

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.9
(351 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
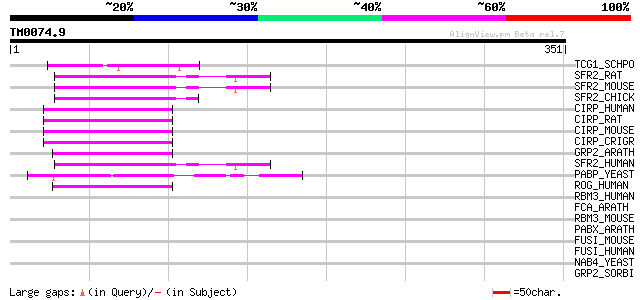
Score E
Sequences producing significant alignments: (bits) Value
TCG1_SCHPO (Q9HEQ9) Single-stranded TG1-3 binding protein 48 4e-05
SFR2_RAT (Q6PDU1) Splicing factor, arginine/serine-rich 2 (Splic... 48 4e-05
SFR2_MOUSE (Q62093) Splicing factor, arginine/serine-rich 2 (Spl... 48 4e-05
SFR2_CHICK (P30352) Splicing factor, arginine/serine-rich 2 (Spl... 47 9e-05
CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein (Glycine-... 46 1e-04
CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-ri... 46 2e-04
CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein (Glycine-... 46 2e-04
CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein (Glycine-... 46 2e-04
GRP2_ARATH (Q9SVM8) Glycine-rich RNA-binding protein 2, mitochon... 45 3e-04
SFR2_HUMAN (Q01130) Splicing factor, arginine/serine-rich 2 (Spl... 45 4e-04
PABP_YEAST (P04147) Polyadenylate-binding protein, cytoplasmic a... 44 5e-04
ROG_HUMAN (P38159) Heterogeneous nuclear ribonucleoprotein G (hn... 44 8e-04
RBM3_HUMAN (P98179) Putative RNA-binding protein 3 (RNA binding ... 43 0.001
FCA_ARATH (O04425) Flowering time control protein FCA 43 0.001
RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding ... 42 0.002
PABX_ARATH (Q9ZQA8) Probable polyadenylate-binding protein At2g3... 42 0.002
FUSI_MOUSE (Q9R0U0) FUS interacting serine-arginine rich protein... 42 0.002
FUSI_HUMAN (O75494) FUS interacting serine-arginine rich protein... 42 0.002
NAB4_YEAST (Q99383) Nuclear polyadenylated RNA-binding protein 4 42 0.002
GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2 42 0.002
>TCG1_SCHPO (Q9HEQ9) Single-stranded TG1-3 binding protein
Length = 348
Score = 47.8 bits (112), Expect = 4e-05
Identities = 36/108 (33%), Positives = 53/108 (48%), Gaps = 14/108 (12%)
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRR------EKFGFVLFQSK 78
++ R+FV L TK S++R LFE GTV V +P +RVRR FV F ++
Sbjct: 42 DDFRVFVGRLSTSTKKSEIRSLFETVGTVRKVTIP--FRRVRRGTRLVPSGIAFVTFNNQ 99
Query: 79 RDGIAAMRSLHGTRLNSFFLSINPARFV------DRKYKWNPPAKQHE 120
D A+ +L+G L+ + + AR V DRK N ++ E
Sbjct: 100 EDVDKAIETLNGKTLDDREIVVQKARPVQEQPIKDRKKSKNKNGEEPE 147
>SFR2_RAT (Q6PDU1) Splicing factor, arginine/serine-rich 2 (Splicing
factor SC35) (SC-35) (Splicing component, 35 kDa)
Length = 220
Score = 47.8 bits (112), Expect = 4e-05
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 15 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 74
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 75 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 112
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 113 GYGRRSRSPRRRRRSRSRS 131
>SFR2_MOUSE (Q62093) Splicing factor, arginine/serine-rich 2
(Splicing factor SC35) (SC-35) (Splicing component, 35
kDa) (PR264 protein)
Length = 220
Score = 47.8 bits (112), Expect = 4e-05
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 15 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 74
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 75 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 112
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 113 GYGRRSRSPRRRRRSRSRS 131
>SFR2_CHICK (P30352) Splicing factor, arginine/serine-rich 2
(Splicing factor SC35) (SC-35) (Splicing component, 35
kDa) (PR264 protein)
Length = 221
Score = 46.6 bits (109), Expect = 9e-05
Identities = 29/91 (31%), Positives = 42/91 (45%), Gaps = 6/91 (6%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 16 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 75
