

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.14
(267 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
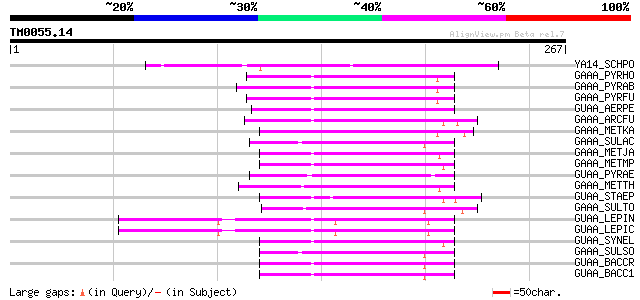
Score E
Sequences producing significant alignments: (bits) Value
YA14_SCHPO (Q09686) Hypothetical protein C13C5.04 in chromosome I 84 4e-16
GAAA_PYRHO (O59071) GMP synthase [glutamine-hydrolyzing] subunit... 76 1e-13
GAAA_PYRAB (Q9V0I6) GMP synthase [glutamine-hydrolyzing] subunit... 74 4e-13
GAAA_PYRFU (Q8U0R9) GMP synthase [glutamine-hydrolyzing] subunit... 69 9e-12
GUAA_AERPE (Q9Y933) GMP synthase [glutamine-hydrolyzing] (EC 6.3... 66 1e-10
GAAA_ARCFU (O28949) GMP synthase [glutamine-hydrolyzing] subunit... 64 3e-10
GAAA_METKA (Q8TXV8) GMP synthase [glutamine-hydrolyzing] subunit... 62 1e-09
GAAA_SULAC (Q8X239) GMP synthase [glutamine-hydrolyzing] subunit... 62 1e-09
GAAA_METJA (Q58970) GMP synthase [glutamine-hydrolyzing] subunit... 61 2e-09
GAAA_METMP (Q6LXA7) GMP synthase [glutamine-hydrolyzing] subunit... 59 9e-09
GUAA_PYRAE (Q8ZT92) GMP synthase [glutamine-hydrolyzing] (EC 6.3... 57 4e-08
GAAA_METTH (O26805) GMP synthase [glutamine-hydrolyzing] subunit... 57 4e-08
GUAA_STAEP (Q8CMQ8) GMP synthase [glutamine-hydrolyzing] (EC 6.3... 56 8e-08
GAAA_SULTO (Q974T4) GMP synthase [glutamine-hydrolyzing] subunit... 55 1e-07
GUAA_LEPIN (Q8F4F4) Probable GMP synthase [glutamine-hydrolyzing... 55 2e-07
GUAA_LEPIC (Q72RB7) Probable GMP synthase [glutamine-hydrolyzing... 55 2e-07
GUAA_SYNEL (Q8DGA5) GMP synthase [glutamine-hydrolyzing] (EC 6.3... 54 4e-07
GAAA_SULSO (Q97VZ9) GMP synthase [glutamine-hydrolyzing] subunit... 54 4e-07
GUAA_BACCR (Q81IS3) GMP synthase [glutamine-hydrolyzing] (EC 6.3... 53 7e-07
GUAA_BACC1 (Q73ER7) GMP synthase [glutamine-hydrolyzing] (EC 6.3... 53 7e-07
>YA14_SCHPO (Q09686) Hypothetical protein C13C5.04 in chromosome I
Length = 248
Score = 84.0 bits (206), Expect = 4e-16
Identities = 49/174 (28%), Positives = 92/174 (52%), Gaps = 8/174 (4%)
Query: 66 WDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINK----KI 121
+++YK N ++P+ ++ P + +ITGS A ++ PWI L+S V D + K KI
Sbjct: 50 YEVYKNPN-DYPQKEDFPNINAIIITGSKASATSDAPWIKKLISFVK--DVLFKYPHIKI 106
Query: 122 LGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRN 181
+G+CFGHQI+ + G + ++ GW++ T ++L + ++I + H+D +
Sbjct: 107 VGLCFGHQIVAKAAGVPIIQNPKGWEVSSTVVQLTENGEKFFG-RKVININQMHQDMAVD 165
Query: 182 LPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQE 235
+P E++G +E ++F L QGHPEF+ +++ + L + E
Sbjct: 166 VPEGFELLGSTEDCEFQIFYKPRQALTFQGHPEFSTEVVNTMVKVLRGTEVFTE 219
>GAAA_PYRHO (O59071) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 189
Score = 75.9 bits (185), Expect = 1e-13
Identities = 38/101 (37%), Positives = 57/101 (55%), Gaps = 2/101 (1%)
Query: 115 DSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKC 174
D N ILGIC GHQ++ + GGKVGR + + +++ I LP L +++
