

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0062.9
(271 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
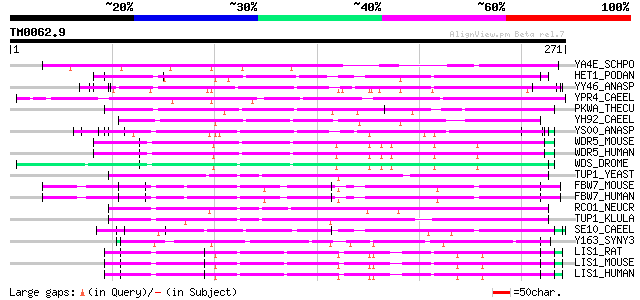
Score E
Sequences producing significant alignments: (bits) Value
YA4E_SCHPO (Q09731) Hypothetical WD-repeat protein C31A2.14 in c... 105 2e-22
HET1_PODAN (Q00808) Vegetatible incompatibility protein HET-E-1 93 6e-19
YY46_ANASP (Q8YRI1) Hypothetical WD-repeat protein alr3466 85 2e-16
YPR4_CAEEL (Q20059) Hypothetical WD-repeat protein F35G12.4 in c... 85 2e-16
PKWA_THECU (P49695) Probable serine/threonine-protein kinase pkw... 85 2e-16
YH92_CAEEL (Q23256) Hypothetical WD-repeat protein ZC302.2 in ch... 81 2e-15
YS00_ANASP (Q8YTC2) Hypothetical WD-repeat protein alr2800 80 4e-15
WDR5_MOUSE (P61965) WD-repeat protein 5 (WD-repeat protein BIG-3) 80 4e-15
WDR5_HUMAN (P61964) WD-repeat protein 5 80 4e-15
WDS_DROME (Q9V3J8) Will die slowly protein 79 9e-15
TUP1_YEAST (P16649) Glucose repression regulatory protein TUP1 (... 76 1e-13
FBW7_MOUSE (Q8VBV4) F-box/WD-repeat protein 7 (F-box and WD-40 d... 75 1e-13
FBW7_HUMAN (Q969H0) F-box/WD-repeat protein 7 (F-box and WD-40 d... 75 1e-13
RCO1_NEUCR (P78706) Transcriptional repressor rco-1 73 6e-13
TUP1_KLULA (P56094) Transcriptional repressor TUP1 72 1e-12
SE10_CAEEL (Q93794) F-box/WD-repeat protein sel-10 (Suppressor/e... 71 2e-12
Y163_SYNY3 (Q55563) Hypothetical WD-repeat protein sll0163 70 5e-12
LIS1_RAT (P63004) Platelet-activating factor acetylhydrolase IB ... 69 2e-11
LIS1_MOUSE (P63005) Platelet-activating factor acetylhydrolase I... 69 2e-11
LIS1_HUMAN (P43034) Platelet-activating factor acetylhydrolase I... 69 2e-11
>YA4E_SCHPO (Q09731) Hypothetical WD-repeat protein C31A2.14 in
chromosome I
Length = 962
Score = 105 bits (261), Expect = 2e-22
Identities = 80/263 (30%), Positives = 118/263 (44%), Gaps = 53/263 (20%)
Query: 17 RKEKRLTYVLND-SDDTKHCAGINCLAVLKSAVSDGSD----YLFTGSRDGRLKRWALGE 71
RK ++TYVL+ +D H NCL + + GS L++G RDG+L W +G
Sbjct: 4 RKRYKVTYVLDSCNDQLGHRLDANCLVLGRYFSEQGSAPVTRALYSGGRDGQLFGWDIGY 63
Query: 72 DAATCS---ATFESHVDWVNDAVLVGDST-LVSCSSDTTLKTWDA-FSTGTCTRTLRQHS 126
D S A ++H WVND L DS ++SCSSD+T+K W +C T+ +H+
Sbjct: 64 DYGKASQPLAKIQAHSAWVNDIALTHDSEGVISCSSDSTVKLWKPHVLNASCLSTIGEHT 123
Query: 127 DYVTCLAASE-KNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSL 185
DYV ++ + S +VASGGL + VWD L
Sbjct: 124 DYVKRVSIPKYSKSPLVASGGLDKRIIVWDYNVGQEVM-----------------RFEQL 166
Query: 186 PLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVW 245
P SL G + VY+LA N ++ +GG +K +++W
Sbjct: 167 PDCSL-----------------------VVGPRSGVYSLAAN--NNIIANGGLQKDIQLW 201
Query: 246 DPRSGSKTMKLKGHTDNIRALLL 268
D S + L GHTDN+R +L+
Sbjct: 202 DCVSKKRITDLVGHTDNVRDILI 224
>HET1_PODAN (Q00808) Vegetatible incompatibility protein HET-E-1
Length = 1356
Score = 93.2 bits (230), Expect = 6e-19
Identities = 70/223 (31%), Positives = 104/223 (46%), Gaps = 20/223 (8%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSC 101
+V A S + +GS D +K W + TC+ T E H WV V D V+
Sbjct: 1011 SVWSVAFSPDGQRVASGSDDKTIKIWDTA--SGTCTQTLEGHGGWVQSVVFSPDGQRVAS 1068
Query: 102 SSDT-TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAH 160
SD T+K WDA S GTCT+TL H D V +A S VASG + G + +WD
Sbjct: 1069 GSDDHTIKIWDAVS-GTCTQTLEGHGDSVWSVAFSPDGQR-VASGSIDGTIKIWD----- 1121
Query: 161 ASAAKCIDATDDDTSNGINGSGNSLPLTS----LRSISSSNSISVHTTQNQGYIPIAAKG 216
A++ C T G G +S+ + + S S +I + + G +G
Sbjct: 1122 AASGTCTQ-----TLEGHGGWVHSVAFSPDGQRVASGSIDGTIKIWDAAS-GTCTQTLEG 1175
Query: 217 HKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
H V ++A + G + SG ++K +++WD SG+ T L+GH
Sbjct: 1176 HGGWVQSVAFSPDGQRVASGSSDKTIKIWDTASGTCTQTLEGH 1218
Score = 79.7 bits (195), Expect = 7e-15
Identities = 69/229 (30%), Positives = 101/229 (43%), Gaps = 24/229 (10%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAA--TCSATFESHVDWVNDAVLVGDSTLV 99
+V A S + + +GS D +K W DAA TC+ T E H V D V
Sbjct: 885 SVWSVAFSPDRERVASGSDDKTIKIW----DAASGTCTQTLEGHGGRVQSVAFSPDGQRV 940
Query: 100 SCSSDT-TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA 158
+ SD T+K WDA S GTCT+TL H V +A S VASG + +WD +
Sbjct: 941 ASGSDDHTIKIWDAAS-GTCTQTLEGHGSSVLSVAFSPDGQR-VASGSGDKTIKIWDTAS 998
Query: 159 AHASAAKCIDATDDDTSNGINGSGNSLPLTS----LRSISSSNSISVHTTQNQGYIPIAA 214
T T G GS S+ + + S S +I + T + G
Sbjct: 999 G----------TCTQTLEGHGGSVWSVAFSPDGQRVASGSDDKTIKIWDTAS-GTCTQTL 1047
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
+GH V ++ + G + SG + +++WD SG+ T L+GH D++
Sbjct: 1048 EGHGGWVQSVVFSPDGQRVASGSDDHTIKIWDAVSGTCTQTLEGHGDSV 1096
Score = 79.3 bits (194), Expect = 9e-15
Identities = 67/219 (30%), Positives = 99/219 (44%), Gaps = 12/219 (5%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-S 100
+VL A S + +GS D +K W T T E H V D V S
Sbjct: 843 SVLSVAFSADGQRVASGSDDKTIKIWDTASGTGT--QTLEGHGGSVWSVAFSPDRERVAS 900
Query: 101 CSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAH 160
S D T+K WDA S GTCT+TL H V +A S VASG + +WD
Sbjct: 901 GSDDKTIKIWDAAS-GTCTQTLEGHGGRVQSVAFSPDGQR-VASGSDDHTIKIWD----- 953
Query: 161 ASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKES 220
A++ C + S+ ++ + S + S S +I + T + G +GH S
Sbjct: 954 AASGTCTQTLEGHGSSVLSVAF-SPDGQRVASGSGDKTIKIWDTAS-GTCTQTLEGHGGS 1011
Query: 221 VYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
V+++A + G + SG +K +++WD SG+ T L+GH
Sbjct: 1012 VWSVAFSPDGQRVASGSDDKTIKIWDTASGTCTQTLEGH 1050
Score = 77.0 bits (188), Expect = 4e-14
Identities = 67/220 (30%), Positives = 97/220 (43%), Gaps = 24/220 (10%)
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAA--TCSATFESHVDWVNDAVLVGDSTLV-SCSS 103
A S + +GS D +K W DAA TC+ T E H V D V S S
Sbjct: 932 AFSPDGQRVASGSDDHTIKIW----DAASGTCTQTLEGHGSSVLSVAFSPDGQRVASGSG 987
Query: 104 DTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASA 163
D T+K WD S GTCT+TL H V +A S VASG + +WD +
Sbjct: 988 DKTIKIWDTAS-GTCTQTLEGHGGSVWSVAFSPDGQR-VASGSDDKTIKIWDTASG---- 1041
Query: 164 AKCIDATDDDTSNGINGSGNSLPLTS----LRSISSSNSISVHTTQNQGYIPIAAKGHKE 219
T T G G S+ + + S S ++I + + G +GH +
Sbjct: 1042 ------TCTQTLEGHGGWVQSVVFSPDGQRVASGSDDHTIKIWDAVS-GTCTQTLEGHGD 1094
Query: 220 SVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
SV+++A + G + SG + +++WD SG+ T L+GH
Sbjct: 1095 SVWSVAFSPDGQRVASGSIDGTIKIWDAASGTCTQTLEGH 1134
Score = 68.6 bits (166), Expect = 2e-11
Identities = 57/193 (29%), Positives = 85/193 (43%), Gaps = 18/193 (9%)
Query: 76 CSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAA 134
C+ T E H V D V S S D T+K WD S GT T+TL H V +A
Sbjct: 833 CTQTLEGHGSSVLSVAFSADGQRVASGSDDKTIKIWDTAS-GTGTQTLEGHGGSVWSVAF 891
Query: 135 SEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTS----L 190
S VASG + +WD A++ C T G G S+ + +
Sbjct: 892 SPDRER-VASGSDDKTIKIWD-----AASGTCTQ-----TLEGHGGRVQSVAFSPDGQRV 940
