

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0076c.7
(656 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
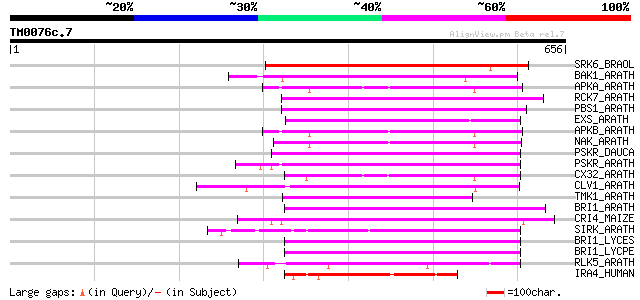
Sequences producing significant alignments: (bits) Value
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 265 3e-70
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 223 2e-57
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 206 2e-52
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 203 1e-51
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 203 1e-51
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 203 1e-51
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 201 5e-51
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 199 2e-50
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 196 2e-49
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 196 2e-49
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 191 5e-48
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 191 7e-48
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 185 3e-46
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 185 3e-46
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 184 5e-46
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 183 1e-45
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 181 4e-45
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 180 9e-45
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 175 3e-43
IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (... 164 9e-40
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 265 bits (677), Expect = 3e-70
Identities = 145/322 (45%), Positives = 199/322 (61%), Gaps = 12/322 (3%)
Query: 303 RRQF-GAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKR 361
+R+F G + F E L + E + KAT F NKLG+GG G VYKG L DG +A+KR
Sbjct: 499 KREFSGEYKFEELELPL-IEMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKR 557
Query: 362 LSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRR 421
LS + Q D F NEV LI LQH NLV++LGC I G E +L+YEY+ NLSL +L +
Sbjct: 558 LSKTSVQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKT 617
Query: 422 NLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARL 481
+L W R I G A GL YLH++S+ +IIHRD+K+SNILLD N PKI+DFG+AR+
Sbjct: 618 RRSKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARI 677
Query: 482 FPEDQSQIST-AICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSF--VQN 538
F D+++ +T + GT GYM+PEY + G +EK+D++SFGV+V+E++SGK F +
Sbjct: 678 FERDETEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDY 737
Query: 539 SCSILHMVWSLYGSNRLCDIVDPILEGN-------YPAEEACKLLKIGLLCTQASAELRP 591
+L VWS + R +IVDP++ + + +E K ++IGLLC Q AE RP
Sbjct: 738 ENDLLSYVWSRWKEGRALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRP 797
Query: 592 PMSVVVKMINNIHKIADPTQPP 613
MS VV M + +PP
Sbjct: 798 AMSSVVWMFGSEATEIPQPKPP 819
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 223 bits (567), Expect = 2e-57
Identities = 138/355 (38%), Positives = 202/355 (56%), Gaps = 18/355 (5%)
Query: 259 PPGTKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAF-LFSENNSK 317
PP GS ++ +A AA A LL I RR++ Q F + +E + +
Sbjct: 214 PPSPAGSNRITGAIAGGVAAGAALLFAVPAIALAW-----WRRKKPQDHFFDVPAEEDPE 268
Query: 318 LNM------PYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWAD 371
+++ L+ A++ F + N LG GG G VYKG L DGT VA+KRL TQ +
Sbjct: 269 VHLGQLKRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGE 328
Query: 372 -HFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQ-LTWE 429
F EV +I H+NL++L G +T E LLVY Y+ N S+ L R Q L W
Sbjct: 329 LQFQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWP 388
Query: 430 VRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQI 489
R +I LG+A GLAYLH+ KIIHRD+K +NILLD+ F + DFGLA+L + +
Sbjct: 389 KRQRIALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHV 448
Query: 490 STAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQ----NSCSILHM 545
+TA+ GT+G++APEY+ GK +EK D++ +GV+++EL++G+ + + +L
Sbjct: 449 TTAVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDW 508
