

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.3
(267 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
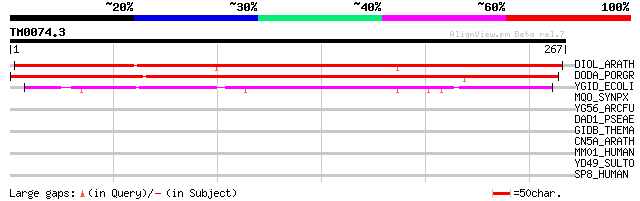
DIOL_ARATH (Q949R4) 4,5-DOPA dioxygenase extradiol-like protein 376 e-104
DODA_PORGR (Q7XA48) 4,5-DOPA dioxygenase extradiol (EC 1.13.-.-) 327 2e-89
YGID_ECOLI (P24197) Hypothetical protein ygiD 169 5e-42
MQO_SYNPX (Q7U5L7) Probable malate:quinone oxidoreductase (EC 1.... 33 0.94
YG56_ARCFU (O28617) Hypothetical protein AF1656 precursor 32 1.2
DAD1_PSEAE (Q9HTQ0) D-amino acid dehydrogenase 1 small subunit (... 31 3.6
GIDB_THEMA (Q9WZG6) Methyltransferase gidB (EC 2.1.-.-) (Glucose... 30 4.7
CN5A_ARATH (Q9FVU9) COP9 signalosome complex subunit 5a (EC 3.4.... 30 4.7
MM01_HUMAN (P03956) Interstitial collagenase precursor (EC 3.4.2... 30 6.1
YD49_SULTO (Q971K8) Hypothetical UPF0219 protein ST1349 30 8.0
SP8_HUMAN (Q8IXZ3) Transcription factor Sp8 (Specificity protein 8) 30 8.0
>DIOL_ARATH (Q949R4) 4,5-DOPA dioxygenase extradiol-like protein
Length = 269
Score = 376 bits (965), Expect = e-104
Identities = 180/267 (67%), Positives = 221/267 (82%), Gaps = 4/267 (1%)
Query: 3 LKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDS 62
+ TF++SHGSPTLSID+SL AR+F +SW ++V P +P SILVIS HWDT P+VN V
Sbjct: 4 VNQTFFLSHGSPTLSIDDSLEARQFFKSWTQKVLPQKPKSILVISAHWDTKFPSVNTV-L 62
Query: 63 TNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELL-KEGGFSRVDEDKKRGLDHGAWVP 121
N+TI+DF GFP PMY+LKY APGA L KRVKELL KEGG RVDED KRGLDHGAWVP
Sbjct: 63 RNNTIHDFSGFPDPMYKLKYEAPGAIELGKRVKELLMKEGGMKRVDEDTKRGLDHGAWVP 122
Query: 122 LLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHA 181
L+LMYPEADIP+CQLSVQSN +G++HYN+GKALA LKDEGVLI+GSGSA HNLR L+ +
Sbjct: 123 LMLMYPEADIPICQLSVQSNQNGSYHYNMGKALASLKDEGVLIIGSGSATHNLRKLDFNI 182
Query: 182 TVAA--PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGE 239
T + PWA+EFD+WL+++LL+GRY DVN +E+KAP+AK AHPWP+H YPLHV +GAAG
Sbjct: 183 TDGSPVPWALEFDHWLRDSLLQGRYGDVNEWEEKAPNAKMAHPWPEHLYPLHVVMGAAGG 242
Query: 240 NSKAKLIHSSIDLGSLSYASYQFTSDV 266
++KA+ IH+S LG+LSY+SY FTS +
Sbjct: 243 DAKAEQIHTSWQLGTLSYSSYSFTSSL 269
>DODA_PORGR (Q7XA48) 4,5-DOPA dioxygenase extradiol (EC 1.13.-.-)
Length = 271
Score = 327 bits (837), Expect = 2e-89
Identities = 158/265 (59%), Positives = 193/265 (72%), Gaps = 2/265 (0%)
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
++ K++F++SHG+P + DES +AR FL WKK VFP +P SILV+S HW+T VP V+
Sbjct: 7 VSFKESFFLSHGNPAMLADESFIARNFLLGWKKNVFPVKPKSILVVSAHWETDVPCVSAG 66
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
N IYDF P M+Q+KYPAPG P LAKRV+ELL GGF D++RG DH +WV
Sbjct: 67 QYPN-VIYDFTEVPASMFQMKYPAPGCPKLAKRVQELLIAGGFKSAKLDEERGFDHSSWV 125
Query: 121 PLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERH 180
PL +M PEADIPVCQLSVQ LD THH+N+G+ALAPLK EGVL +GSG AVH