Query: 191 RSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSG 250
S S ++I + + G +GH SV ++A + G + SG +K +++WD SG
Sbjct: 941 ASGSDDHTIKIWDAAS-GTCTQTLEGHGSSVLSVAFSPDGQRVASGSGDKTIKIWDTASG 999
Query: 251 SKTMKLKGHTDNI 263
+ T L+GH ++
Sbjct: 1000 TCTQTLEGHGGSV 1012
Score = 41.2 bits (95), Expect = 0.003
Identities = 16/52 (30%), Positives = 32/52 (60%)
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+GH SV+++A + + SG +K +++WD SG+ T L+GH ++++
Sbjct: 880 EGHGGSVWSVAFSPDRERVASGSDDKTIKIWDAASGTCTQTLEGHGGRVQSV 931
>YY46_ANASP (Q8YRI1) Hypothetical WD-repeat protein alr3466
Length = 1526
Score = 85.1 bits (209), Expect = 2e-16
Identities = 71/276 (25%), Positives = 117/276 (41%), Gaps = 50/276 (18%)
Query: 40 CLAVLKSAVS---------DGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDA 90
CL +L+ S DGS L +GS D ++ W + +++ C TF+ H WVN
Sbjct: 1192 CLYILQGHTSWVNSVVFNPDGST-LASGSSDQTVRLWEI--NSSKCLCTFQGHTSWVNSV 1248
Query: 91 VLVGD-STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGG 149
V D S L S SSD T++ WD S+ C T + H+++V +A + S ++ASG
Sbjct: 1249 VFNPDGSMLASGSSDKTVRLWD-ISSSKCLHTFQGHTNWVNSVAFNPDGS-MLASGSGDQ 1306
Query: 150 EVFVWDLEAA--------HASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISV 201
V +W++ ++ H S + + D T ++ L S+ S +
Sbjct: 1307 TVRLWEISSSKCLHTFQGHTSWVSSVTFSPDGTMLASGSDDQTVRLWSISSGECLYTFLG 1366
Query: 202 HTTQ---------------------------NQGYIPIAAKGHKESVYALAMNEGGTLLV 234
HT + G +GH V ++ + GTLL
Sbjct: 1367 HTNWVGSVIFSPDGAILASGSGDQTVRLWSISSGKCLYTLQGHNNWVGSIVFSPDGTLLA 1426
Query: 235 SGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
SG ++ VR+W+ SG L GH +++R++ S
Sbjct: 1427 SGSDDQTVRLWNISSGECLYTLHGHINSVRSVAFSS 1462
Score = 81.6 bits (200), Expect = 2e-15
Identities = 70/239 (29%), Positives = 106/239 (44%), Gaps = 36/239 (15%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRW--ALGEDAATCSATFESHVDWVNDAVLVGD-STL 98
+VL A S TG G ++ W A G++ TC + H WVN D L
Sbjct: 866 SVLTVAFSPDGKLFATGDSGGIVRFWEAATGKELLTC----KGHNSWVNSVGFSQDGKML 921
Query: 99 VSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA 158
S S D T++ WD S+G C +T + H+ V + S NS ++ASG V +WD+
Sbjct: 922 ASGSDDQTVRLWD-ISSGQCLKTFKGHTSRVRSVVFS-PNSLMLASGSSDQTVRLWDI-- 977
Query: 159 AHASAAKCIDATDDDT----SNGINGSGNSLPLTS------LRSISSSNSISVHTTQNQG 208
S+ +C+ T S N G+ L S L ISSS +
Sbjct: 978 ---SSGECLYIFQGHTGWVYSVAFNLDGSMLATGSGDQTVRLWDISSSQCFYIF------ 1028
Query: 209 YIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALL 267
+GH V ++ + G +L SG ++ VR+WD SG+ L+GHT +R+++
Sbjct: 1029 ------QGHTSCVRSVVFSSDGAMLASGSDDQTVRLWDISSGNCLYTLQGHTSCVRSVV 1081
Score = 80.5 bits (197), Expect = 4e-15
Identities = 66/239 (27%), Positives = 108/239 (44%), Gaps = 23/239 (9%)
Query: 35 CAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVG 94
C G N DG L +GS D ++ W + + C TF+ H V V
Sbjct: 902 CKGHNSWVNSVGFSQDGK-MLASGSDDQTVRLWDIS--SGQCLKTFKGHTSRVRSVVFSP 958
Query: 95 DST-LVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFV 153
+S L S SSD T++ WD S+G C + H+ +V +A + S ++A+G V +
Sbjct: 959 NSLMLASGSSDQTVRLWD-ISSGECLYIFQGHTGWVYSVAFNLDGS-MLATGSGDQTVRL 1016
Query: 154 WDLEAAHA-----SAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQG 208
WD+ ++ C+ + + + SG+ L ISS N + +T Q
Sbjct: 1017 WDISSSQCFYIFQGHTSCVRSVVFSSDGAMLASGSDDQTVRLWDISSGNCL--YTLQ--- 1071
Query: 209 YIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALL 267
GH V ++ + G +L SGG +++VR+WD SG+ L+G+T +R L+
Sbjct: 1072 -------GHTSCVRSVVFSPDGAMLASGGDDQIVRLWDISSGNCLYTLQGYTSWVRFLV 1123
Score = 78.2 bits (191), Expect = 2e-14
Identities = 69/231 (29%), Positives = 105/231 (44%), Gaps = 33/231 (14%)
Query: 50 DGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDTTLK 108
DGS L TGS D ++ W + ++ C F+ H V V D L S S D T++
Sbjct: 1001 DGS-MLATGSGDQTVRLWDIS--SSQCFYIFQGHTSCVRSVVFSSDGAMLASGSDDQTVR 1057
Query: 109 TWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCID 168
WD S+G C TL+ H+ V + S + ++ASGG V +WD+ S+ C+
Sbjct: 1058 LWD-ISSGNCLYTLQGHTSCVRSVVFSPDGA-MLASGGDDQIVRLWDI-----SSGNCLY 1110
Query: 169 ATDDDTS---------NGIN-GSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHK 218
TS NG+ +G+S + L ISS + +T Q GH
Sbjct: 1111 TLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLWDISSKKCL--YTLQ----------GHT 1158
Query: 219 ESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
V A+A + G L SG ++ VR+WD S L+GHT + +++ +
Sbjct: 1159 NWVNAVAFSPDGATLASGSGDQTVRLWDISSSKCLYILQGHTSWVNSVVFN 1209
Score = 75.1 bits (183), Expect = 2e-13
Identities = 61/209 (29%), Positives = 101/209 (48%), Gaps = 15/209 (7%)
Query: 50 DGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDTTLK 108
DGS L +GS D ++ W + ++ C TF+ H WV+ D T L S S D T++
Sbjct: 1295 DGS-MLASGSGDQTVRLWEIS--SSKCLHTFQGHTSWVSSVTFSPDGTMLASGSDDQTVR 1351
Query: 109 TWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCID 168
W + S+G C T H+++V + S + I+ASG V +W + S+ KC+
Sbjct: 1352 LW-SISSGECLYTFLGHTNWVGSVIFSPDGA-ILASGSGDQTVRLWSI-----SSGKCL- 1403
Query: 169 ATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNE 228
T +N + S T L S S ++ + + G GH SV ++A +
Sbjct: 1404 YTLQGHNNWVGSIVFSPDGTLLASGSDDQTVRLWNISS-GECLYTLHGHINSVRSVAFSS 1462
Query: 229 GGTLLVSGGTEKVVRVWDPRSGS--KTMK 255
G +L SG ++ +++WD ++G KT+K
Sbjct: 1463 DGLILASGSDDETIKLWDVKTGECIKTLK 1491
Score = 72.0 bits (175), Expect = 1e-12
Identities = 59/222 (26%), Positives = 101/222 (44%), Gaps = 13/222 (5%)
Query: 49 SDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDTTL 107
SDG+ L +GS D ++ W + + C T + H V V D L S D +
Sbjct: 1042 SDGA-MLASGSDDQTVRLWDIS--SGNCLYTLQGHTSCVRSVVFSPDGAMLASGGDDQIV 1098
Query: 108 KTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCI 167
+ WD S+G C TL+ ++ +V L S N +A+G V +WD+ S+ KC+
Sbjct: 1099 RLWD-ISSGNCLYTLQGYTSWVRFLVFSP-NGVTLANGSSDQIVRLWDI-----SSKKCL 1151
Query: 168 DATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMN 227
T N +N S +L S S ++ + + + I +GH V ++ N
Sbjct: 1152 YTLQGHT-NWVNAVAFSPDGATLASGSGDQTVRLWDISSSKCLYIL-QGHTSWVNSVVFN 1209
Query: 228 EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
G+ L SG +++ VR+W+ S +GHT + +++ +
Sbjct: 1210 PDGSTLASGSSDQTVRLWEINSSKCLCTFQGHTSWVNSVVFN 1251
>YPR4_CAEEL (Q20059) Hypothetical WD-repeat protein F35G12.4 in
chromosome III
Length = 683
Score = 84.7 bits (208), Expect = 2e-16
Identities = 73/291 (25%), Positives = 124/291 (42%), Gaps = 41/291 (14%)
Query: 4 VGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGR 63
+ S NT S T P+K ++++++ D + + + ++ L + LFTG D
Sbjct: 1 MSSCVNT-SQTGPKK--KISFIIRDEHEYSNRSAVSALQY-----DAQNGRLFTGGSDTI 52
Query: 64 LKRWALGEDAATCSA-----------------TFESHVDWVNDAVLVGDST-LVSCSSDT 105
++ W++ SA + E H DWVND +L G L+S S+DT
Sbjct: 53 IRTWSVPHHKDAFSARGGVRSPGKNSPVQYQGSLEQHTDWVNDMILCGHGKILISASNDT 112