Query: 546 VWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
V L +L +VD L+GNY EE +L+++ LLCTQ+S RP MS VV+M+
Sbjct: 509 VKGLLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRML 563
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 206 bits (523), Expect = 2e-52
Identities = 125/323 (38%), Positives = 184/323 (56%), Gaps = 20/323 (6%)
Query: 300 RRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPD------ 353
R R G L S N + + L+ AT F + LGEGG G V+KG + +
Sbjct: 38 RPSPRTEGEILQSPNLKSFS--FAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTAS 95
Query: 354 ----GTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVP 409
G +A+K+L+ + Q + EVN + H++LVKL+G + LLVYE++P
Sbjct: 96 RPGTGLVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMP 155
Query: 410 NLSLHDHLSVRRNL--QQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDD 467
SL +HL RR L Q L+W++R K+ LG A+GLA+LH S+ ++I+RD K SNILLD
Sbjct: 156 RGSLENHL-FRRGLYFQPLSWKLRLKVALGAAKGLAFLHS-SETRVIYRDFKTSNILLDS 213
Query: 468 NFTPKIADFGLARLFP-EDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIEL 526
+ K++DFGLA+ P D+S +ST + GT GY APEY+ G LT K+D+YSFGV+++EL
Sbjct: 214 EYNAKLSDFGLAKDGPIGDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLEL 273
Query: 527 LSGKSRTSFVQNSCSILHMVWS---LYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCT 583
LSG+ + S + W+ L ++ ++D L+ Y EEACK+ + L C
Sbjct: 274 LSGRRAVDKNRPSGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCL 333
Query: 584 QASAELRPPMSVVVKMINNIHKI 606
+LRP MS VV + +I +
Sbjct: 334 TTEIKLRPNMSEVVSHLEHIQSL 356
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 203 bits (517), Expect = 1e-51
Identities = 124/316 (39%), Positives = 177/316 (55%), Gaps = 7/316 (2%)
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPD-GTTVAIKRLSFNTTQWADHFFNEVNLI 380
++ L +AT F LGEGG G V+KG + VAIK+L N Q F EV +
Sbjct: 93 FQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFVVEVLTL 152
Query: 381 KDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQ-LTWEVRHKIILGTA 439
H NLVKL+G G + LLVYEY+P SL DHL V + ++ L W R KI G A
Sbjct: 153 SLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKIAAGAA 212
Query: 440 EGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPE-DQSQISTAICGTLG 498
GL YLH+ +I+RD+K SNILL +++ PK++DFGLA++ P D++ +ST + GT G
Sbjct: 213 RGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTRVMGTYG 272
Query: 499 YMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWS--LYGSNR-L 555
Y AP+Y + G+LT K+DIYSFGV+++EL++G+ + + W+ L+ R
Sbjct: 273 YCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARPLFKDRRNF 332
Query: 556 CDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIA-DPTQPPF 614
+VDP+L+G YP + L I +C Q +RP +S VV +N + DP P
Sbjct: 333 PKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLALNFLASSKYDPNSPSS 392
Query: 615 LNFGSGEFSRSSLPEE 630
+ + F R EE
Sbjct: 393 SSGKNPSFHRDRDDEE 408
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 203 bits (517), Expect = 1e-51
Identities = 118/297 (39%), Positives = 174/297 (57%), Gaps = 7/297 (2%)
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPD-GTTVAIKRLSFNTTQWADHFFNEVNLI 380
+ L AT FH LGEGG G VYKG L G VA+K+L N Q F EV ++
Sbjct: 76 FRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVLML 135
Query: 381 KDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHL-SVRRNLQQLTWEVRHKIILGTA 439
L H NLV L+G G + LLVYE++P SL DHL + + + L W +R KI G A
Sbjct: 136 SLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAGAA 195
Query: 440 EGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPE-DQSQISTAICGTLG 498
+GL +LH+++ +I+RD K SNILLD+ F PK++DFGLA+L P D+S +ST + GT G
Sbjct: 196 KGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGTYG 255
Query: 499 YMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSR--TSFVQNSCSILHMVWSLYGSNR-L 555
Y APEY + G+LT K+D+YSFGV+ +EL++G+ + +++ L+ R
Sbjct: 256 YCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFNDRRKF 315
Query: 556 CDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI-HKIADPTQ 611