Sbjct: 126 PLSMMCPEADIPVCQLSVQPGLDATHHFNVGRALAPLKGEGVLFIGSGGAVHPSDDTPHW 185
Query: 181 ATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHA-KKAHPWPDHFYPLHVAIGAAGE 239
APWA EFD WL++ALLEGRYEDVN+Y+ KAP K AHP P+HF PLHVA+GA GE
Sbjct: 186 FDGVAPWAAEFDQWLEDALLEGRYEDVNNYQTKAPEGWKLAHPIPEHFLPLHVAMGAGGE 245
Query: 240 NSKAKLIHSSIDLGSLSYASYQFTS 264
SKA+LI+ + D G+L YASY+FTS
Sbjct: 246 KSKAELIYRTWDHGTLGYASYKFTS 270
>YGID_ECOLI (P24197) Hypothetical protein ygiD
Length = 271
Score = 169 bits (429), Expect = 5e-42
Identities = 101/261 (38%), Positives = 153/261 (57%), Gaps = 17/261 (6%)
Query: 8 YISHGSPTLSIDESLVARKFLQSWKK-EVFPPRPTSILVISGHWDTAVPTVNVVDSTNDT 66
++ HGSP ++++L R SW+K + PRP +I+V+S HW T V ++ T T
Sbjct: 19 FLGHGSPMNVLEDNLYTR----SWQKLGMTLPRPQAIVVVSAHWFTRGTGVTAME-TPPT 73
Query: 67 IYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKR-GLDHGAWVPLLLM 125
I+DF GFP+ +Y YPAPG+P LA+R+ ELL V DK+ G DHG+W L+ M
Sbjct: 74 IHDFGGFPQALYDTHYPAPGSPALAQRLVELLAP---IPVTLDKEAWGFDHGSWGVLIKM 130
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVAA 185
YP+ADIP+ QLS+ S+ H+ +G+ LA L+DEG+++V SG+ VHNLR ++ H +
Sbjct: 131 YPDADIPMVQLSIDSSKPAAWHFEMGRKLAALRDEGIMLVASGNVVHNLRTVKWHGDSSP 190
Query: 186 -PWAVEFDNWLKEALL-EGRYED---VNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGEN 240
PWA F+ ++K L +G E VN+ + + ++P P+H+ PL +GA
Sbjct: 191 YPWATSFNEYVKANLTWQGPVEQHPLVNYLDHEG--GTLSNPTPEHYLPLLYVLGAWDGQ 248
Query: 241 SKAKLIHSSIDLGSLSYASYQ 261
+ I++GSLS S Q
Sbjct: 249 EPITIPVEGIEMGSLSMLSVQ 269
>MQO_SYNPX (Q7U5L7) Probable malate:quinone oxidoreductase (EC
1.1.99.16) (Malate dehydrogenase [acceptor]) (MQO)
Length = 502
Score = 32.7 bits (73), Expect = 0.94
Identities = 25/108 (23%), Positives = 46/108 (42%), Gaps = 1/108 (0%)
Query: 82 YPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLMYPEADIPVCQLSVQSN 141
+ A L +R ++L + F+ + ++R + W+PL++ + PV ++
Sbjct: 125 WTAENIAFLRQRFEQLSEIPAFAAMRWSEER-TELTDWMPLVMAGRDLKQPVAATRIERG 183
Query: 142 LDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVAAPWAV 189
D L PL+ G L V G+ V +++ L R A W V
Sbjct: 184 TDVDFGALTRAYLEPLQRSGALCVEYGTQVRDIKRLRRGDMTEADWRV 231
>YG56_ARCFU (O28617) Hypothetical protein AF1656 precursor
Length = 144
Score = 32.3 bits (72), Expect = 1.2
Identities = 19/51 (37%), Positives = 24/51 (46%), Gaps = 6/51 (11%)
Query: 47 SGHWDTAV-PTVNVVDSTNDTIYDFYGFPKPMYQLK-----YPAPGAPHLA 91
SGHW + + PTV+ V + DFYG P Y APG P L+
Sbjct: 36 SGHWASEIYPTVSFVKKESPVKIDFYGKPNEEYSYDVIFRIVDAPGIPKLS 86
>DAD1_PSEAE (Q9HTQ0) D-amino acid dehydrogenase 1 small subunit (EC
1.4.99.1)
Length = 432
Score = 30.8 bits (68), Expect = 3.6
Identities = 22/79 (27%), Positives = 34/79 (42%), Gaps = 7/79 (8%)
Query: 154 LAPLKDEGVLIVGSGSAVHNLRALERHATVAAPWAVEFDNWLKEALLEGRYE------DV 207
L P +G IVG+ + NL H T+ A +L + + + R + D+