Query: 106 TLKTWDAFSTGT-----CTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAH 160
T+K W+ C RT H DYV+CLA + V S +FV+D+ A
Sbjct: 113 TVKVWNIERDNKHGFIDCIRT---HKDYVSCLAYAPIVEKAV-SASFDHNIFVYDINANF 168
Query: 161 ASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKES 220
+ I D I + L+ + + I + + I + +GH ++
Sbjct: 169 KTVNNLIGCKDS-----IYSLATTPNLSLVLGAGTEKCIRLFDPRTNEKI-MKLRGHTDN 222
Query: 221 VYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDST 271
V AL +N+ GT +S G++ +R+WD H + + L +DS+
Sbjct: 223 VRALVVNDDGTRALSAGSDATIRLWDIGQQRCIATCIAHEEGVWTLQVDSS 273
>PKWA_THECU (P49695) Probable serine/threonine-protein kinase pkwA
(EC 2.7.1.37)
Length = 742
Score = 84.7 bits (208), Expect = 2e-16
Identities = 67/228 (29%), Positives = 110/228 (47%), Gaps = 27/228 (11%)
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSS-DT 105
A S L +GS D ++ W + AA A FE H +V D D ++V+ S D
Sbjct: 508 AFSPDGALLASGSDDATVRLWDVA--AAEERAVFEGHTHYVLDIAFSPDGSMVASGSRDG 565
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDL------EAA 159
T + W+ +TGT L+ H+DYV +A S S +VASG G + +WD+ +
Sbjct: 566 TARLWNV-ATGTEHAVLKGHTDYVYAVAFSPDGS-MVASGSRDGTIRLWDVATGKERDVL 623
Query: 160 HASAAKCID-ATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHK 218
A A + A D S ++GS +++ L + S ++HT + GH
Sbjct: 624 QAPAENVVSLAFSPDGSMLVHGSDSTVHLWDVAS-----GEALHTFE----------GHT 668
Query: 219 ESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+ V A+A + G LL SG ++ +R+WD + + L+GHT+ + ++
Sbjct: 669 DWVRAVAFSPDGALLASGSDDRTIRLWDVAAQEEHTTLEGHTEPVHSV 716
Score = 48.9 bits (115), Expect = 1e-05
Identities = 33/86 (38%), Positives = 46/86 (53%), Gaps = 8/86 (9%)
Query: 184 SLPLTSLRSISS---SNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEK 240
+LP T LR SS S S H P +E+V A+A + GG+LL G +K
Sbjct: 427 TLPETPLRPDSSTAPSESADPHELNE----PRILTTDREAV-AVAFSPGGSLLAGGSGDK 481
Query: 241 VVRVWDPRSGSKTMKLKGHTDNIRAL 266
++ VWD SG + L+GHTD +RA+
Sbjct: 482 LIHVWDVASGDELHTLEGHTDWVRAV 507
Score = 38.9 bits (89), Expect = 0.014
Identities = 17/46 (36%), Positives = 26/46 (55%)
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHT 260
+GH + V A+A + G LL SG + VR+WD + + +GHT
Sbjct: 498 EGHTDWVRAVAFSPDGALLASGSDDATVRLWDVAAAEERAVFEGHT 543
>YH92_CAEEL (Q23256) Hypothetical WD-repeat protein ZC302.2 in
chromosome V
Length = 501
Score = 81.3 bits (199), Expect = 2e-15
Identities = 62/212 (29%), Positives = 100/212 (46%), Gaps = 21/212 (9%)
Query: 54 YLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDA 112
YL TGS D ++K W + T T SH +ND +S + S S DTT+K +D
Sbjct: 227 YLGTGSADKQIKVWNTVD--MTYLQTLASHQLGINDFSWSSNSQFIASASDDTTVKIFDV 284
Query: 113 FSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDA-TD 171
S G C RT+R H++YV C + + + S+++AS G V VWD + KCI A +D
Sbjct: 285 IS-GACLRTMRGHTNYVFCCSFNPQ-SSLIASAGFDETVRVWDFKT--GLCVKCIPAHSD 340
Query: 172 DDTSNGINGSGNSLPLTS----LRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMN 227
TS N GN++ +S +R +++ + T + + P+ + +
Sbjct: 341 PITSISYNHDGNTMATSSYDGCIRVWDAASGSCLKTLVDTDHAPVT---------FVCFS 391
Query: 228 EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
G L+S + +++WDP+ GH
Sbjct: 392 PNGKYLLSAQLDSSLKLWDPKKAKPLKYYNGH 423
Score = 38.1 bits (87), Expect = 0.023
Identities = 16/52 (30%), Positives = 29/52 (55%)
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+GH V+ + N +L+ S G ++ VRVWD ++G + H+D I ++
Sbjct: 294 RGHTNYVFCCSFNPQSSLIASAGFDETVRVWDFKTGLCVKCIPAHSDPITSI 345
>YS00_ANASP (Q8YTC2) Hypothetical WD-repeat protein alr2800
Length = 1258
Score = 80.5 bits (197), Expect = 4e-15
Identities = 64/220 (29%), Positives = 100/220 (45%), Gaps = 20/220 (9%)
Query: 49 SDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDS-TLVSCSSDTTL 107
S + L + D +K W++ + C T H V D TL S S D T+
Sbjct: 693 SPDGEILASCGADENVKLWSVRD--GVCIKTLTGHEHEVFSVAFHPDGETLASASGDKTI 750
Query: 108 KTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCI 167
K WD GTC +TL H+D+V C+A S + N +AS + +WD+ S KC+
Sbjct: 751 KLWD-IQDGTCLQTLTGHTDWVRCVAFSP-DGNTLASSAADHTIKLWDV-----SQGKCL 803
Query: 168 DATDDDTSNGINGSGNSLPLTSLRSISSSNSISV---HTTQN-QGYIPIAAKGHKESVYA 223
T + S +L S S +I + HT + + YI GH SVY+
Sbjct: 804 RTLKSHTG-WVRSVAFSADGQTLASGSGDRTIKIWNYHTGECLKTYI-----GHTNSVYS 857
Query: 224 LAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
+A + +LVSG ++ +++WD ++ L GHT+ +
Sbjct: 858 IAYSPDSKILVSGSGDRTIKLWDCQTHICIKTLHGHTNEV 897
Score = 79.0 bits (193), Expect = 1e-14
Identities = 64/233 (27%), Positives = 103/233 (43%), Gaps = 17/233 (7%)
Query: 32 TKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAV 91
T H + C+A S + L + + D +K W + + C T +SH WV
Sbjct: 765 TGHTDWVRCVAF-----SPDGNTLASSAADHTIKLWDVSQ--GKCLRTLKSHTGWVRSVA 817
Query: 92 LVGDS-TLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGE 150
D TL S S D T+K W+ + TG C +T H++ V +A S +S I+ SG
Sbjct: 818 FSADGQTLASGSGDRTIKIWN-YHTGECLKTYIGHTNSVYSIAYSP-DSKILVSGSGDRT 875
Query: 151 VFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYI 210
+ +WD + H T++ S + G +L SL S+ + + G
Sbjct: 876 IKLWDCQT-HICIKTLHGHTNEVCSVAFSPDGQTLACVSL-----DQSVRLWNCRT-GQC 928
Query: 211 PIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
A G+ + +A + +L SG +K V++WD ++G L+GHTD I
Sbjct: 929 LKAWYGNTDWALPVAFSPDRQILASGSNDKTVKLWDWQTGKYISSLEGHTDFI 981
Score = 75.1 bits (183), Expect = 2e-13
Identities = 59/219 (26%), Positives = 102/219 (45%), Gaps = 12/219 (5%)
Query: 44 LKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDS-TLVSCS 102
L A S L +GS D +K W ++ E H D++ DS TL S S
Sbjct: 940 LPVAFSPDRQILASGSNDKTVKLWDW--QTGKYISSLEGHTDFIYGIAFSPDSQTLASAS 997
Query: 103 SDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHAS 162
+D++++ W+ STG C + L +H+D+V + + I+A+G V +W++ S
Sbjct: 998 TDSSVRLWN-ISTGQCFQILLEHTDWVYAVVFHPQGK-IIATGSADCTVKLWNI-----S 1050
Query: 163 AAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVY 222
+C+ T + S+ I G S L S S+ S+ + + I +GH VY
Sbjct: 1051 TGQCLK-TLSEHSDKILGMAWSPDGQLLASASADQSVRLWDCCTGRCVGIL-RGHSNRVY 1108
Query: 223 ALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTD 261
+ + G ++ + T++ V++WD + G L GHT+
Sbjct: 1109 SAIFSPNGEIIATCSTDQTVKIWDWQQGKCLKTLTGHTN 1147
Score = 71.2 bits (173), Expect = 2e-12
Identities = 57/215 (26%), Positives = 93/215 (42%), Gaps = 12/215 (5%)
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSC-SSDT 105
A S S L + S D ++ W + C H DWV V +++ S+D
Sbjct: 985 AFSPDSQTLASASTDSSVRLWNIS--TGQCFQILLEHTDWVYAVVFHPQGKIIATGSADC 1042
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
T+K W+ STG C +TL +HSD + +A S + ++AS V +WD +
Sbjct: 1043 TVKLWN-ISTGQCLKTLSEHSDKILGMAWSP-DGQLLASASADQSVRLWD-----CCTGR 1095
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
C+ SN + + S + + S+ ++ + Q QG GH V+ +A
Sbjct: 1096 CVGILRGH-SNRVYSAIFSPNGEIIATCSTDQTVKIWDWQ-QGKCLKTLTGHTNWVFDIA 1153
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHT 260