+ DP L+G +P + L + +C Q A RP ++ VV ++ + ++ DP++
Sbjct: 316 IKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTALSYLANQAYDPSK 372
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 203 bits (517), Expect = 1e-51
Identities = 121/282 (42%), Positives = 165/282 (57%), Gaps = 6/282 (2%)
Query: 327 KATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHK 386
+AT++F N +G+GG G+VYK LP TVA+K+LS TQ F E+ + ++H
Sbjct: 912 EATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEMETLGKVKHP 971
Query: 387 NLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRN-LQQLTWEVRHKIILGTAEGLAYL 445
NLV LLG E LLVYEY+ N SL L + L+ L W R KI +G A GLA+L
Sbjct: 972 NLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARGLAFL 1031
Query: 446 HEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYI 505
H IIHRDIK SNILLD +F PK+ADFGLARL +S +ST I GT GY+ PEY
Sbjct: 1032 HHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTVIAGTFGYIPPEYG 1091
Query: 506 VLGKLTEKADIYSFGVLVIELLSGKSRT--SFVQNSCSILHMVWSLYGSN--RLCDIVDP 561
+ T K D+YSFGV+++EL++GK T F ++ L + W++ N + D++DP
Sbjct: 1092 QSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNL-VGWAIQKINQGKAVDVIDP 1150
Query: 562 ILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
+L +LL+I +LC + RP M V+K + I
Sbjct: 1151 LLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKALKEI 1192
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 201 bits (511), Expect = 5e-51
Identities = 122/322 (37%), Positives = 179/322 (54%), Gaps = 18/322 (5%)
Query: 300 RRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPD------ 353
R R G L S N + L+ AT F + LGEGG GSV+KG + +
Sbjct: 39 RTNPRTEGEILQSPNLKSFT--FAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTAS 96
Query: 354 ----GTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVP 409
G +A+K+L+ + Q + EVN + H NLVKL+G + LLVYE++P
Sbjct: 97 KPGTGVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMP 156
Query: 410 NLSLHDHLSVRRN-LQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDN 468
SL +HL R + Q L+W +R K+ LG A+GLA+LH ++ +I+RD K SNILLD
Sbjct: 157 RGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLHN-AETSVIYRDFKTSNILLDSE 215
Query: 469 FTPKIADFGLARLFPE-DQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELL 527
+ K++DFGLA+ P D+S +ST I GT GY APEY+ G LT K+D+YS+GV+++E+L
Sbjct: 216 YNAKLSDFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVL 275
Query: 528 SGKSRTSFVQNSCSILHMVWS---LYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQ 584
SG+ + + W+ L +L ++D L+ Y EEACK+ + L C
Sbjct: 276 SGRRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLT 335
Query: 585 ASAELRPPMSVVVKMINNIHKI 606
+LRP M+ VV + +I +
Sbjct: 336 FEIKLRPNMNEVVSHLEHIQTL 357
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 199 bits (506), Expect = 2e-50
Identities = 119/307 (38%), Positives = 171/307 (54%), Gaps = 16/307 (5%)
Query: 313 ENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPD----------GTTVAIKRL 362
+N + N L+ AT F + +GEGG G V+KG + + G +A+KRL
Sbjct: 49 QNANLKNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRL 108
Query: 363 SFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRN 422
+ Q + E+N + L H NLVKL+G + LLVYE++ SL +HL R
Sbjct: 109 NQEGFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGT 168
Query: 423 LQQ-LTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARL 481
Q L+W R ++ LG A GLA+LH +Q ++I+RD K SNILLD N+ K++DFGLAR
Sbjct: 169 FYQPLSWNTRVRMALGAARGLAFLHN-AQPQVIYRDFKASNILLDSNYNAKLSDFGLARD 227
Query: 482 FPE-DQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSC 540
P D S +ST + GT GY APEY+ G L+ K+D+YSFGV+++ELLSG+ Q
Sbjct: 228 GPMGDNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVG 287
Query: 541 SILHMVWS---LYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVV 597
+ W+ L RL ++DP L+G Y A K+ + L C A+ RP M+ +V
Sbjct: 288 EHNLVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEIV 347
Query: 598 KMINNIH 604
K + +H
Sbjct: 348 KTMEELH 354
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 196 bits (498), Expect = 2e-49
