

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.11
(605 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
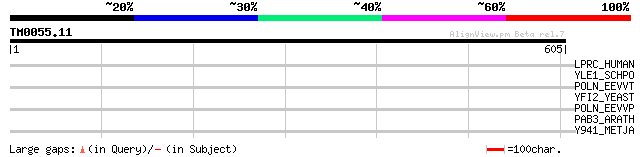
Score E
Sequences producing significant alignments: (bits) Value
LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP13... 40 0.013
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 37 0.15
POLN_EEVVT (P27282) Nonstructural polyprotein [Contains: Nonstru... 33 1.6
YFI2_YEAST (P43595) Hypothetical 22.4 kDa protein in GCN20-CMK1 ... 33 2.2
POLN_EEVVP (P36328) Nonstructural polyprotein [Contains: Nonstru... 32 3.7
PAB3_ARATH (O64380) Polyadenylate-binding protein 3 (Poly(A)-bin... 32 3.7
Y941_METJA (Q57711) Hypothetical protein MJ0941 32 6.3
>LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP130)
(Leucine-rich PPR-motif containing protein)
Length = 1273
Score = 40.4 bits (93), Expect = 0.013
Identities = 45/211 (21%), Positives = 97/211 (45%), Gaps = 17/211 (8%)
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLH 378
DV +++++ Y Q + KM +PN VT +I++ N+ ++ +
Sbjct: 40 DVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIASYCNVGDIEGASKIL 99
Query: 379 GFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNR----DSISWSTMISAYGLH 434
GF+ L + ++ +AL+ +A+ G +E + I M++ ++ +++AY
Sbjct: 100 GFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIEPGPDTYLALLNAYAEK 159
Query: 435 G---HGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYE-IPL 490
G H +Q L+ + + + D L ++ + + AG ++ Q+ +K+ + IP
Sbjct: 160 GDIDHVKQTLEKVEKFELHLMDRD---LLQIIFSFSKAGYLSMSQKFWKKFTCERRYIPD 216
Query: 491 TIEHYACLVDLLGRSGKVED-ALEIMRTMPM 520
+ LV + K+ED AL+I+ P+
Sbjct: 217 AMNLILLLV-----TEKLEDVALQILLACPV 242
Score = 34.3 bits (77), Expect = 0.97
Identities = 25/99 (25%), Positives = 47/99 (47%), Gaps = 4/99 (4%)
Query: 180 DGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKN----EVSWTAMISGCIANLDYD 235
+ ++ + V L+ Y D AS++ M+ K+ E ++A+++G D +
Sbjct: 69 EANIQPNRVTYQRLIASYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDME 128
Query: 236 AAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKE 274
A M+ G++P T +ALL A A+ G + H K+
Sbjct: 129 NAENILTVMRDAGIEPGPDTYLALLNAYAEKGDIDHVKQ 167
Score = 33.9 bits (76), Expect = 1.3
Identities = 23/75 (30%), Positives = 33/75 (43%), Gaps = 5/75 (6%)
Query: 474 EGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRI-----WS 528
E Q ++ A YE + YA L++L R KVEDAL + S+ + +
Sbjct: 572 ENMQKALELKAKYESDMVTGGYAALINLCCRHDKVEDALNLKEEFDRLDSSAVLDTGNYL 631
Query: 529 SLVSACKLHGRLDIA 543
LV HG+L A
Sbjct: 632 GLVRVLAKHGKLQDA 646
Score = 31.2 bits (69), Expect = 8.2
Identities = 22/98 (22%), Positives = 48/98 (48%), Gaps = 9/98 (9%)
Query: 244 MQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGE 303
M+ + PNRVT L+ + G ++ +I G+ + + SAL+ + ++G+
Sbjct: 67 MEEANIQPNRVTYQRLIASYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGD 126
Query: 304 SLHLAEQIFERSSFRDVVL------WSSIIGSYSQRGD 335
+ AE I + RD + + +++ +Y+++GD
Sbjct: 127 -MENAENIL--TVMRDAGIEPGPDTYLALLNAYAEKGD 161
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 37.0 bits (84), Expect = 0.15
Identities = 23/84 (27%), Positives = 40/84 (47%), Gaps = 1/84 (1%)
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
AL +F E K VKP + AVLS A TE ++F+++ +P ++ Y ++
Sbjct: 910 ALNIFEETKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVT-YGTVI 968
Query: 500 DLLGRSGKVEDALEIMRTMPMKPS 523
+ R G A ++ M +P+
Sbjct: 969 NAACRIGDESLAEKLFAEMENQPN 992
Score = 32.7 bits (73), Expect = 2.8
Identities = 16/59 (27%), Positives = 33/59 (55%)
Query: 424 WSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV 482
++ ++S G + KLF EMKE G+ P ++T+ V++A G + +++F ++
Sbjct: 929 YNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEM 987
Score = 32.3 bits (72), Expect = 3.7
Identities = 37/191 (19%), Positives = 82/191 (42%), Gaps = 10/191 (5%)
Query: 323 WSSIIGSYSQRGDS---YKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
++ +I + ++RGD+ AL +F + +P+ AV+S L