+ G +L S ++ VR+WD +G GHT
Sbjct: 1154 FSPDGKILASASHDQTVRIWDVNTGKCHHICIGHT 1188
Score = 61.2 bits (147), Expect = 3e-09
Identities = 53/195 (27%), Positives = 82/195 (41%), Gaps = 12/195 (6%)
Query: 57 TGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAFST 115
TGS D +K W + C T H D + D L+ S S+D +++ WD T
Sbjct: 1037 TGSADCTVKLWNIS--TGQCLKTLSEHSDKILGMAWSPDGQLLASASADQSVRLWDC-CT 1093
Query: 116 GTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTS 175
G C LR HS+ V A N I+A+ V +WD + KC+ T
Sbjct: 1094 GRCVGILRGHSNRVYS-AIFSPNGEIIATCSTDQTVKIWDWQQG-----KCLKTLTGHT- 1146
Query: 176 NGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVS 235
N + S L S S ++ + N G GH V ++A + G ++ S
Sbjct: 1147 NWVFDIAFSPDGKILASASHDQTVRIWDV-NTGKCHHICIGHTHLVSSVAFSPDGEVVAS 1205
Query: 236 GGTEKVVRVWDPRSG 250
G ++ VR+W+ ++G
Sbjct: 1206 GSQDQTVRIWNVKTG 1220
Score = 47.4 bits (111), Expect = 4e-05
Identities = 56/239 (23%), Positives = 94/239 (38%), Gaps = 18/239 (7%)
Query: 36 AGINCLAVLKSAVSDGSDYLFTGS-------RDGRLKRWALGEDAATCSATFESHVDWVN 88
AG N L +L A D S Y F+G + L +C E+ + ++
Sbjct: 588 AGGNVLNLLHHAQVDLSGYDFSGLTVWQAYLQGVNLHDVDFANSDLSCCVFTETLGNILS 647
Query: 89 DAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLG 148
A L +C +D ++ W+ S G R HS++V + S + I+AS G
Sbjct: 648 AAFSPEGQLLATCDTDCHVRVWEVKS-GKLLLICRGHSNWVRFVVFSP-DGEILASCGAD 705
Query: 149 GEVFVWDLEAAHASAAKCIDATDDDT-SNGINGSGNSLPLTSLRSISSSNSISVHTTQNQ 207
V +W + K + + + S + G +L S S +I + Q+
Sbjct: 706 ENVKLWSVR--DGVCIKTLTGHEHEVFSVAFHPDGETLA-----SASGDKTIKLWDIQD- 757
Query: 208 GYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
G GH + V +A + G L S + +++WD G LK HT +R++
Sbjct: 758 GTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQGKCLRTLKSHTGWVRSV 816
>WDR5_MOUSE (P61965) WD-repeat protein 5 (WD-repeat protein BIG-3)
Length = 334
Score = 80.5 bits (197), Expect = 4e-15
Identities = 72/260 (27%), Positives = 109/260 (41%), Gaps = 44/260 (16%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTL-VS 100
AV S ++L + S D +K W G T H ++D DS L VS
Sbjct: 47 AVSSVKFSPNGEWLASSSADKLIKIW--GAYDGKFEKTISGHKLGISDVAWSSDSNLLVS 104
Query: 101 CSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA-- 158
S D TLK WD S+G C +TL+ HS+YV C + + SN++ SG V +WD++
Sbjct: 105 ASDDKTLKIWDV-SSGKCLKTLKGHSNYVFCCNFNPQ-SNLIVSGSFDESVRIWDVKTGK 162
Query: 159 ------AHASAAKCIDATDDDT---SNGING--------SGNSL---------PLTSLRS 192
AH+ + D + S+ +G SG L P++ ++
Sbjct: 163 CLKTLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKF 222
Query: 193 ISSSNSISVHTTQN--------QGYIPIAAKGHKESVYALAMN---EGGTLLVSGGTEKV 241
+ I T N +G GHK Y + N GG +VSG + +
Sbjct: 223 SPNGKYILAATLDNTLKLWDYSKGKCLKTYTGHKNEKYCIFANFSVTGGKWIVSGSEDNL 282
Query: 242 VRVWDPRSGSKTMKLKGHTD 261
V +W+ ++ KL+GHTD
Sbjct: 283 VYIWNLQTKEIVQKLQGHTD 302
Score = 47.0 bits (110), Expect = 5e-05
Identities = 45/203 (22%), Positives = 73/203 (35%), Gaps = 62/203 (30%)
Query: 64 LKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLR 123
LK G A S F + +W L S S+D +K W A+ G +T+
Sbjct: 37 LKFTLAGHTKAVSSVKFSPNGEW-----------LASSSADKLIKIWGAYD-GKFEKTIS 84
Query: 124 QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGN 183
H ++ +A S +SN++ S + +WD+ S+ KC+
Sbjct: 85 GHKLGISDVAWSS-DSNLLVSASDDKTLKIWDV-----SSGKCLKTL------------- 125
Query: 184 SLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVR 243
KGH V+ N L+VSG ++ VR
Sbjct: 126 -------------------------------KGHSNYVFCCNFNPQSNLIVSGSFDESVR 154
Query: 244 VWDPRSGSKTMKLKGHTDNIRAL 266
+WD ++G L H+D + A+
Sbjct: 155 IWDVKTGKCLKTLPAHSDPVSAV 177
Score = 32.0 bits (71), Expect = 1.7
Identities = 17/67 (25%), Positives = 30/67 (44%), Gaps = 1/67 (1%)
Query: 194 SSSNSISVHTTQNQGY-IPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSK 252
SSS + S T Y + GH ++V ++ + G L S +K++++W G
Sbjct: 20 SSSATQSKPTPVKPNYALKFTLAGHTKAVSSVKFSPNGEWLASSSADKLIKIWGAYDGKF 79
Query: 253 TMKLKGH 259
+ GH
Sbjct: 80 EKTISGH 86
>WDR5_HUMAN (P61964) WD-repeat protein 5
Length = 334
Score = 80.5 bits (197), Expect = 4e-15
Identities = 72/260 (27%), Positives = 109/260 (41%), Gaps = 44/260 (16%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTL-VS 100
AV S ++L + S D +K W G T H ++D DS L VS
Sbjct: 47 AVSSVKFSPNGEWLASSSADKLIKIW--GAYDGKFEKTISGHKLGISDVAWSSDSNLLVS 104
Query: 101 CSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA-- 158
S D TLK WD S+G C +TL+ HS+YV C + + SN++ SG V +WD++
Sbjct: 105 ASDDKTLKIWDV-SSGKCLKTLKGHSNYVFCCNFNPQ-SNLIVSGSFDESVRIWDVKTGK 162
Query: 159 ------AHASAAKCIDATDDDT---SNGING--------SGNSL---------PLTSLRS 192
AH+ + D + S+ +G SG L P++ ++
Sbjct: 163 CLKTLPAHSDPVSAVHFNRDGSLIVSSSYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKF 222
Query: 193 ISSSNSISVHTTQN--------QGYIPIAAKGHKESVYALAMN---EGGTLLVSGGTEKV 241
+ I T N +G GHK Y + N GG +VSG + +
Sbjct: 223 SPNGKYILAATLDNTLKLWDYSKGKCLKTYTGHKNEKYCIFANFSVTGGKWIVSGSEDNL 282
Query: 242 VRVWDPRSGSKTMKLKGHTD 261
V +W+ ++ KL+GHTD
Sbjct: 283 VYIWNLQTKEIVQKLQGHTD 302
Score = 47.0 bits (110), Expect = 5e-05
Identities = 45/203 (22%), Positives = 73/203 (35%), Gaps = 62/203 (30%)
Query: 64 LKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLR 123
LK G A S F + +W L S S+D +K W A+ G +T+
Sbjct: 37 LKFTLAGHTKAVSSVKFSPNGEW-----------LASSSADKLIKIWGAYD-GKFEKTIS 84
Query: 124 QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGN 183
H ++ +A S +SN++ S + +WD+ S+ KC+
Sbjct: 85 GHKLGISDVAWSS-DSNLLVSASDDKTLKIWDV-----SSGKCLKTL------------- 125
Query: 184 SLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVR 243
KGH V+ N L+VSG ++ VR
Sbjct: 126 -------------------------------KGHSNYVFCCNFNPQSNLIVSGSFDESVR 154
Query: 244 VWDPRSGSKTMKLKGHTDNIRAL 266
+WD ++G L H+D + A+
Sbjct: 155 IWDVKTGKCLKTLPAHSDPVSAV 177
Score = 32.0 bits (71), Expect = 1.7
Identities = 17/67 (25%), Positives = 30/67 (44%), Gaps = 1/67 (1%)
Query: 194 SSSNSISVHTTQNQGY-IPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSK 252
SSS + S T Y + GH ++V ++ + G L S +K++++W G
Sbjct: 20 SSSATQSKPTPVKPNYALKFTLAGHTKAVSSVKFSPNGEWLASSSADKLIKIWGAYDGKF 79
Query: 253 TMKLKGH 259
+ GH
Sbjct: 80 EKTISGH 86
>WDS_DROME (Q9V3J8) Will die slowly protein
Length = 361
Score = 79.3 bits (194), Expect = 9e-15
Identities = 82/300 (27%), Positives = 120/300 (39%), Gaps = 45/300 (15%)
Query: 4 VGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGR 63
VG G T SS K V + AG + AV S ++L + S D
Sbjct: 37 VGQPGATTSSNSSASNKSSLSVKPNYTLKFTLAG-HTKAVSAVKFSPNGEWLASSSADKL 95
Query: 64 LKRWALGEDAATCSATFESHVDWVNDAVLVGDSTL-VSCSSDTTLKTWDAFSTGTCTRTL 122
+K W G T H ++D DS L VS S D TLK W+ STG +TL
Sbjct: 96 IKIW--GAYDGKFEKTISGHKLGISDVAWSSDSRLLVSGSDDKTLKVWE-LSTGKSLKTL 152
Query: 123 RQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA--------AHASAAKCIDATDDDT 174
+ HS+YV C + + SN++ SG V +WD+ AH+ + D +