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQH 119
G L+ L + AR+ PP H
Sbjct: 76 DGAVLDGRELRVQMARY------GRPPDSHH 100
>CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 46.2 bits (108), Expect = 1e-04
Identities = 24/82 (29%), Positives = 44/82 (53%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
+A + +LFV GL DT + ++F ++G + V V K + R FGFV F++ D
Sbjct: 1 MASDEGKLFVGGLSFDTNEQSLEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDA 60
Query: 82 IAAMRSLHGTRLNSFFLSINPA 103
AM +++G ++ + ++ A
Sbjct: 61 KDAMMAMNGKSVDGRQIRVDQA 82
>CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-rich
RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 45.8 bits (107), Expect = 2e-04
Identities = 24/82 (29%), Positives = 44/82 (53%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
+A + +LFV GL DT + ++F ++G + V V K + R FGFV F++ D
Sbjct: 1 MASDEGKLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDA 60
Query: 82 IAAMRSLHGTRLNSFFLSINPA 103
AM +++G ++ + ++ A
Sbjct: 61 KDAMMAMNGKSVDGRQIRVDQA 82
>CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 45.8 bits (107), Expect = 2e-04
Identities = 24/82 (29%), Positives = 44/82 (53%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
+A + +LFV GL DT + ++F ++G + V V K + R FGFV F++ D
Sbjct: 1 MASDEGKLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDA 60
Query: 82 IAAMRSLHGTRLNSFFLSINPA 103
AM +++G ++ + ++ A
Sbjct: 61 KDAMMAMNGKSVDGRQIRVDQA 82
>CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 45.8 bits (107), Expect = 2e-04
Identities = 24/82 (29%), Positives = 44/82 (53%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
+A + +LFV GL DT + ++F ++G + V V K + R FGFV F++ D
Sbjct: 1 MASDEGKLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDA 60
Query: 82 IAAMRSLHGTRLNSFFLSINPA 103
AM +++G ++ + ++ A
Sbjct: 61 KDAMMAMNGKSVDGRQIRVDQA 82
>GRP2_ARATH (Q9SVM8) Glycine-rich RNA-binding protein 2,
mitochondrial precursor (AtGRP2)
Length = 158
Score = 45.1 bits (105), Expect = 3e-04
Identities = 26/76 (34%), Positives = 36/76 (47%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LF+ GL T + +R F FG VV V + R FGFV F + AA+
Sbjct: 36 KLFIGGLSWGTDDASLRDAFAHFGDVVDAKVIVDRETGRSRGFGFVNFNDEGAATAAISE 95
Query: 88 LHGTRLNSFFLSINPA 103
+ G LN + +NPA
Sbjct: 96 MDGKELNGRHIRVNPA 111
>SFR2_HUMAN (Q01130) Splicing factor, arginine/serine-rich 2
(Splicing factor SC35) (SC-35) (Splicing component, 35
kDa) (PR264 protein)
Length = 220
Score = 44.7 bits (104), Expect = 4e-04
Identities = 38/139 (27%), Positives = 57/139 (40%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++ V V++P+ F FV F KRD AM ++
Sbjct: 15 LKVDNLTYRTSPDTLRRVFEKYRRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 74
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 75 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 112
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 113 GYGRRSRSPRRRRRSRSRS 131
>PABP_YEAST (P04147) Polyadenylate-binding protein, cytoplasmic and
nuclear (Poly(A)-binding protein) (PABP) (ARS consensus
binding protein ACBP-67) (Polyadenylate tail-binding
protein)
Length = 576
Score = 44.3 bits (103), Expect = 5e-04
Identities = 43/178 (24%), Positives = 77/178 (43%), Gaps = 28/178 (15%)
Query: 12 PRWSCHERSEVAEEN----IRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRR 67
P S ER EE L+V ++ +T Q ++LF +FG +VS + K +
Sbjct: 199 PHLSRKERDSQLEETKAHYTNLYVKNINSETTDEQFQELFAKFGPIVSASLEKDADG-KL 257
Query: 68 EKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKK 127
+ FGFV ++ D + A+ +L+ + LN L + A+ K++E + KK
Sbjct: 258 KGFGFVNYEKHEDAVKAVEALNDSELNGEKLYVGRAQ------------KKNERMHVLKK 305
