
Query: 60 NKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVS----------- 108
N GNK P P R++ SL P + + + LS +RK + + +
Sbjct: 232 NSGNKKAGAPFPARLIAVSLPPEKALISKTRLLSENRRKGRVVQAETLEAAGHVLLLTSL 291
Query: 109 -EEEKSALKAREAARKRVQ-----EREKSLLGL 135
E+E SA + + R R Q +R KSLL L
Sbjct: 292 PEDEYSAEQVADCYRLRWQIELAFKRLKSLLHL 324
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.306 0.125 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,152,817
Number of Sequences: 164201
Number of extensions: 669901
Number of successful extensions: 2815
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 2713
Number of HSP's gapped (non-prelim): 145
length of query: 139
length of database: 59,974,054
effective HSP length: 99
effective length of query: 40
effective length of database: 43,718,155
effective search space: 1748726200
effective search space used: 1748726200
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0079a.1