Identities = 119/297 (40%), Positives = 164/297 (55%), Gaps = 3/297 (1%)
Query: 310 LFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQW 369
LF +S + + + K+T+ F+ +N +G GG G VYK LPDGT VAIKRLS +T Q
Sbjct: 721 LFHNKDSNNELSLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQM 780
Query: 370 ADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRN-LQQLTW 428
F EV + QH NLV LLG + LL+Y Y+ N SL L + + L W
Sbjct: 781 DREFQAEVETLSRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDW 840
Query: 429 EVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQ 488
+ R +I G AEGLAYLH+ + I+HRDIK SNILL D F +ADFGLARL +
Sbjct: 841 KTRLRIARGAAEGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTH 900
Query: 489 ISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFV--QNSCSILHMV 546
++T + GTLGY+ PEY T K D+YSFGV+++ELL+G+ + S ++ V
Sbjct: 901 VTTDLVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWV 960
Query: 547 WSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
+ R +I DP + AEE +L+I C + + RP +V + NI
Sbjct: 961 LQMKTEKRESEIFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLENI 1017
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 196 bits (498), Expect = 2e-49
Identities = 125/353 (35%), Positives = 189/353 (53%), Gaps = 19/353 (5%)
Query: 268 VGIIVAESFAAVATLLIVATIIFFVRK-----------NVLRQRRERRQFGA---FLFSE 313
+G+ + +F +V L +++ I+ R+ + R+E + G+ LF
Sbjct: 658 IGMAIGIAFGSVFLLTLLSLIVLRARRRSGEVDPEIEESESMNRKELGEIGSKLVVLFQS 717
Query: 314 NNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHF 373
N+ +L+ Y+ L +TN F +N +G GG G VYK LPDG VAIK+LS + Q F
Sbjct: 718 NDKELS--YDDLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQIEREF 775
Query: 374 FNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLT-WEVRH 432
EV + QH NLV L G + LL+Y Y+ N SL L R + L W+ R
Sbjct: 776 EAEVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRL 835
Query: 433 KIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTA 492
+I G A+GL YLHE I+HRDIK SNILLD+NF +ADFGLARL ++ +ST
Sbjct: 836 RIAQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTD 895
Query: 493 ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQ-NSC-SILHMVWSLY 550
+ GTLGY+ PEY T K D+YSFGV+++ELL+ K + C ++ V +
Sbjct: 896 LVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMK 955
Query: 551 GSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
+R ++ DP++ +E ++L+I LC + + RP +V ++++
Sbjct: 956 HESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVSWLDDV 1008
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 191 bits (485), Expect = 5e-48
Identities = 115/293 (39%), Positives = 173/293 (58%), Gaps = 17/293 (5%)
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKG----------ALPDGTTVAIKRLSFNTTQWADHFF 374
L+ AT F + LG+GG G VY+G + G VAIKRL+ + Q +
Sbjct: 79 LKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQGFAEWR 138
Query: 375 NEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKI 434
+EVN + L H+NLVKLLG E LLVYE++P SL HL RRN W++R KI
Sbjct: 139 SEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHL-FRRN-DPFPWDLRIKI 196
Query: 435 ILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQ-SQISTAI 493
++G A GLA+LH Q ++I+RD K SNILLD N+ K++DFGLA+L P D+ S ++T I
Sbjct: 197 VIGAARGLAFLH-SLQREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPADEKSHVTTRI 255
Query: 494 CGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVW---SLY 550
GT GY APEY+ G L K+D+++FGV+++E+++G + + + + W L
Sbjct: 256 MGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQESLVDWLRPELS 315
Query: 551 GSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
+R+ I+D ++G Y + A ++ +I L C + + RP M VV+++ +I
Sbjct: 316 NKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVLEHI 368
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 191 bits (484), Expect = 7e-48
Identities = 130/399 (32%), Positives = 204/399 (50%), Gaps = 23/399 (5%)
Query: 221 AVTRIDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNIAPPGTKGSTKVGIIVAESFAA-- 278