Query: 128 IRQEWRIKHRVGAPFSGTGGFSQESQKVWRVKKSTRKSPEKKVEEQERSGMTMNQVRV 185
+ +R++ A + G F VK ++K+EE+ T+ +V
Sbjct: 306 QYEAYRLEKM--AKYQGVNLF---------VKNLDDSVDDEKLEEEFAPYGTITSAKV 352
Score = 32.3 bits (72), Expect = 1.8
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 1/61 (1%)
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAA 84
+ + LFV LD ++ + F +GT+ S V +T + + + FGFV F + + A
Sbjct: 319 QGVNLFVKNLDDSVDDEKLEEEFAPYGTITSAKVMRT-ENGKSKGFGFVCFSTPEEATKA 377
Query: 85 M 85
+
Sbjct: 378 I 378
Score = 30.8 bits (68), Expect = 5.4
Identities = 22/89 (24%), Positives = 42/89 (46%), Gaps = 1/89 (1%)
Query: 14 WSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFV 73
WS + S + + +F+ L D + + F FG ++S + T + + + FGFV
Sbjct: 112 WSQRDPSLRKKGSGNIFIKNLHPDIDNKALYDTFSVFGDILSSKIA-TDENGKSKGFGFV 170
Query: 74 LFQSKRDGIAAMRSLHGTRLNSFFLSINP 102
F+ + A+ +L+G LN + + P
Sbjct: 171 HFEEEGAAKEAIDALNGMLLNGQEIYVAP 199
>ROG_HUMAN (P38159) Heterogeneous nuclear ribonucleoprotein G (hnRNP
G) (RNA binding motif protein, X chromosome)
(Glycoprotein p43)
Length = 391
Score = 43.5 bits (101), Expect = 8e-04
Identities = 21/76 (27%), Positives = 39/76 (50%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LF+ GL+ +T + +F ++G +V V + K + + F FV F+S D A R
Sbjct: 9 KLFIGGLNTETNEKALEAVFGKYGRIVEVLLMKDRETNKSRGFAFVTFESPADAKDAARD 68
Query: 88 LHGTRLNSFFLSINPA 103
++G L+ + + A
Sbjct: 69 MNGKSLDGKAIKVEQA 84
>RBM3_HUMAN (P98179) Putative RNA-binding protein 3 (RNA binding
motif protein 3) (RNPL)
Length = 157
Score = 43.1 bits (100), Expect = 0.001
Identities = 24/82 (29%), Positives = 41/82 (49%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
++ E +LFV GL+ +T + F FG + V V K + R FGF+ F +
Sbjct: 1 MSSEEGKLFVGGLNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHA 60
Query: 82 IAAMRSLHGTRLNSFFLSINPA 103
AMR+++G L+ + ++ A
Sbjct: 61 SVAMRAMNGESLDGRQIRVDHA 82
>FCA_ARATH (O04425) Flowering time control protein FCA
Length = 747
Score = 43.1 bits (100), Expect = 0.001
Identities = 25/79 (31%), Positives = 42/79 (52%), Gaps = 1/79 (1%)
Query: 13 RWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGF 72
R++ ER + +LFV L++ +V ++F +FG V V++ + R R GF
Sbjct: 197 RYADGERERIGTLEFKLFVGSLNKQATEKEVEEIFLQFGHVEDVYLMRDEYRQSR-GCGF 255
Query: 73 VLFQSKRDGIAAMRSLHGT 91
V + SK +AA+ L+GT
Sbjct: 256 VKYSSKETAMAAIDGLNGT 274
Score = 37.4 bits (85), Expect = 0.057
Identities = 18/63 (28%), Positives = 35/63 (54%)
Query: 27 IRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMR 86
++LFV + R ++R FE+ G V+ V + K + +++ FV + + +D A+R
Sbjct: 120 VKLFVGSVPRTATEEEIRPYFEQHGNVLEVALIKDKRTGQQQGCCFVKYATSKDADRAIR 179
Query: 87 SLH 89
+LH
Sbjct: 180 ALH 182
>RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding
motif protein 3)
Length = 153
Score = 42.4 bits (98), Expect = 0.002
Identities = 24/82 (29%), Positives = 41/82 (49%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
++ E +LFV GL+ +T + F FG + V V K + R FGF+ F +
Sbjct: 1 MSSEEGKLFVGGLNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHA 60
Query: 82 IAAMRSLHGTRLNSFFLSINPA 103
AMR+++G L+ + ++ A
Sbjct: 61 SDAMRAMNGESLDGRQIRVDHA 82
>PABX_ARATH (Q9ZQA8) Probable polyadenylate-binding protein
At2g36660 (Poly(A)-binding protein At2g36660) (PABP)
Length = 609
Score = 42.4 bits (98), Expect = 0.002
Identities = 25/87 (28%), Positives = 44/87 (49%), Gaps = 1/87 (1%)
Query: 18 ERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
+R + E+ L++ LD D +R+ F FG +VS+ + K R+ R + FV F +
Sbjct: 192 DRVKPEEKYTNLYMKNLDADVSEDLLREKFAEFGKIVSLAIAKDENRLCR-GYAFVNFDN 250
Query: 78 KRDGIAAMRSLHGTRLNSFFLSINPAR 104
D A +++GT+ S L + A+
Sbjct: 251 PEDARRAAETVNGTKFGSKCLYVGRAQ 277