Sbjct: 153 KGHSNYVFCCNFNPQ-SNLIVSGSFDESVRIWDVRTGKCLKTLPAHSDPVSAVHFNRDGS 211
Query: 175 ---SNGING--------SGNSL---------PLTSLRSISSSNSISVHTTQN-------- 206
S+ +G SG L P++ ++ + I T N
Sbjct: 212 LIVSSSYDGLCRIWDTASGQCLKTLIDDDNPPVSFVKFSPNGKYILAATLDNTLKLWDYS 271
Query: 207 QGYIPIAAKGHKESVYALAMN---EGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
+G GHK Y + N GG +VSG + +V +W+ +S KL+GHTD +
Sbjct: 272 KGKCLKTYTGHKNEKYCIFANFSVTGGKWIVSGSEDNMVYIWNLQSKEVVQKLQGHTDTV 331
Score = 44.7 bits (104), Expect = 2e-04
Identities = 46/203 (22%), Positives = 72/203 (34%), Gaps = 62/203 (30%)
Query: 64 LKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLR 123
LK G A + F + +W L S S+D +K W A+ G +T+
Sbjct: 64 LKFTLAGHTKAVSAVKFSPNGEW-----------LASSSADKLIKIWGAYD-GKFEKTIS 111
Query: 124 QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGN 183
H ++ +A S +S ++ SG + VW+L +G
Sbjct: 112 GHKLGISDVAWSS-DSRLLVSGSDDKTLKVWELS-----------------------TGK 147
Query: 184 SLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVR 243
SL KGH V+ N L+VSG ++ VR
Sbjct: 148 SLK--------------------------TLKGHSNYVFCCNFNPQSNLIVSGSFDESVR 181
Query: 244 VWDPRSGSKTMKLKGHTDNIRAL 266
+WD R+G L H+D + A+
Sbjct: 182 IWDVRTGKCLKTLPAHSDPVSAV 204
>TUP1_YEAST (P16649) Glucose repression regulatory protein TUP1
(Flocculation suppressor protein) (Repressor AER2)
Length = 713
Score = 75.9 bits (185), Expect = 1e-13
Identities = 62/230 (26%), Positives = 99/230 (42%), Gaps = 21/230 (9%)
Query: 49 SDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLK 108
S +L TG+ D ++ W + + D GD LVS S D T++
Sbjct: 452 SPDGKFLATGAEDRLIRIWDIENRKIVMILQGHEQDIYSLDYFPSGDK-LVSGSGDRTVR 510
Query: 109 TWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAA--------- 159
WD TG C+ TL D VT +A S + +A+G L V VWD E
Sbjct: 511 IWD-LRTGQCSLTL-SIEDGVTTVAVSPGDGKYIAAGSLDRAVRVWDSETGFLVERLDSE 568
Query: 160 ------HASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIA 213
H + + T D S S+ L +L++ +N+ S T N G +
Sbjct: 569 NESGTGHKDSVYSVVFTRDGQSVVSGSLDRSVKLWNLQN---ANNKSDSKTPNSGTCEVT 625
Query: 214 AKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
GHK+ V ++A + ++SG ++ V WD +SG+ + L+GH +++
Sbjct: 626 YIGHKDFVLSVATTQNDEYILSGSKDRGVLFWDKKSGNPLLMLQGHRNSV 675
Score = 32.3 bits (72), Expect = 1.3
Identities = 22/110 (20%), Positives = 49/110 (44%), Gaps = 13/110 (11%)
Query: 170 TDDDTSNGINGS---GNSLPLTSLRSISSSNSISVHTTQNQGYIPIA--------AKGHK 218
+DD +N S N+ T +++++ + ++ TT +A +
Sbjct: 382 SDDSAANNHRNSITENNTTTSTDNNTMTTTTTTTITTTAMTSAAELAKDVENLNTSSSPS 441
Query: 219 ESVY--ALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+Y ++ + G L +G ++++R+WD + M L+GH +I +L
Sbjct: 442 SDLYIRSVCFSPDGKFLATGAEDRLIRIWDIENRKIVMILQGHEQDIYSL 491
>FBW7_MOUSE (Q8VBV4) F-box/WD-repeat protein 7 (F-box and WD-40
domain protein 7) (F-box protein FBW7) (F-box protein
Fbxw6) (F-box-WD40 repeat protein 6) (SEL-10)
Length = 629
Score = 75.5 bits (184), Expect = 1e-13
Identities = 71/251 (28%), Positives = 104/251 (41%), Gaps = 45/251 (17%)
Query: 17 RKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATC 76
R E + VL DD I CL + + GSD D LK W+ C
Sbjct: 289 RGELKSPKVLKGHDDHV----ITCLQFCGNRIVSGSD-------DNTLKVWSAV--TGKC 335
Query: 77 SATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASE 136
T H V + + D+ ++S S+D TLK W+A TG C TL H+ V C+ E
Sbjct: 336 LRTLVGHTGGVWSSQM-RDNIIISGSTDRTLKVWNA-ETGECIHTLYGHTSTVRCMHLHE 393
Query: 137 KNSNIVASGGLGGEVFVWDLEAA--------HASAAKCIDATDDDTSNGINGSGNSLPLT 188
K V SG + VWD+E H +A +C+ D ++G+ + +
Sbjct: 394 KR---VVSGSRDATLRVWDIETGQCLHVLMGHVAAVRCVQY---DGRRVVSGAYDFM--- 444
Query: 189 SLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPR 248
++ +HT Q GH VY+L + G +VSG + +RVWD
Sbjct: 445 -VKVWDPETETCLHTLQ----------GHTNRVYSLQFD--GIHVVSGSLDTSIRVWDVE 491
Query: 249 SGSKTMKLKGH 259
+G+ L GH
Sbjct: 492 TGNCIHTLTGH 502
Score = 64.7 bits (156), Expect = 2e-10
Identities = 58/204 (28%), Positives = 88/204 (42%), Gaps = 19/204 (9%)
Query: 67 WALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHS 126
W GE + H D V + + +VS S D TLK W A TG C RTL H+
Sbjct: 287 WRRGE--LKSPKVLKGHDDHVITCLQFCGNRIVSGSDDNTLKVWSAV-TGKCLRTLVGHT 343
Query: 127 DYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLP 186
V +S+ NI+ SG + VW+ E +CI T G + +
Sbjct: 344 GGVW---SSQMRDNIIISGSTDRTLKVWNAETG-----ECIH-----TLYGHTSTVRCMH 390
Query: 187 LTSLRSISSSNSISVHTTQNQ-GYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVW 245
L R +S S ++ + G GH +V + + G +VSG + +V+VW
Sbjct: 391 LHEKRVVSGSRDATLRVWDIETGQCLHVLMGHVAAVRCVQYD--GRRVVSGAYDFMVKVW 448
Query: 246 DPRSGSKTMKLKGHTDNIRALLLD 269
DP + + L+GHT+ + +L D
Sbjct: 449 DPETETCLHTLQGHTNRVYSLQFD 472
Score = 52.0 bits (123), Expect = 2e-06
Identities = 34/107 (31%), Positives = 56/107 (51%), Gaps = 10/107 (9%)
Query: 54 YLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAF 113
++ +GS D ++ W + + C T H + + + D+ LVS ++D+T+K WD
Sbjct: 475 HVVSGSLDTSIRVWDV--ETGNCIHTLTGHQS-LTSGMELKDNILVSGNADSTVKIWD-I 530
Query: 114 STGTCTRTLR---QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
TG C +TL+ +H VTCL + N N V + G V +WDL+
Sbjct: 531 KTGQCLQTLQGPSKHQSAVTCL---QFNKNFVITSSDDGTVKLWDLK 574
>FBW7_HUMAN (Q969H0) F-box/WD-repeat protein 7 (F-box and WD-40
domain protein 7) (F-box protein FBX30) (hCdc4)
(Archipelago homolog) (hAgo) (SEL-10)
Length = 707
Score = 75.5 bits (184), Expect = 1e-13
Identities = 71/251 (28%), Positives = 104/251 (41%), Gaps = 45/251 (17%)
Query: 17 RKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATC 76
R E + VL DD I CL + + GSD D LK W+ C
Sbjct: 367 RGELKSPKVLKGHDDHV----ITCLQFCGNRIVSGSD-------DNTLKVWSAV--TGKC 413
Query: 77 SATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASE 136
T H V + + D+ ++S S+D TLK W+A TG C TL H+ V C+ E
Sbjct: 414 LRTLVGHTGGVWSSQM-RDNIIISGSTDRTLKVWNA-ETGECIHTLYGHTSTVRCMHLHE 471
Query: 137 KNSNIVASGGLGGEVFVWDLEAA--------HASAAKCIDATDDDTSNGINGSGNSLPLT 188
K V SG + VWD+E H +A +C+ D ++G+ + +
Sbjct: 472 KR---VVSGSRDATLRVWDIETGQCLHVLMGHVAAVRCVQY---DGRRVVSGAYDFM--- 522
Query: 189 SLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPR 248
++ +HT Q GH VY+L + G +VSG + +RVWD
Sbjct: 523 -VKVWDPETETCLHTLQ----------GHTNRVYSLQFD--GIHVVSGSLDTSIRVWDVE 569
Query: 249 SGSKTMKLKGH 259
+G+ L GH
Sbjct: 570 TGNCIHTLTGH 580
Score = 64.7 bits (156), Expect = 2e-10
Identities = 58/204 (28%), Positives = 88/204 (42%), Gaps = 19/204 (9%)
Query: 67 WALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHS 126
W GE + H D V + + +VS S D TLK W A TG C RTL H+
Sbjct: 365 WRRGE--LKSPKVLKGHDDHVITCLQFCGNRIVSGSDDNTLKVWSAV-TGKCLRTLVGHT 421
Query: 127 DYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLP 186
V +S+ NI+ SG + VW+ E +CI T G + +
Sbjct: 422 GGVW---SSQMRDNIIISGSTDRTLKVWNAETG-----ECIH-----TLYGHTSTVRCMH 468