++T +D GRV G +L F+ F N+ P T+ G + A
Sbjct: 577 SLTTLDLSFNDLSGRVPLGGQFLVFNETSFAGNTYLCLPHRVSCPTRPGQTSDHNHTALF 636
Query: 279 -----VATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYE-VLEKATNYF 332
V T++ T + + + + +++ Q KL+ E VLE
Sbjct: 637 SPSRIVITVIAAITGLILISVAIRQMNKKKNQKSLAWKLTAFQKLDFKSEDVLE----CL 692
Query: 333 HDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFN-EVNLIKDLQHKNLVKL 391
+ N +G+GG+G VY+G++P+ VAIKRL T +DH F E+ + ++H+++V+L
Sbjct: 693 KEENIIGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRL 752
Query: 392 LGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQL 451
LG +LL+YEY+PN SL + L + L WE RH++ + A+GL YLH +
Sbjct: 753 LGYVANKDTNLLLYEYMPNGSLGELLHGSKG-GHLQWETRHRVAVEAAKGLCYLHHDCSP 811
Query: 452 KIIHRDIKLSNILLDDNFTPKIADFGLARLFPED-QSQISTAICGTLGYMAPEYIVLGKL 510
I+HRD+K +NILLD +F +ADFGLA+ + S+ ++I G+ GY+APEY K+
Sbjct: 812 LILHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLKV 871
Query: 511 TEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSL-------YGSNRLCDIVDPIL 563
EK+D+YSFGV+++EL++GK I+ V + + + IVDP L
Sbjct: 872 DEKSDVYSFGVVLLELIAGKKPVGEFGEGVDIVRWVRNTEEEITQPSDAAIVVAIVDPRL 931
Query: 564 EGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINN 602
G YP + KI ++C + A RP M VV M+ N
Sbjct: 932 TG-YPLTSVIHVFKIAMMCVEEEAAARPTMREVVHMLTN 969
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 185 bits (470), Expect = 3e-46
Identities = 101/229 (44%), Positives = 136/229 (59%), Gaps = 4/229 (1%)
Query: 323 EVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWAD--HFFNEVNLI 380
+VL TN F N LG GG G VYKG L DGT +A+KR+ F +E+ ++
Sbjct: 579 QVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAVL 638
Query: 381 KDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSV--RRNLQQLTWEVRHKIILGT 438
++H++LV LLG + G E LLVYEY+P +L HL L+ L W+ R + L
Sbjct: 639 TKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLALDV 698
Query: 439 AEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLG 498
A G+ YLH + IHRD+K SNILL D+ K+ADFGL RL PE + I T I GT G
Sbjct: 699 ARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETRIAGTFG 758
Query: 499 YMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVW 547
Y+APEY V G++T K D+YSFGV+++EL++G+ Q SI + W
Sbjct: 759 YLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSW 807
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 185 bits (470), Expect = 3e-46
Identities = 118/314 (37%), Positives = 171/314 (53%), Gaps = 5/314 (1%)
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
L +ATN FH+ + +G GG G VYK L DG+ VAIK+L + Q F E+ I ++
Sbjct: 876 LLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQGDREFMAEMETIGKIK 935
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHL-SVRRNLQQLTWEVRHKIILGTAEGLA 443
H+NLV LLG G E LLVYE++ SL D L ++ +L W R KI +G+A GLA
Sbjct: 936 HRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIAIGSARGLA 995
Query: 444 YLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIS-TAICGTLGYMAP 502
+LH IIHRD+K SN+LLD+N +++DFG+ARL + +S + + GT GY+ P
Sbjct: 996 FLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPP 1055
Query: 503 EYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVDPI 562
EY + + K D+YS+GV+++ELL+GK T + L + R+ D+ DP
Sbjct: 1056 EYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSPDFGDNNLVGWVKQHAKLRISDVFDPE 1115
Query: 563 LEGNYPAEEACKL--LKIGLLCTQASAELRPPMSVVVKMINNIHKIAD-PTQPPFLNFGS 619
L PA E L LK+ + C A RP M V+ M I + +Q +
Sbjct: 1116 LMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFKEIQAGSGIDSQSTIRSIED 1175
Query: 620 GEFSRSSLPEESLQ 633
G FS + + S++
Sbjct: 1176 GGFSTIEMVDMSIK 1189
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 184 bits (468), Expect = 5e-46
Identities = 131/405 (32%), Positives = 201/405 (49%), Gaps = 31/405 (7%)
Query: 270 IIVAES-FAAVATLLIVATIIFFVRKNVLRQRRERRQFG-----AFLFSENNSKLN---- 319
I VAE FA V L + T +VR + + R+ A+ F ++N K+
Sbjct: 424 IFVAEIVFAVVLVLSVSVTTCLYVRHKLRHCQCSNRELRLAKSTAYSFRKDNMKIQPDME 483
Query: 320 ---------MPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRL--SFNTTQ 368