Score = 35.4 bits (80), Expect = 0.22
Identities = 18/74 (24%), Positives = 39/74 (52%), Gaps = 1/74 (1%)
Query: 17 HERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQ 76
HE ++ + ++V ++ ++RK F + GT+ S + ++ + + FGFV F
Sbjct: 294 HEEQKMIAKVSNIYVKNVNVAVTEEELRKHFSQCGTITSTKL-MCDEKGKSKGFGFVCFS 352
Query: 77 SKRDGIAAMRSLHG 90
+ + I A+++ HG
Sbjct: 353 TPEEAIDAVKTFHG 366
Score = 34.3 bits (77), Expect = 0.48
Identities = 26/91 (28%), Positives = 47/91 (51%), Gaps = 10/91 (10%)
Query: 1 MRPTGSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPK 60
+R S RAP R R+ V +FV L ++ ++ +F++FG +VS V
Sbjct: 95 IRVMWSVRAPDAR-----RNGVGN----VFVKNLPESVTNAVLQDMFKKFGNIVSCKV-A 144
Query: 61 TIKRVRREKFGFVLFQSKRDGIAAMRSLHGT 91
T++ + +GFV F+ + AA+++L+ T
Sbjct: 145 TLEDGKSRGYGFVQFEQEDAAHAAIQTLNST 175
>FUSI_MOUSE (Q9R0U0) FUS interacting serine-arginine rich protein
1 (TLS-associated protein with Ser-Arg repeats)
(TLS-associated protein with SR repeats) (TASR)
(TLS-associated serine-arginine protein)
(TLS-associated SR protein) (Neural specific SR prot
Length = 262
Score = 42.4 bits (98), Expect = 0.002
Identities = 22/63 (34%), Positives = 33/63 (51%)
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAM 85
N LFV + DT+ +R+ F R+G +V V+VP R F +V F+ RD A+
Sbjct: 9 NTSLFVRNVADDTRSEDLRREFGRYGPIVDVYVPLDFYTRRPRGFAYVQFEDVRDAEDAL 68
Query: 86 RSL 88
+L
Sbjct: 69 HNL 71
>FUSI_HUMAN (O75494) FUS interacting serine-arginine rich protein
1 (TLS-associated protein with Ser-Arg repeats)
(TLS-associated protein with SR repeats) (TASR)
(TLS-associated serine-arginine protein)
(TLS-associated SR protein) (40 kDa SR-repressor pro
Length = 262
Score = 42.4 bits (98), Expect = 0.002
Identities = 22/63 (34%), Positives = 33/63 (51%)
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAM 85
N LFV + DT+ +R+ F R+G +V V+VP R F +V F+ RD A+
Sbjct: 9 NTSLFVRNVADDTRSEDLRREFGRYGPIVDVYVPLDFYTRRPRGFAYVQFEDVRDAEDAL 68
Query: 86 RSL 88
+L
Sbjct: 69 HNL 71
>NAB4_YEAST (Q99383) Nuclear polyadenylated RNA-binding protein 4
Length = 534
Score = 42.0 bits (97), Expect = 0.002
Identities = 22/89 (24%), Positives = 46/89 (50%), Gaps = 5/89 (5%)
Query: 6 SARAPPPRWSCHERSE-----VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPK 60
S+++PP + + E +++E+ ++F+ GL+ DT +R+ F ++GTV + + K
Sbjct: 133 SSQSPPQQQVTQTKEERSKADLSKESCKMFIGGLNWDTTEDNLREYFGKYGTVTDLKIMK 192
Query: 61 TIKRVRREKFGFVLFQSKRDGIAAMRSLH 89
R FGF+ F+ +++ H
Sbjct: 193 DPATGRSRGFGFLSFEKPSSVDEVVKTQH 221
>GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2
Length = 168
Score = 42.0 bits (97), Expect = 0.002
Identities = 24/82 (29%), Positives = 42/82 (50%)
Query: 23 AEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGI 82
A+ R FV GL T + + + F FG V+ V + R FGFV F S++ +
Sbjct: 4 ADVEYRCFVGGLAWATNNETLEQAFANFGQVIDSKVITDRETGRSRGFGFVTFSSEQSML 63
Query: 83 AAMRSLHGTRLNSFFLSINPAR 104
A+ +++G L+ +++N A+
Sbjct: 64 DAIENMNGKELDGRNITVNQAQ 85
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,106,685
Number of Sequences: 164201
Number of extensions: 1660183
Number of successful extensions: 4519
Number of sequences better than 10.0: 209
Number of HSP's better than 10.0 without gapping: 131
Number of HSP's successfully gapped in prelim test: 78
Number of HSP's that attempted gapping in prelim test: 4253
Number of HSP's gapped (non-prelim): 328
length of query: 351
length of database: 59,974,054
effective HSP length: 111
effective length of query: 240
effective length of database: 41,747,743
effective search space: 10019458320
effective search space used: 10019458320
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0074.9