Query: 187 LTSLRSISSSNSISVHTTQNQ-GYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVW 245
L R +S S ++ + G GH +V + + G +VSG + +V+VW
Sbjct: 469 LHEKRVVSGSRDATLRVWDIETGQCLHVLMGHVAAVRCVQYD--GRRVVSGAYDFMVKVW 526
Query: 246 DPRSGSKTMKLKGHTDNIRALLLD 269
DP + + L+GHT+ + +L D
Sbjct: 527 DPETETCLHTLQGHTNRVYSLQFD 550
Score = 52.0 bits (123), Expect = 2e-06
Identities = 34/107 (31%), Positives = 56/107 (51%), Gaps = 10/107 (9%)
Query: 54 YLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAF 113
++ +GS D ++ W + + C T H + + + D+ LVS ++D+T+K WD
Sbjct: 553 HVVSGSLDTSIRVWDV--ETGNCIHTLTGHQS-LTSGMELKDNILVSGNADSTVKIWD-I 608
Query: 114 STGTCTRTLR---QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
TG C +TL+ +H VTCL + N N V + G V +WDL+
Sbjct: 609 KTGQCLQTLQGPNKHQSAVTCL---QFNKNFVITSSDDGTVKLWDLK 652
>RCO1_NEUCR (P78706) Transcriptional repressor rco-1
Length = 604
Score = 73.2 bits (178), Expect = 6e-13
Identities = 62/223 (27%), Positives = 98/223 (43%), Gaps = 13/223 (5%)
Query: 49 SDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDS-TLVSCSSDTTL 107
S YL TG+ D ++ W + + T TF H + D T+ S S D T+
Sbjct: 353 SPDGKYLATGAEDKLIRVWDI--QSRTIRNTFHGHEQDIYSLDFSRDGRTIASGSGDRTV 410
Query: 108 KTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCI 167
+ WD TG T L D VT +A S + VA+G L V VWD+ A +
Sbjct: 411 RLWD-IETGQNTSVL-SIEDGVTTVAISP-DKQFVAAGSLDKSVRVWDMRGYLAERLEGP 467
Query: 168 DATDDD------TSNGINGSGNSLPLT-SLRSISSSNSISVHTTQNQGYIPIAAKGHKES 220
D D + +G N SL T + +S+ I G +GH++
Sbjct: 468 DGHKDSVYSVAFSPDGRNLVSGSLDKTIKMWELSAPRGIPSSAPPKGGRCIKTFEGHRDF 527
Query: 221 VYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
V ++A+ ++SG ++ V+ WDPR+G + L+GH +++
Sbjct: 528 VLSVALTPDSQWVLSGSKDRGVQFWDPRTGHTQLMLQGHKNSV 570
Score = 30.4 bits (67), Expect = 4.8
Identities = 13/46 (28%), Positives = 25/46 (54%)
Query: 221 VYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+ ++ + G L +G +K++RVWD +S + GH +I +L
Sbjct: 347 IRSVCFSPDGKYLATGAEDKLIRVWDIQSRTIRNTFHGHEQDIYSL 392
>TUP1_KLULA (P56094) Transcriptional repressor TUP1
Length = 682
Score = 72.4 bits (176), Expect = 1e-12
Identities = 60/231 (25%), Positives = 96/231 (40%), Gaps = 28/231 (12%)
Query: 49 SDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHV-DWVNDAVLVGDSTLVSCSSDTTL 107
S +L TG+ D ++ W L + T + H D + + LVS S D T+
Sbjct: 415 SPDGKFLATGAEDKLIRIWDL--ETKKIVMTLKGHEQDIYSLDYFPSGNKLVSGSGDRTV 472
Query: 108 KTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD------------ 155
+ WD +TGTC+ TL D VT +A S +A+G L V VWD
Sbjct: 473 RIWD-LTTGTCSLTL-SIEDGVTTVAVSPGEGKFIAAGSLDRTVRVWDSDTGFLVERLDS 530
Query: 156 ---LEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPI 212
L H + + T D S+ L +L +S S +
Sbjct: 531 ENELGTGHRDSVYSVVFTRDGKGVVSGSLDRSVKLWNLNGLSGQKS--------HAECEV 582
Query: 213 AAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNI 263
GHK+ V ++A + ++SG ++ V WD +SG+ + L+GH +++
Sbjct: 583 TYTGHKDFVLSVATTQNDEYILSGSKDRGVLFWDTKSGNPLLMLQGHRNSV 633
Score = 42.7 bits (99), Expect = 0.001
Identities = 35/146 (23%), Positives = 61/146 (40%), Gaps = 13/146 (8%)
Query: 124 QHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGN 183
+HS V C+ S + +V+ S + + DD+++
Sbjct: 319 EHSSVVCCVRFSSDGEFLATGCNKTTQVY-------KVSTGELVARLSDDSASQPQPQPQ 371
Query: 184 SLPLTSLRSISSSNSISVHT-TQNQGYIPIAAKGHKESVY--ALAMNEGGTLLVSGGTEK 240
+ +T+ S S+SN S T NQ AA +Y ++ + G L +G +K
Sbjct: 372 NQTVTAETSTSNSNGSSAEDGTGNQNS---AASTASSDLYIRSVCFSPDGKFLATGAEDK 428
Query: 241 VVRVWDPRSGSKTMKLKGHTDNIRAL 266
++R+WD + M LKGH +I +L
Sbjct: 429 LIRIWDLETKKIVMTLKGHEQDIYSL 454
>SE10_CAEEL (Q93794) F-box/WD-repeat protein sel-10
(Suppressor/enhancer of lin-12) (Sel-10 protein)
Length = 587
Score = 71.2 bits (173), Expect = 2e-12
Identities = 70/230 (30%), Positives = 98/230 (42%), Gaps = 25/230 (10%)
Query: 43 VLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCS 102
V S +S Y+ +GS D +K W+ + + T + H V + G S LV+ S
Sbjct: 299 VWTSQISQCGRYIVSGSTDRTVKVWSTVDGSLL--HTLQGHTSTVRCMAMAG-SILVTGS 355
Query: 103 SDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHAS 162
DTTL+ WD S G TL H V C+ + + V SGG V +W+ A
Sbjct: 356 RDTTLRVWDVES-GRHLATLHGHHAAVRCV---QFDGTTVVSGGYDFTVKIWN-----AH 406
Query: 163 AAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHT-----TQNQGYIPIAA-KG 216
+CI T G N SL S RSI S S+ T+ +G +A +G
Sbjct: 407 TGRCIR-----TLTGHNNRVYSLLFESERSIVCSGSLDTSIRVWDFTRPEGQECVALLQG 461
Query: 217 HKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
H + + G +LVS + VRVWD G+ L GH I +L
Sbjct: 462 HTSLTSGMQLR--GNILVSCNADSHVRVWDIHEGTCVHMLSGHRSAITSL 509
Score = 56.2 bits (134), Expect = 8e-08
Identities = 51/218 (23%), Positives = 85/218 (38%), Gaps = 56/218 (25%)
Query: 53 DYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDA 112
D L TGS D LK W + + + + W + G +VS S+D T+K W
Sbjct: 267 DVLVTGSDDNTLKVWCIDKGEVMYTLVGHTGGVWTSQISQCG-RYIVSGSTDRTVKVWST 325
Query: 113 FSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDD 172
G+ TL+ H+ V C+A + +I+ +G + VWD+E+ A
Sbjct: 326 VD-GSLLHTLQGHTSTVRCMAMA---GSILVTGSRDTTLRVWDVESGRHLA--------- 372
Query: 173 DTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTL 232
++H GH +V + + GT
Sbjct: 373 ---------------------------TLH-------------GHHAAVRCVQFD--GTT 390
Query: 233 LVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+VSGG + V++W+ +G L GH + + +LL +S
Sbjct: 391 VVSGGYDFTVKIWNAHTGRCIRTLTGHNNRVYSLLFES 428
Score = 50.8 bits (120), Expect = 3e-06
Identities = 53/195 (27%), Positives = 70/195 (35%), Gaps = 53/195 (27%)
Query: 77 SATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASE 136
SA H D V + + D LV+ S D TLK W G TL H+ V S+
Sbjct: 248 SAVLRGHEDHVITCMQIHDDVLVTGSDDNTLKVW-CIDKGEVMYTLVGHTGGVWTSQISQ 306
Query: 137 KNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSS 196
IV SG V VW ++GS
Sbjct: 307 CGRYIV-SGSTDRTVKVWST---------------------VDGS--------------- 329
Query: 197 NSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKL 256
+HT Q GH +V +AM G++LV+G + +RVWD SG L
Sbjct: 330 ---LLHTLQ----------GHTSTVRCMAM--AGSILVTGSRDTTLRVWDVESGRHLATL 374
Query: 257 KGHTDNIRALLLDST 271
GH +R + D T
Sbjct: 375 HGHHAAVRCVQFDGT 389
Score = 47.4 bits (111), Expect = 4e-05
Identities = 31/102 (30%), Positives = 49/102 (47%), Gaps = 5/102 (4%)
Query: 57 TGSRDGRLKRWALGE-DAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFST 115
+GS D ++ W + C A + H + L G+ LVSC++D+ ++ WD
Sbjct: 435 SGSLDTSIRVWDFTRPEGQECVALLQGHTSLTSGMQLRGN-ILVSCNADSHVRVWDIHE- 492
Query: 116 GTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLE 157
GTC L H +T L +N +VA+ G V +WD+E
Sbjct: 493 GTCVHMLSGHRSAITSLQWFGRN--MVATSSDDGTVKLWDIE 532
>Y163_SYNY3 (Q55563) Hypothetical WD-repeat protein sll0163
Length = 1693
Score = 70.1 bits (170), Expect = 5e-12
Identities = 58/212 (27%), Positives = 92/212 (43%), Gaps = 19/212 (8%)