Sbjct: 69 DEFNVPILGICLGHQLIAKFFGGKVGRGEKA-EYSLVEIEIIDEDEIFKGLPKRLKVWES 127
Query: 175 HRDEVRNLPPKAEIIGWSEKTGIEMFRYGD-HMLGIQGHPE 214
H DEV+ LPPK +I+ SE IE ++ + + G+Q HPE
Sbjct: 128 HMDEVKELPPKFKILARSETCPIEAMKHEELPIYGVQFHPE 168
>GAAA_PYRAB (Q9V0I6) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 188
Score = 73.9 bits (180), Expect = 4e-13
Identities = 36/106 (33%), Positives = 59/106 (54%), Gaps = 2/106 (1%)
Query: 110 LVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSEL 169
++ D N ILGIC GHQ++ + GGKVGR + + +++ I LP +L
Sbjct: 63 ILENYDEFNVPILGICLGHQLIAKFFGGKVGRGEKA-EYSLVEIEILEEDEIFKGLPRKL 121
Query: 170 SIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD-HMLGIQGHPE 214
+++ H DEV+ LPP +++ SE IE ++ + + G+Q HPE
Sbjct: 122 RVWESHMDEVKELPPNFKVLARSETCPIEAMKHEELPIYGVQFHPE 167
>GAAA_PYRFU (Q8U0R9) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 188
Score = 69.3 bits (168), Expect = 9e-12
Identities = 32/101 (31%), Positives = 59/101 (57%), Gaps = 2/101 (1%)
Query: 115 DSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKC 174
+ N ILGIC GHQ++ + GG+VGR + + +++ + I LP ++ +++
Sbjct: 68 EEFNVPILGICLGHQLIAKFFGGEVGRGEKA-EYSLVEIEIIDENDIFKGLPRKVRVWES 126
Query: 175 HRDEVRNLPPKAEIIGWSEKTGIEMFRYGD-HMLGIQGHPE 214
H DEV+ LPPK +++ S+ +E ++ + + G+Q HPE
Sbjct: 127 HMDEVKKLPPKFKLLARSDTCPVEAMKHEELPIYGVQFHPE 167
>GUAA_AERPE (Q9Y933) GMP synthase [glutamine-hydrolyzing] (EC
6.3.5.2) (Glutamine amidotransferase) (GMP synthetase)
Length = 512
Score = 65.9 bits (159), Expect = 1e-10
Identities = 32/98 (32%), Positives = 54/98 (54%), Gaps = 1/98 (1%)
Query: 117 INKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHR 176
+ K +LGIC+GHQ+L +VLGG+VGRS + G T +++ + LPS ++ H
Sbjct: 79 LGKPVLGICYGHQLLAKVLGGEVGRSPLP-EFGPTEVEVLDYGILLNGLPSRFKVWMSHY 137
Query: 177 DEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPE 214
D V P +A+++ + + + + G+Q HPE
Sbjct: 138 DAVLRPPGEAKVLARTPGSPVAAMELWGRVFGVQWHPE 175
>GAAA_ARCFU (O28949) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 183
Score = 64.3 bits (155), Expect = 3e-10
Identities = 35/118 (29%), Positives = 64/118 (53%), Gaps = 7/118 (5%)
Query: 114 LDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFK 173
L ++ ++GIC GHQ++ +V GG+VG+ S G +++ + +P E++++
Sbjct: 63 LKELDVPMIGICLGHQLMAKVFGGEVGKGSMG-GYSEVKVRIVEDDELFEGIPREITVWA 121
Query: 174 CHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHML-GIQGHPE-----FTIDILFHFID 225
H DEV+ LP + + S+ IE R+ L G+Q HPE F +++ +FI+
Sbjct: 122 SHMDEVKKLPEGFKRLAESDICKIEAMRHEKKPLYGVQWHPEVYHSQFGVELYRNFIE 179
>GAAA_METKA (Q8TXV8) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 187
Score = 62.4 bits (150), Expect = 1e-09
Identities = 35/109 (32%), Positives = 58/109 (53%), Gaps = 6/109 (5%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
ILGIC GHQ++ V GGKV ++ + T +++ I LP ++ + HRDEV+
Sbjct: 72 ILGICLGHQLMAEVFGGKVDWAAGREEYARTEVEILDHEGIFEGLPDKIVAWASHRDEVK 131