Sbjct: 891 FAHLINNSTRRGDTDDATTALNIFEETKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQ 950
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS-----WSTMIS-AYGL 433
+ + GL + ++N + G + ++F EM+N+ + ++TMI
Sbjct: 951 EMKESGLLPTSVTYGTVINAACRIGDESLAEKLFAEMENQPNYQPRVAPYNTMIQFEVQT 1010
Query: 434 HGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQ-QVFKQVNADYEIPLTI 492
+ E+AL ++ + ++P + T+ ++ A V G + ++ ++P+
Sbjct: 1011 MFNREKALFYYNRLCATDIEPSSHTYKLLMDAYGTLKPVNVGSVKAVLELMERTDVPILS 1070
Query: 493 EHYACLVDLLG 503
HYA + +LG
Sbjct: 1071 MHYAAYIHILG 1081
>POLN_EEVVT (P27282) Nonstructural polyprotein [Contains:
Nonstructural protein NSP1; Nonstructural protein NSP2;
Nonstructural protein NSP3; Nonstructural protein NSP4]
Length = 2492
Score = 33.5 bits (75), Expect = 1.6
Identities = 28/94 (29%), Positives = 44/94 (46%), Gaps = 3/94 (3%)
Query: 24 SLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACS-SLHSHTLGTLLHGLALTTGSH 82
SL+S G HQ L++ +H P S+SS S+ A + S ++ L G ++T+G+
Sbjct: 1712 SLLSDGPTHQVLQVEADIH--GPPSVSSSSWSIPHASDFDVDSLSILDTLEGASVTSGAT 1769
Query: 83 SDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDP 116
S S S+ + + R VF PH P
Sbjct: 1770 SAETNSYFAKSMEFLARPVPAPRTVFRNPPHPAP 1803
>YFI2_YEAST (P43595) Hypothetical 22.4 kDa protein in GCN20-CMK1
intergenic region precursor
Length = 202
Score = 33.1 bits (74), Expect = 2.2
Identities = 21/88 (23%), Positives = 42/88 (46%), Gaps = 1/88 (1%)
Query: 45 APHSISSVLPSV-IKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQS 103
A H+I +L S+ + C + S L ++ L T G++ + SN+++ + S I +
Sbjct: 87 ARHTIGGLLISIPVATCLTFISFALPLVIIFLFQTGGTNVSLITSNAILHILTLLSTIFA 146
Query: 104 ARQVFDTMPHRDPISWNSMINAYLQNGC 131
+ HRDP++ +S+ + C
Sbjct: 147 CTVILLLCMHRDPVTISSLYDLVWLANC 174
>POLN_EEVVP (P36328) Nonstructural polyprotein [Contains:
Nonstructural protein NSP1; Nonstructural protein NSP2;
Nonstructural protein NSP3; Nonstructural protein NSP4]
Length = 2492
Score = 32.3 bits (72), Expect = 3.7
Identities = 28/94 (29%), Positives = 45/94 (47%), Gaps = 3/94 (3%)
Query: 24 SLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACS-SLHSHTLGTLLHGLALTTGSH 82
SL+S G HQ L++ +H S S+SS S+ A + S ++ L G ++T+G+
Sbjct: 1712 SLLSDGPTHQVLQVEADIHGSP--SVSSSSWSIPHASDFDVDSLSILDTLDGASVTSGAV 1769
Query: 83 SDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDP 116
S S S+ + + + R VF PH P
Sbjct: 1770 SAETNSYFARSMEFRARPVPAPRTVFRNPPHPAP 1803
>PAB3_ARATH (O64380) Polyadenylate-binding protein 3
(Poly(A)-binding protein 3) (PABP 3)
Length = 660
Score = 32.3 bits (72), Expect = 3.7
Identities = 27/78 (34%), Positives = 38/78 (48%), Gaps = 7/78 (8%)
Query: 519 PMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYT--LLNMIYAEHGHWLH 576
P+ P +++ SSL SA + E L P + R EP + A T LL M AE H +
Sbjct: 568 PLLPISKLTSSLASASPADRTRMLGEQLYPLVERHEPLHVAKVTGMLLEMDQAEILHLME 627
Query: 577 -----KEQVMEAMKLQRL 589
K +V EA+ + RL
Sbjct: 628 SPEALKSKVSEALDVLRL 645
>Y941_METJA (Q57711) Hypothetical protein MJ0941
Length = 338
Score = 31.6 bits (70), Expect = 6.3
Identities = 26/111 (23%), Positives = 55/111 (49%), Gaps = 6/111 (5%)
Query: 414 LEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSAC-NHAGLV 472
L ++++ I+W + YG+ G+ ++ALK ++ K G++ L+ + + C G
Sbjct: 90 LSYESKNPITWVFVGQLYGMSGNCDEALKCYN--KALGIENRFLSAFLLKTICLEFLGEY 147
Query: 473 TEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIM-RTMPMKP 522
E + + +V P + + ++L + G+ EDAL + R + +KP
Sbjct: 148 DELLKCYNEVLT--YTPNFVPMWVKKAEILRKLGRYEDALLCLNRALELKP 196
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,774,878
Number of Sequences: 164201
Number of extensions: 2683862
Number of successful extensions: 6306
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 6284
Number of HSP's gapped (non-prelim): 24
length of query: 605
length of database: 59,974,054
effective HSP length: 116
effective length of query: 489
effective length of database: 40,926,738
effective search space: 20013174882
effective search space used: 20013174882
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0055.11