Query: 55 LFTGSRDGRLKRWAL-GEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDTTLKTWDA 112
L T SRDG + W L G + C + H WV +A D +V+CS+D T + WD
Sbjct: 1152 LLTASRDGTARLWDLEGREIGLC----QGHTSWVRNAQFSPDGQWIVTCSADGTARLWDL 1207
Query: 113 FSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDD 172
S C L+ H ++V S +I+ S G VW KC+
Sbjct: 1208 SSQ--CFAVLKGHQNWVNNALWSPDGQHIITSSS-DGTARVWSRHG------KCLGTLRG 1258
Query: 173 DTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTL 232
N I+G+ SL + + S+ N+ + T +G + +GH++ VY + G
Sbjct: 1259 HDHN-IHGARFSLDGQKIVTYSTDNTARLWT--KEGTLLTILRGHQKEVYDADFSADGRF 1315
Query: 233 LVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIR 264
+ + ++ R WD S T+ L GH+ +R
Sbjct: 1316 VFTVSADQTARQWD-ISQKDTITLTGHSHWVR 1346
Score = 47.8 bits (112), Expect = 3e-05
Identities = 60/241 (24%), Positives = 90/241 (36%), Gaps = 42/241 (17%)
Query: 53 DYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVSCSSDTTLKTWD 111
D L T SRD + W + C A H WV + D +V+ S+D T + W+
Sbjct: 1355 DRLLTVSRDKTARLWTTEGE---CVAVLADHQGWVREGQFSPDGQWIVTGSADKTAQLWN 1411
Query: 112 AFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD---------------L 156
G LR H D V + S +S + + G VW+ +
Sbjct: 1412 VL--GKKLTVLRGHQDAVLNVRFSP-DSQYIVTASKDGTARVWNNTGRELAVLRHYEKNI 1468
Query: 157 EAAHASAAK--CIDATDDDTSN---------GINGSGNSLPLTSLRSISSSNSISVHTTQ 205
AA SA + A+DD+T+ GI G+ P+ + + S I +
Sbjct: 1469 FAAEFSADGQFIVTASDDNTAGIWEIVGREVGIC-RGHEGPVYFAQFSADSRYILTASVD 1527
Query: 206 NQGYI-------PIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKG 258
N I + GH+ VY + G L+ + + R+WD RSG L G
Sbjct: 1528 NTARIWDFLGRPLLTLAGHQSIVYQARFSPEGNLIATVSADHTARLWD-RSGKTVAVLYG 1586
Query: 259 H 259
H
Sbjct: 1587 H 1587
Score = 35.4 bits (80), Expect = 0.15
Identities = 15/52 (28%), Positives = 29/52 (54%), Gaps = 1/52 (1%)
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+GH+E++ AL + G + + V++W R G + L+GH D +R++
Sbjct: 1052 RGHQEAITALDWSADGQYFATASADHTVKLWQ-RHGEEVATLRGHEDWVRSV 1102
Score = 35.4 bits (80), Expect = 0.15
Identities = 50/234 (21%), Positives = 83/234 (35%), Gaps = 35/234 (14%)
Query: 42 AVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDST-LVS 100
AVL S S Y+ T S+DG + W + A + + A D +V+
Sbjct: 1426 AVLNVRFSPDSQYIVTASKDGTARVW---NNTGRELAVLRHYEKNIFAAEFSADGQFIVT 1482
Query: 101 CSSDTTLKTWDAFS--TGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA 158
S D T W+ G C R H V A +S + + + +WD
Sbjct: 1483 ASDDNTAGIWEIVGREVGIC----RGHEGPVY-FAQFSADSRYILTASVDNTARIWDFLG 1537
Query: 159 AHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISV----HTTQ---NQGYIP 211
+ +G+ + R N I+ HT + G
Sbjct: 1538 RPL----------------LTLAGHQSIVYQARFSPEGNLIATVSADHTARLWDRSGKTV 1581
Query: 212 IAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRA 265
GH+ V + + G +LV+ + R+WD SG + + L+GH + +R+
Sbjct: 1582 AVLYGHQGLVGTVDWSPDGQMLVTASNDGTARLWD-LSGRELLTLEGHGNWVRS 1634
Score = 32.0 bits (71), Expect = 1.7
Identities = 51/221 (23%), Positives = 81/221 (36%), Gaps = 26/221 (11%)
Query: 39 NCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTL 98
N ++ +A ++G YL+ S G L G A + +DW D
Sbjct: 1024 NTPPLVLTATTNGIAYLW--SFHGELINVLRGHQEAITA------LDWSADG-----QYF 1070
Query: 99 VSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEA 158
+ S+D T+K W G TLR H D+V + S + +V S G +W+
Sbjct: 1071 ATASADHTVKLWQRH--GEEVATLRGHEDWVRSVHFSPHHQFLVTS-GQDNTARIWNF-- 1125
Query: 159 AHASAAKCIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHK 218
A C D + N G L +++S + +G +GH
Sbjct: 1126 AGEQLTLCQGHADWVRNAEFNCHGQIL-------LTASRDGTARLWDLEGREIGLCQGHT 1178
Query: 219 ESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
V + G +V+ + R+WD S + LKGH
Sbjct: 1179 SWVRNAQFSPDGQWIVTCSADGTARLWDLSSQCFAV-LKGH 1218
Score = 30.0 bits (66), Expect = 6.3
Identities = 17/71 (23%), Positives = 34/71 (46%), Gaps = 1/71 (1%)
Query: 194 SSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKT 253
++S +V Q G +GH++ V ++ + LV+ G + R+W+ +G +
Sbjct: 1072 TASADHTVKLWQRHGEEVATLRGHEDWVRSVHFSPHHQFLVTSGQDNTARIWN-FAGEQL 1130
Query: 254 MKLKGHTDNIR 264
+GH D +R
Sbjct: 1131 TLCQGHADWVR 1141
>LIS1_RAT (P63004) Platelet-activating factor acetylhydrolase IB
alpha subunit (PAF acetylhydrolase 45 kDa subunit)
(PAF-AH 45 kDa subunit) (PAF-AH alpha) (PAFAH alpha)
(Lissencephaly-1 protein) (LIS-1)
Length = 409
Score = 68.6 bits (166), Expect = 2e-11
Identities = 57/221 (25%), Positives = 92/221 (40%), Gaps = 34/221 (15%)
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDT 105
A+ D++ + SRD +K W + C TF H +WV D TL+ SCS+D
Sbjct: 198 AIMPNGDHIVSASRDKTIKMWEV--QTGYCVKTFTGHREWVRMVRPNQDGTLIASCSNDQ 255
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
T++ W +T C LR+H V C++ W E++++S +
Sbjct: 256 TVRVW-VVATKECKAELREHEHVVECIS--------------------WAPESSYSSIS- 293
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
+AT +T SG P L S S +I + G + GH V +
Sbjct: 294 --EATGSETKK----SGKPGPF--LLSGSRDKTIKMWDVST-GMCLMTLVGHDNWVRGVL 344
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+ GG ++S +K +RVWD ++ L H + +L
Sbjct: 345 FHSGGKFILSCADDKTLRVWDYKNKRCMKTLNAHEHFVTSL 385
Score = 58.5 bits (140), Expect = 2e-08
Identities = 46/165 (27%), Positives = 78/165 (46%), Gaps = 11/165 (6%)
Query: 96 STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD 155
S +VS S D T+K WD + TG RTL+ H+D V ++ + + ++AS + +WD
Sbjct: 120 SVMVSASEDATIKVWD-YETGDFERTLKGHTDSVQDISF-DHSGKLLASCSADMTIKLWD 177
Query: 156 LEAAHASAAKCIDATDDDTSN-GINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAA 214
+ + + D + S+ I +G+ + S S +I + Q GY
Sbjct: 178 FQGFEC--IRTMHGHDHNVSSVAIMPNGDHIV-----SASRDKTIKMWEVQT-GYCVKTF 229
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
GH+E V + N+ GTL+ S ++ VRVW + +L+ H
Sbjct: 230 TGHREWVRMVRPNQDGTLIASCSNDQTVRVWVVATKECKAELREH 274
Score = 57.4 bits (137), Expect = 4e-08
Identities = 52/230 (22%), Positives = 93/230 (39%), Gaps = 18/230 (7%)
Query: 55 LFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAF 113
+ + S D +K W + T + H D V D L+ SCS+D T+K WD F
Sbjct: 122 MVSASEDATIKVWDY--ETGDFERTLKGHTDSVQDISFDHSGKLLASCSADMTIKLWD-F 178
Query: 114 STGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAA--------HASAAK 165
C RT+ H V+ +A +IV S + +W+++ H +
Sbjct: 179 QGFECIRTMHGHDHNVSSVAIMPNGDHIV-SASRDKTIKMWEVQTGYCVKTFTGHREWVR 237
Query: 166 CIDATDDDT--SNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGH-KESVY 222
+ D T ++ N + + + + + H + + P ++ E+
Sbjct: 238 MVRPNQDGTLIASCSNDQTVRVWVVATKECKAELREHEHVVECISWAPESSYSSISEATG 297
Query: 223 ALAMNEG--GTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+ G G L+SG +K +++WD +G M L GH + +R +L S
Sbjct: 298 SETKKSGKPGPFLLSGSRDKTIKMWDVSTGMCLMTLVGHDNWVRGVLFHS 347
Score = 40.0 bits (92), Expect = 0.006
Identities = 17/57 (29%), Positives = 31/57 (53%)