Query: 181 NLPPKAEIIGWSEKTGIEMFRYGD-HMLGIQGHPEFTI-----DILFHF 223
+P + + S++ +E R+ + + G+Q HPE DIL +F
Sbjct: 132 EVPDEFVVTARSDRCEVEAMRHEELPLYGVQFHPELKFTEYGPDILKNF 180
>GAAA_SULAC (Q8X239) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase) (Fragment)
Length = 168
Score = 62.0 bits (149), Expect = 1e-09
Identities = 34/100 (34%), Positives = 54/100 (54%), Gaps = 2/100 (2%)
Query: 116 SINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCH 175
S+N LGIC GHQ++ +VLGG+V R + + G+TT+ + +I L + ++ H
Sbjct: 50 SLNVPKLGICLGHQLIAKVLGGEV-RKANKPEYGLTTVNIVDEDTILRGLKPSIKAWESH 108
Query: 176 RDEVRNLPPKAEIIGWSEKTGIE-MFRYGDHMLGIQGHPE 214
DEV P I+ SE ++ M + + G+Q HPE
Sbjct: 109 NDEVVRPPSGFRILASSENAKVQAMVNNDNSIFGVQFHPE 148
>GAAA_METJA (Q58970) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 188
Score = 61.2 bits (147), Expect = 2e-09
Identities = 31/95 (32%), Positives = 54/95 (56%), Gaps = 2/95 (2%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
ILGIC GHQ++ GG+VGR+ + +T + + + + N+P E + + H+DEV+
Sbjct: 72 ILGICLGHQLIALAYGGEVGRAEAE-EYALTKVYVDKENDLFKNVPREFNAWASHKDEVK 130
Query: 181 NLPPKAEIIGWSEKTGIEMFRYGDH-MLGIQGHPE 214
+P EI+ S+ +E ++ + G+Q HPE
Sbjct: 131 KVPEGFEILAHSDICQVEAMKHKTKPIYGVQFHPE 165
>GAAA_METMP (Q6LXA7) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 189
Score = 59.3 bits (142), Expect = 9e-09
Identities = 28/95 (29%), Positives = 54/95 (56%), Gaps = 2/95 (2%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
+LGIC GHQ++ + GG+V R+ + + + + + + +PSE + + H DEV+
Sbjct: 72 VLGICLGHQLISKAYGGEVSRADSE-EYASIKIYVKQENDLFKGVPSEFTAWASHMDEVK 130
Query: 181 NLPPKAEIIGWSEKTGIEMFRYGDHML-GIQGHPE 214
P E++ +S+ G+E ++ + + G+Q HPE
Sbjct: 131 VTPDCFEVLAYSDICGVESIKHKEKSIYGVQFHPE 165
>GUAA_PYRAE (Q8ZT92) GMP synthase [glutamine-hydrolyzing] (EC
6.3.5.2) (Glutamine amidotransferase) (GMP synthetase)
Length = 505
Score = 57.4 bits (137), Expect = 4e-08
Identities = 33/99 (33%), Positives = 53/99 (53%), Gaps = 3/99 (3%)
Query: 116 SINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCH 175
+++K +LGIC+GHQ++ + LGGKV R + G T +K+ + + E +++ H
Sbjct: 71 ALSKPVLGICYGHQMIAKKLGGKVERGK--GEYGKTIVKILVNDPLFDGWKPEEAVWMSH 128
Query: 176 RDEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPE 214
D V PP ++ SE I R G + G+Q HPE
Sbjct: 129 SDFVEEPPPGFHVLAISENGYIAAMRKG-LIYGVQFHPE 166
>GAAA_METTH (O26805) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 186
Score = 57.4 bits (137), Expect = 4e-08
Identities = 30/105 (28%), Positives = 54/105 (50%), Gaps = 2/105 (1%)
Query: 111 VHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELS 170
V + + ILGIC GHQ++ + GG+V ++ +++ I L +S
Sbjct: 63 VEYIQKLEIPILGICLGHQLIAKAYGGEVA-TAEAESYAQIELEILDEDDIFRGLGPRMS 121
Query: 171 IFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDH-MLGIQGHPE 214
++ H+DEV+ LP + +++ S +E ++ D + GIQ HPE
Sbjct: 122 VWASHKDEVKKLPDEFDVLATSSICEVEAMKHHDKPVYGIQFHPE 166