YE LE+AT F + +++G+G V+KG L DGT VA+KR + + +
Sbjct: 484 DLKIRRAQEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKK 543
Query: 369 WADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRR-NLQQ-L 426
+ F NE++L+ L H +L+ LLG G E LLVYE++ + SL+ HL + NL++ L
Sbjct: 544 SSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRL 603
Query: 427 TWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQ 486
W R I + A G+ YLH + +IHRDIK SNIL+D++ ++ADFGL+ L P D
Sbjct: 604 NWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADS 663
Query: 487 -SQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHM 545
+ +S GTLGY+ PEY L LT K+D+YSFGV+++E+LSG+ +I+
Sbjct: 664 GTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEGNIVEW 723
Query: 546 VWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHK 605
L + + I+DP+L E K+ + C + + RP M V + +
Sbjct: 724 AVPLIKAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTALEHALA 783
Query: 606 -------IADPTQPPFLNFGSGEFSRSSLPEESLQPGSNTQSSGD 643
I P P + GS + S + N + G+
Sbjct: 784 LLMGSPCIEQPILPTEVVLGSSRMHKVSQMSSNHSCSENELADGE 828
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 183 bits (465), Expect = 1e-45
Identities = 127/374 (33%), Positives = 194/374 (50%), Gaps = 18/374 (4%)
Query: 235 RVLNAGCYLRFSTDK---FYDNSSNIAPPGTKGSTKVGIIVAESFAAVATLLIVATIIFF 291
R N LRF + D+ SN TK K G I+ + +L+ A +F
Sbjct: 485 RSKNGSLSLRFGRNPDLCLSDSCSN-----TKKKNKNGYIIPLVVVGIIVVLLTALALF- 538
Query: 292 VRKNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGAL 351
+++++R G +K Y + TN F +G+GG G VY G +
Sbjct: 539 ---RRFKKKQQRGTLGERNGPLKTAKRYFKYSEVVNITNNFE--RVIGKGGFGKVYHGVI 593
Query: 352 PDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNL 411
+G VA+K LS + Q F EV+L+ + H NL L+G +L+YEY+ N
Sbjct: 594 -NGEQVAVKVLSEESAQGYKEFRAEVDLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANE 652
Query: 412 SLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTP 471
+L D+L+ +R+ L+WE R KI L A+GL YLH + I+HRD+K +NILL++
Sbjct: 653 NLGDYLAGKRSFI-LSWEERLKISLDAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQA 711
Query: 472 KIADFGLARLFP-EDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK 530
K+ADFGL+R F E QIST + G++GY+ PEY ++ EK+D+YS GV+++E+++G+
Sbjct: 712 KMADFGLSRSFSVEGSGQISTVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQ 771
Query: 531 -SRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAEL 589
+ S I V S+ + + IVD L Y A K+ +I L CT+ ++
Sbjct: 772 PAIASSKTEKVHISDHVRSILANGDIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQ 831
Query: 590 RPPMSVVVKMINNI 603
RP MS VV + I
Sbjct: 832 RPTMSQVVMELKQI 845
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 181 bits (460), Expect = 4e-45
Identities = 111/283 (39%), Positives = 159/283 (55%), Gaps = 4/283 (1%)
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
L +ATN FH+ + +G GG G VYK L DG+ VAIK+L + Q F E+ I ++
Sbjct: 881 LLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETIGKIK 940
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQ-QLTWEVRHKIILGTAEGLA 443
H+NLV LLG G E LLVYEY+ SL D L R+ + +L W R KI +G A GLA
Sbjct: 941 HRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIGAARGLA 1000
Query: 444 YLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIS-TAICGTLGYMAP 502
+LH IIHRD+K SN+LLD+N +++DFG+ARL + +S + + GT GY+ P
Sbjct: 1001 FLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPP 1060
Query: 503 EYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVD-P 561
EY + + K D+YS+GV+++ELL+GK T + L L+ ++ D+ D
Sbjct: 1061 EYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLVGWVKLHAKGKITDVFDRE 1120
Query: 562 ILEGNYPAE-EACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
+L+ + E E + LK+ C RP M V+ M I
Sbjct: 1121 LLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFKEI 1163
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 180 bits (457), Expect = 9e-45
Identities = 111/283 (39%), Positives = 158/283 (55%), Gaps = 4/283 (1%)
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