Sbjct: 355 LRPATPDGTPIVGA-TRYRNLFLNTGHGTLGWTMACGSGRYLADLMAKKRPQISTEGLDI 413
Query: 208 NHYEQKAPHAKKAHPWPDH 226
+ Y +AK AHP P H
Sbjct: 414 SRYSNSPENAKNAHPAPAH 432
>GIDB_THEMA (Q9WZG6) Methyltransferase gidB (EC 2.1.-.-) (Glucose
inhibited division protein B)
Length = 238
Score = 30.4 bits (67), Expect = 4.7
Identities = 27/76 (35%), Positives = 37/76 (48%), Gaps = 15/76 (19%)
Query: 136 LSVQSNLDGTHHYNIGKALAPLKDE--GVLI-VGSGSAVHNLRALERHATVAAPWAVEFD 192
L+ +LD H N+ + L PLK+E G L+ VGSG+ V L A+ F
Sbjct: 47 LTAHRDLDSAVHKNVVEILLPLKEELKGTLLDVGSGNGVPGLIL-----------AIFFS 95
Query: 193 NWLKEALLEGRYEDVN 208
LK LL+ R + VN
Sbjct: 96 K-LKVVLLDSREKSVN 110
>CN5A_ARATH (Q9FVU9) COP9 signalosome complex subunit 5a (EC
3.4.-.-) (Signalosome subunit 5a) (Jun activation
domain-binding homolog 2)
Length = 358
Score = 30.4 bits (67), Expect = 4.7
Identities = 15/45 (33%), Positives = 25/45 (55%)
Query: 51 DTAVPTVNVVDSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVK 95
+ ++ TV+ DST+D I+ + + +Q + P PH KRVK
Sbjct: 16 ENSILTVDSPDSTSDNIFYYDDTSQTRFQQEKPWENDPHYFKRVK 60
>MM01_HUMAN (P03956) Interstitial collagenase precursor (EC
3.4.24.7) (Matrix metalloproteinase-1) (MMP-1)
(Fibroblast collagenase)
Length = 469
Score = 30.0 bits (66), Expect = 6.1
Identities = 16/51 (31%), Positives = 26/51 (50%)
Query: 216 HAKKAHPWPDHFYPLHVAIGAAGENSKAKLIHSSIDLGSLSYASYQFTSDV 266
H + W ++F ++ AA E + + S D+G+L Y SY F+ DV
Sbjct: 196 HFDEDERWTNNFREYNLHRVAAHELGHSLGLSHSTDIGALMYPSYTFSGDV 246
>YD49_SULTO (Q971K8) Hypothetical UPF0219 protein ST1349
Length = 348
Score = 29.6 bits (65), Expect = 8.0
Identities = 22/73 (30%), Positives = 33/73 (45%), Gaps = 5/73 (6%)
Query: 42 SILVISGHWDTAVPTVNVVDSTNDTIYDFY---GFPKPMYQLKYPAPGA--PHLAKRVKE 96
S+ I G D + + S + DF+ G P P++ + A H+ V+
Sbjct: 158 SVAYIVGPADESSAIIEYATSYTTDMPDFWRRDGMPYPLHGEAFTGEPAYFAHIISAVQL 217
Query: 97 LLKEGGFSRVDED 109
LLKEGG+S D D
Sbjct: 218 LLKEGGYSISDFD 230
>SP8_HUMAN (Q8IXZ3) Transcription factor Sp8 (Specificity protein 8)
Length = 508
Score = 29.6 bits (65), Expect = 8.0
Identities = 13/29 (44%), Positives = 15/29 (50%)
Query: 215 PHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
P AHP+ F P H +GAAGE A
Sbjct: 195 PRVGMAHPYESWFKPSHPGLGAAGEVGSA 223
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,889,145
Number of Sequences: 164201
Number of extensions: 1572339
Number of successful extensions: 3441
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 3425
Number of HSP's gapped (non-prelim): 12
length of query: 267
length of database: 59,974,054
effective HSP length: 108
effective length of query: 159
effective length of database: 42,240,346
effective search space: 6716215014
effective search space used: 6716215014
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0074.3