>GUAA_STAEP (Q8CMQ8) GMP synthase [glutamine-hydrolyzing] (EC
6.3.5.2) (Glutamine amidotransferase) (GMP synthetase)
Length = 513
Score = 56.2 bits (134), Expect = 8e-08
Identities = 37/113 (32%), Positives = 60/113 (52%), Gaps = 8/113 (7%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
+LGIC+G Q+ ++LGGKV R++ + G T+ A S + LPSE +++ H D+V
Sbjct: 82 VLGICYGMQLTTKLLGGKVERANER-EYGKATIN-AKSDELFFGLPSEQTVWMSHSDKVI 139
Query: 181 NLPPKAEIIGWSEKTGIEMFRYGDHML-GIQGHP-----EFTIDILFHFIDRL 227
+P E+I S T + G+Q HP E+ D+L +F+ R+
Sbjct: 140 EIPEGFEVIADSPSTNYAAIEDKKRRIYGVQFHPEVRHTEYGNDLLRNFVRRV 192
>GAAA_SULTO (Q974T4) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 188
Score = 55.5 bits (132), Expect = 1e-07
Identities = 34/110 (30%), Positives = 59/110 (52%), Gaps = 7/110 (6%)
Query: 122 LGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRN 181
LGIC GHQ+L +VLGG+V + +T + G+ + + +I L ++ ++ H DEV +
Sbjct: 76 LGICLGHQLLAKVLGGEVTK-ATKPEYGLVKVNINDEDTILRGLSPSINAWESHTDEVIS 134
Query: 182 LPPKAEIIGWSEKTGIE-MFRYGDHMLGIQGHPEFT-----IDILFHFID 225
P I+ SE ++ M + + G+Q HPE I++ +FI+
Sbjct: 135 PPQGFRILANSENAKVQAMVNKDNTIFGVQFHPEVKHTEKGIEVFKNFIE 184
>GUAA_LEPIN (Q8F4F4) Probable GMP synthase [glutamine-hydrolyzing]
(EC 6.3.5.2) (Glutamine amidotransferase) (GMP
synthetase)
Length = 603
Score = 54.7 bits (130), Expect = 2e-07
Identities = 44/166 (26%), Positives = 73/166 (43%), Gaps = 11/166 (6%)
Query: 53 GFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHA-NHPWIHDLLSLV 111
G + ++A R Y + Y G +++G + N P ++
Sbjct: 14 GQYAHLIASRIRRLGAYTEILSNEEPISNYQKYSGIILSGGPESVYEPNSP------TVT 67
Query: 112 HTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKL--ASSSSIALNLPSEL 169
+ + ILGIC+GHQ++ ++LGG V RS TG + G +++L + +SI N
Sbjct: 68 TRIFDLGIPILGICYGHQLIMKLLGGVVERSGTG-EYGPASLQLHGTNGNSILKNFVGGE 126
Query: 170 SIFKCHRDEVRNLPPKAEIIGWSEKTGIEMF-RYGDHMLGIQGHPE 214
++ H DEV LP I +S+ G + + GIQ H E
Sbjct: 127 QVWMNHADEVVKLPEGFSKIAFSKDCGYAVVANSSKKIFGIQFHAE 172
>GUAA_LEPIC (Q72RB7) Probable GMP synthase [glutamine-hydrolyzing]
(EC 6.3.5.2) (Glutamine amidotransferase) (GMP
synthetase)
Length = 603
Score = 54.7 bits (130), Expect = 2e-07
Identities = 44/166 (26%), Positives = 73/166 (43%), Gaps = 11/166 (6%)
Query: 53 GFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHA-NHPWIHDLLSLV 111
G + ++A R Y + Y G +++G + N P ++
Sbjct: 14 GQYAHLIASRIRRLGAYTEILSNEEPISNYQKYSGIILSGGPESVYEPNSP------TVT 67
Query: 112 HTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKL--ASSSSIALNLPSEL 169
+ + ILGIC+GHQ++ ++LGG V RS TG + G +++L + +SI N
Sbjct: 68 TRIFDLGIPILGICYGHQLIMKLLGGVVERSGTG-EYGPASLQLHGTNGNSILKNFVGGE 126
Query: 170 SIFKCHRDEVRNLPPKAEIIGWSEKTGIEMF-RYGDHMLGIQGHPE 214
++ H DEV LP I +S+ G + + GIQ H E
Sbjct: 127 QVWMNHADEVVKLPEGFSKIAFSKDCGYAVVANSSKKIFGIQFHAE 172
>GUAA_SYNEL (Q8DGA5) GMP synthase [glutamine-hydrolyzing] (EC