L +ATN FH+ + +G GG G VYK L DG+ VAIK+L + Q F E+ I ++
Sbjct: 881 LLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETIGKIK 940
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQ-QLTWEVRHKIILGTAEGLA 443
H+NLV LLG G E LLVYEY+ SL D L R+ +L W R KI +G A GLA
Sbjct: 941 HRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIGAARGLA 1000
Query: 444 YLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIS-TAICGTLGYMAP 502
+LH IIHRD+K SN+LLD+N +++DFG+ARL + +S + + GT GY+ P
Sbjct: 1001 FLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPP 1060
Query: 503 EYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVD-P 561
EY + + K D+YS+GV+++ELL+GK T + L L+ ++ D+ D
Sbjct: 1061 EYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLVGWVKLHAKGKITDVFDRE 1120
Query: 562 ILEGNYPAE-EACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
+L+ + E E + LK+ C RP M V+ M I
Sbjct: 1121 LLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFKEI 1163
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 175 bits (444), Expect = 3e-43
Identities = 120/354 (33%), Positives = 180/354 (49%), Gaps = 34/354 (9%)
Query: 271 IVAESFAAVATLLIVATIIFFVRKNVLRQRRER-------RQFGAFLFSENNSKLNMPYE 323
I+ F + +V ++F + LR + R F FSE+
Sbjct: 627 ILLTIFLLAGLVFVVGIVMFIAKCRKLRALKSSTLAASKWRSFHKLHFSEH--------- 677
Query: 324 VLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFN-------- 375
+ + + N +G G SG VYK L G VA+K+L+ + D + +
Sbjct: 678 ---EIADCLDEKNVIGFGSSGKVYKVELRGGEVVAVKKLNKSVKGGDDEYSSDSLNRDVF 734
Query: 376 --EVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHL-SVRRNLQQLTWEVRH 432
EV + ++HK++V+L C +G LLVYEY+PN SL D L R+ L W R
Sbjct: 735 AAEVETLGTIRHKSIVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLGWPERL 794
Query: 433 KIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTA 492
+I L AEGL+YLH + I+HRD+K SNILLD ++ K+ADFG+A++ S+ A
Sbjct: 795 RIALDAAEGLSYLHHDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAKVGQMSGSKTPEA 854
Query: 493 ---ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSL 549
I G+ GY+APEY+ ++ EK+DIYSFGV+++EL++GK T + V +
Sbjct: 855 MSGIAGSCGYIAPEYVYTLRVNEKSDIYSFGVVLLELVTGKQPTDSELGDKDMAKWVCTA 914
Query: 550 YGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI 603
L ++DP L+ + EE K++ IGLLCT RP M VV M+ +
Sbjct: 915 LDKCGLEPVIDPKLDLKF-KEEISKVIHIGLLCTSPLPLNRPSMRKVVIMLQEV 967
>IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4) (NY-REN-64 antigen)
Length = 460
Score = 164 bits (414), Expect = 9e-40
Identities = 98/216 (45%), Positives = 136/216 (62%), Gaps = 16/216 (7%)
Query: 325 LEKATNYFHD------SNKLGEGGSGSVYKGALPDGTTVAIKRLS----FNTTQWADHFF 374
L+ TN F + NK+GEGG G VYKG + + TTVA+K+L+ T + F
Sbjct: 173 LKNVTNNFDERPISVGGNKMGEGGFGVVYKGYV-NNTTVAVKKLAAMVDITTEELKQQFD 231
Query: 375 NEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKI 434
E+ ++ QH+NLV+LLG S G + LVY Y+PN SL D LS L+W +R KI
Sbjct: 232 QEIKVMAKCQHENLVELLGFSSDGDDLCLVYVYMPNGSLLDRLSCLDGTPPLSWHMRCKI 291
Query: 435 ILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPE-DQSQISTAI 493
G A G+ +LHE IHRDIK +NILLD+ FT KI+DFGLAR + Q+ +++ I
Sbjct: 292 AQGAANGINFLHENHH---IHRDIKSANILLDEAFTAKISDFGLARASEKFAQTVMTSRI 348
Query: 494 CGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSG 529
GT YMAPE + G++T K+DIYSFGV+++E+++G
Sbjct: 349 VGTTAYMAPE-ALRGEITPKSDIYSFGVVLLEIITG 383
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 75,992,890
Number of Sequences: 164201
Number of extensions: 3178476
Number of successful extensions: 11056
Number of sequences better than 10.0: 1617
Number of HSP's better than 10.0 without gapping: 1174
Number of HSP's successfully gapped in prelim test: 443
Number of HSP's that attempted gapping in prelim test: 7435
Number of HSP's gapped (non-prelim): 1856
length of query: 656
length of database: 59,974,054
effective HSP length: 117
effective length of query: 539
effective length of database: 40,762,537
effective search space: 21971007443
effective search space used: 21971007443
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0076c.7