Query: 213 AAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
A GH+ V + + +++VS + ++VWD +G LKGHTD+++ + D
Sbjct: 102 ALSGHRSPVTRVIFHPVFSVMVSASEDATIKVWDYETGDFERTLKGHTDSVQDISFD 158
>LIS1_MOUSE (P63005) Platelet-activating factor acetylhydrolase IB
alpha subunit (PAF acetylhydrolase 45 kDa subunit)
(PAF-AH 45 kDa subunit) (PAF-AH alpha) (PAFAH alpha)
(Lissencephaly-1 protein) (LIS-1)
Length = 409
Score = 68.6 bits (166), Expect = 2e-11
Identities = 57/221 (25%), Positives = 92/221 (40%), Gaps = 34/221 (15%)
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDT 105
A+ D++ + SRD +K W + C TF H +WV D TL+ SCS+D
Sbjct: 198 AIMPNGDHIVSASRDKTIKMWEV--QTGYCVKTFTGHREWVRMVRPNQDGTLIASCSNDQ 255
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
T++ W +T C LR+H V C++ W E++++S +
Sbjct: 256 TVRVW-VVATKECKAELREHEHVVECIS--------------------WAPESSYSSIS- 293
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
+AT +T SG P L S S +I + G + GH V +
Sbjct: 294 --EATGSETKK----SGKPGPF--LLSGSRDKTIKMWDVST-GMCLMTLVGHDNWVRGVL 344
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+ GG ++S +K +RVWD ++ L H + +L
Sbjct: 345 FHSGGKFILSCADDKTLRVWDYKNKRCMKTLNAHEHFVTSL 385
Score = 58.5 bits (140), Expect = 2e-08
Identities = 46/165 (27%), Positives = 78/165 (46%), Gaps = 11/165 (6%)
Query: 96 STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD 155
S +VS S D T+K WD + TG RTL+ H+D V ++ + + ++AS + +WD
Sbjct: 120 SVMVSASEDATIKVWD-YETGDFERTLKGHTDSVQDISF-DHSGKLLASCSADMTIKLWD 177
Query: 156 LEAAHASAAKCIDATDDDTSN-GINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAA 214
+ + + D + S+ I +G+ + S S +I + Q GY
Sbjct: 178 FQGFEC--IRTMHGHDHNVSSVAIMPNGDHIV-----SASRDKTIKMWEVQT-GYCVKTF 229
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
GH+E V + N+ GTL+ S ++ VRVW + +L+ H
Sbjct: 230 TGHREWVRMVRPNQDGTLIASCSNDQTVRVWVVATKECKAELREH 274
Score = 57.4 bits (137), Expect = 4e-08
Identities = 52/230 (22%), Positives = 93/230 (39%), Gaps = 18/230 (7%)
Query: 55 LFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAF 113
+ + S D +K W + T + H D V D L+ SCS+D T+K WD F
Sbjct: 122 MVSASEDATIKVWDY--ETGDFERTLKGHTDSVQDISFDHSGKLLASCSADMTIKLWD-F 178
Query: 114 STGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAA--------HASAAK 165
C RT+ H V+ +A +IV S + +W+++ H +
Sbjct: 179 QGFECIRTMHGHDHNVSSVAIMPNGDHIV-SASRDKTIKMWEVQTGYCVKTFTGHREWVR 237
Query: 166 CIDATDDDT--SNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGH-KESVY 222
+ D T ++ N + + + + + H + + P ++ E+
Sbjct: 238 MVRPNQDGTLIASCSNDQTVRVWVVATKECKAELREHEHVVECISWAPESSYSSISEATG 297
Query: 223 ALAMNEG--GTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+ G G L+SG +K +++WD +G M L GH + +R +L S
Sbjct: 298 SETKKSGKPGPFLLSGSRDKTIKMWDVSTGMCLMTLVGHDNWVRGVLFHS 347
Score = 40.0 bits (92), Expect = 0.006
Identities = 17/57 (29%), Positives = 31/57 (53%)
Query: 213 AAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
A GH+ V + + +++VS + ++VWD +G LKGHTD+++ + D
Sbjct: 102 ALSGHRSPVTRVIFHPVFSVMVSASEDATIKVWDYETGDFERTLKGHTDSVQDISFD 158
>LIS1_HUMAN (P43034) Platelet-activating factor acetylhydrolase IB
alpha subunit (PAF acetylhydrolase 45 kDa subunit)
(PAF-AH 45 kDa subunit) (PAF-AH alpha) (PAFAH alpha)
(Lissencephaly-1 protein) (LIS-1)
Length = 409
Score = 68.6 bits (166), Expect = 2e-11
Identities = 57/221 (25%), Positives = 92/221 (40%), Gaps = 34/221 (15%)
Query: 47 AVSDGSDYLFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDT 105
A+ D++ + SRD +K W + C TF H +WV D TL+ SCS+D
Sbjct: 198 AIMPNGDHIVSASRDKTIKMWEV--QTGYCVKTFTGHREWVRMVRPNQDGTLIASCSNDQ 255
Query: 106 TLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAK 165
T++ W +T C LR+H V C++ W E++++S +
Sbjct: 256 TVRVW-VVATKECKAELREHEHVVECIS--------------------WAPESSYSSIS- 293
Query: 166 CIDATDDDTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALA 225
+AT +T SG P L S S +I + G + GH V +
Sbjct: 294 --EATGSETKK----SGKPGPF--LLSGSRDKTIKMWDVST-GMCLMTLVGHDNWVRGVL 344
Query: 226 MNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRAL 266
+ GG ++S +K +RVWD ++ L H + +L
Sbjct: 345 FHSGGKFILSCADDKTLRVWDYKNKRCMKTLNAHEHFVTSL 385
Score = 58.5 bits (140), Expect = 2e-08
Identities = 46/165 (27%), Positives = 78/165 (46%), Gaps = 11/165 (6%)
Query: 96 STLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWD 155
S +VS S D T+K WD + TG RTL+ H+D V ++ + + ++AS + +WD
Sbjct: 120 SVMVSASEDATIKVWD-YETGDFERTLKGHTDSVQDISF-DHSGKLLASCSADMTIKLWD 177
Query: 156 LEAAHASAAKCIDATDDDTSN-GINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAA 214
+ + + D + S+ I +G+ + S S +I + Q GY
Sbjct: 178 FQGFEC--IRTMHGHDHNVSSVAIMPNGDHIV-----SASRDKTIKMWEVQT-GYCVKTF 229
Query: 215 KGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGH 259
GH+E V + N+ GTL+ S ++ VRVW + +L+ H
Sbjct: 230 TGHREWVRMVRPNQDGTLIASCSNDQTVRVWVVATKECKAELREH 274
Score = 57.4 bits (137), Expect = 4e-08
Identities = 52/230 (22%), Positives = 93/230 (39%), Gaps = 18/230 (7%)
Query: 55 LFTGSRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLV-SCSSDTTLKTWDAF 113
+ + S D +K W + T + H D V D L+ SCS+D T+K WD F
Sbjct: 122 MVSASEDATIKVWDY--ETGDFERTLKGHTDSVQDISFDHSGKLLASCSADMTIKLWD-F 178
Query: 114 STGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAA--------HASAAK 165
C RT+ H V+ +A +IV S + +W+++ H +
Sbjct: 179 QGFECIRTMHGHDHNVSSVAIMPNGDHIV-SASRDKTIKMWEVQTGYCVKTFTGHREWVR 237
Query: 166 CIDATDDDT--SNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGH-KESVY 222
+ D T ++ N + + + + + H + + P ++ E+
Sbjct: 238 MVRPNQDGTLIASCSNDQTVRVWVVATKECKAELREHEHVVECISWAPESSYSSISEATG 297
Query: 223 ALAMNEG--GTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLDS 270
+ G G L+SG +K +++WD +G M L GH + +R +L S
Sbjct: 298 SETKKSGKPGPFLLSGSRDKTIKMWDVSTGMCLMTLVGHDNWVRGVLFHS 347
Score = 40.0 bits (92), Expect = 0.006
Identities = 17/57 (29%), Positives = 31/57 (53%)
Query: 213 AAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
A GH+ V + + +++VS + ++VWD +G LKGHTD+++ + D
Sbjct: 102 ALSGHRSPVTRVIFHPVFSVMVSASEDATIKVWDYETGDFERTLKGHTDSVQDISFD 158
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.127 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,496,801
Number of Sequences: 164201
Number of extensions: 1308496
Number of successful extensions: 5047
Number of sequences better than 10.0: 413
Number of HSP's better than 10.0 without gapping: 302
Number of HSP's successfully gapped in prelim test: 111
Number of HSP's that attempted gapping in prelim test: 2726
Number of HSP's gapped (non-prelim): 1607
length of query: 271
length of database: 59,974,054
effective HSP length: 108
effective length of query: 163
effective length of database: 42,240,346
effective search space: 6885176398
effective search space used: 6885176398
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0062.9