6.3.5.2) (Glutamine amidotransferase) (GMP synthetase)
Length = 535
Score = 53.9 bits (128), Expect = 4e-07
Identities = 29/95 (30%), Positives = 50/95 (52%), Gaps = 2/95 (2%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
ILG+C+G Q++ + LGG V R+ G + G + + + + N+ E ++ H D V
Sbjct: 94 ILGVCYGMQLMVQQLGGTVERAIAG-EYGKAALFIDDPTDLLTNVEQETIMWMSHGDSVT 152
Query: 181 NLPPKAEIIGWSEKTGIEMFRYGDHML-GIQGHPE 214
LPP ++ ++ T I + + L G+Q HPE
Sbjct: 153 ELPPGFRVLAHTDNTPIAAIAHPERKLYGVQFHPE 187
>GAAA_SULSO (Q97VZ9) GMP synthase [glutamine-hydrolyzing] subunit A
(EC 6.3.5.2) (Glutamine amidotransferase)
Length = 188
Score = 53.9 bits (128), Expect = 4e-07
Identities = 29/95 (30%), Positives = 50/95 (52%), Gaps = 2/95 (2%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
+LGIC GHQ++ VLGG V R + + G+T + + +I +L++++ H DEV
Sbjct: 75 MLGICLGHQLIAYVLGGVV-RRALNPEYGLTRINIFDEDTILKGFSQQLNVWESHNDEVV 133
Query: 181 NLPPKAEIIGWSEKTGIE-MFRYGDHMLGIQGHPE 214
P ++ S ++ M + + G+Q HPE
Sbjct: 134 EPPSGFRVLASSANARVQAMANSSNSIFGVQFHPE 168
>GUAA_BACCR (Q81IS3) GMP synthase [glutamine-hydrolyzing] (EC
6.3.5.2) (Glutamine amidotransferase) (GMP synthetase)
Length = 515
Score = 53.1 bits (126), Expect = 7e-07
Identities = 32/95 (33%), Positives = 49/95 (50%), Gaps = 2/95 (2%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
I GIC+G Q++ + GGKV R++ + G +K+ + S + NLP E ++ H D V
Sbjct: 83 IFGICYGMQLMTQQFGGKVERANHR-EYGKAVLKVENESKLYANLPEEQVVWMSHGDLVT 141
Query: 181 NLPPKAEIIGWSEKTGIE-MFRYGDHMLGIQGHPE 214
LP + SE I M ++ G+Q HPE
Sbjct: 142 GLPEGFVVDATSESCPIAGMSNEAKNLYGVQFHPE 176
>GUAA_BACC1 (Q73ER7) GMP synthase [glutamine-hydrolyzing] (EC
6.3.5.2) (Glutamine amidotransferase) (GMP synthetase)
Length = 512
Score = 53.1 bits (126), Expect = 7e-07
Identities = 32/95 (33%), Positives = 49/95 (50%), Gaps = 2/95 (2%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVR 180
I GIC+G Q++ + GGKV R++ + G +K+ + S + NLP E ++ H D V
Sbjct: 80 IFGICYGMQLMTQKFGGKVERANHR-EYGKAVLKVENESKLYANLPEEQVVWMSHGDLVT 138
Query: 181 NLPPKAEIIGWSEKTGIE-MFRYGDHMLGIQGHPE 214
LP + SE I M ++ G+Q HPE
Sbjct: 139 GLPEGFVVDATSESCPIAGMSNEAKNLYGVQFHPE 173
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.143 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,849,433
Number of Sequences: 164201
Number of extensions: 1602466
Number of successful extensions: 15725
Number of sequences better than 10.0: 334
Number of HSP's better than 10.0 without gapping: 192
Number of HSP's successfully gapped in prelim test: 144
Number of HSP's that attempted gapping in prelim test: 13498
Number of HSP's gapped (non-prelim): 1288
length of query: 267
length of database: 59,974,054
effective HSP length: 108
effective length of query: 159
effective length of database: 42,240,346
effective search space: 6716215014
effective search space used: 6716215014
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0055.14


