

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.5
(638 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
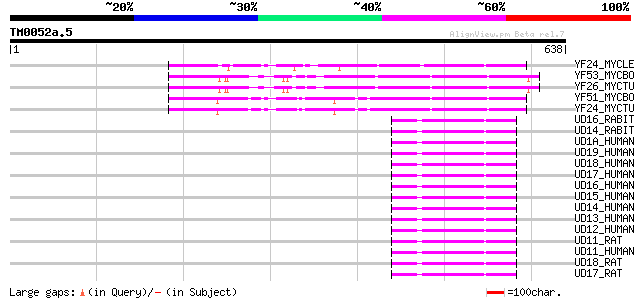
Sequences producing significant alignments: (bits) Value
YF24_MYCLE (Q49929) Hypothetical glycosyl transferase ML2348 (EC... 144 9e-34
YF53_MYCBO (P64870) Hypothetical protein Mb1553c 132 3e-30
YF26_MYCTU (P64869) Hypothetical protein Rv1526c/MT1577 132 3e-30
YF51_MYCBO (P64866) Hypothetical glycosyl transferase Mb1551 (EC... 130 1e-29
YF24_MYCTU (P64865) Hypothetical glycosyl transferase Rv1524/MT1... 130 1e-29
UD16_RABIT (Q28611) UDP-glucuronosyltransferase 1-6 precursor, m... 49 4e-05
UD14_RABIT (Q28612) UDP-glucuronosyltransferase 1-4 precursor, m... 49 4e-05
UD1A_HUMAN (Q9HAW8) UDP-glucuronosyltransferase 1-10 precursor, ... 46 3e-04
UD19_HUMAN (O60656) UDP-glucuronosyltransferase 1-9 precursor, m... 46 3e-04
UD18_HUMAN (Q9HAW9) UDP-glucuronosyltransferase 1-8 precursor, m... 46 3e-04
UD17_HUMAN (Q9HAW7) UDP-glucuronosyltransferase 1-7 precursor, m... 46 3e-04
UD16_HUMAN (P19224) UDP-glucuronosyltransferase 1-6 precursor, m... 46 3e-04
UD15_HUMAN (P35504) UDP-glucuronosyltransferase 1-5 precursor, m... 46 3e-04
UD14_HUMAN (P22310) UDP-glucuronosyltransferase 1-4 precursor, m... 46 3e-04
UD13_HUMAN (P35503) UDP-glucuronosyltransferase 1-3 precursor, m... 46 3e-04
UD12_HUMAN (P36509) UDP-glucuronosyltransferase 1-2 precursor, m... 46 3e-04
UD11_RAT (Q64550) UDP-glucuronosyltransferase 1-1 precursor, mic... 46 3e-04
UD11_HUMAN (P22309) UDP-glucuronosyltransferase 1-1 precursor, m... 46 3e-04
UD18_RAT (Q64634) UDP-glucuronosyltransferase 1-8 precursor, mic... 46 3e-04
UD17_RAT (Q64633) UDP-glucuronosyltransferase 1-7 precursor, mic... 46 3e-04
>YF24_MYCLE (Q49929) Hypothetical glycosyl transferase ML2348 (EC
2.-.-.-)
Length = 421
Score = 144 bits (362), Expect = 9e-34
Identities = 128/433 (29%), Positives = 201/433 (45%), Gaps = 51/433 (11%)
Query: 183 LQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRIL 242
++ + G+RGDV+PF A+ LQ GH VR+ + FV+SAG+ G D +
Sbjct: 1 MKFTLAASGSRGDVEPFAALGLELQRRGHEVRIGVPPDMLRFVESAGLAAVAYGPDTQ-- 58
Query: 243 AGYMARNK--------GLIPSGPGEVSIQLKQL-KDIIDSLLPACTAPDLE-TGVPFRAQ 292
++AR+ ++P P + QL+Q D+ L DL TGV + Q
Sbjct: 59 -EFLARDTYSQWRQWWKILP--PIKALQQLRQAWADMATDLKSLADGADLVMTGVVY--Q 113
Query: 293 AIIANPPAYGHAHVAEALGVPLHIFFTMPWTPTY----DFPHPLARVPQSAGYWMSYIIV 348
++AN VAE G+P + +P P PL R A W ++
Sbjct: 114 GVVAN--------VAEYYGIPFGVLHFVPARVNGKIIPSLPSPLNRAIL-ATVWRAH--- 161
Query: 349 DLMTWWGMRGIINDFRKKKLKLPPIAYFS----MYRGSISHLPTGYMWSPHVVPKPSDWG 404
W + D ++++L LP S + RG++ + P + + +++G
Sbjct: 162 -----WLLAKKPEDAQRRELGLPKATSLSTRRIVKRGALEIQAYDELCFPGLAAEWAEYG 216
Query: 405 PLVDVVGYCFLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTTDVILEALKDT 464
VG L L + + + WI G PP+YFGFGSMP+ PT T +I A D
Sbjct: 217 DRRPFVGALTLELPTAAD--NEVLSWIAAGTPPIYFGFGSMPIVSPTDTVAMIAAACADL 274
Query: 465 EQRGIIDRGWGNLGSLAEVP--DNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGC 522
+R +I L +VP D+V ++ H +FP C AVVHHGGAGTTA L+AG
Sbjct: 275 GERALISV---KPKDLTQVPKFDHVKIVTSVSHAAVFPACRAVVHHGGAGTTAASLRAGV 331
Query: 523 PTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVMLIA-K 581
PT I+ F +Q W +I ++G A S +L+ ++ +L P+ +R +A +
Sbjct: 332 PTLILWIFIEQPVWAAQIKRLKVG-AGRRFSATTQRSLAADLRTILAPQYATRAREVANR 390
Query: 582 LVENEDGVAAAVD 594
+ + ++ V AA D
Sbjct: 391 MSKPDESVNAAAD 403
>YF53_MYCBO (P64870) Hypothetical protein Mb1553c
Length = 426
Score = 132 bits (332), Expect = 3e-30
Identities = 129/459 (28%), Positives = 193/459 (41%), Gaps = 67/459 (14%)
Query: 183 LQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDP--- 239
++ + V GTRGDV+P A+ L+ GH V +A N FV+SAG+ G D
Sbjct: 1 MKFVLAVHGTRGDVEPCAAVGVELRRRGHAVHMAVPPNLIEFVESAGLTGVAYGPDSDEQ 60
Query: 240 -RILAGYM-----ARN--------KGLIPSGPGEVSIQLKQLKDIIDSLLPACTAPDLET 285
+A ++ A+N K L G E+ L L D D ++ T
Sbjct: 61 INTVAAFVRNLTRAQNPLNLARAVKELFVEGWAEMGTTLTTLADGADLVM---------T 111
Query: 286 GVPFRAQAIIANPPAYGHAHVAEALGVP---LHIF-------FTMPWTPTYDFPHPLARV 335
G + A A+VAE +P LH F +P PT P L R
Sbjct: 112 GQTYHGVA----------ANVAEYYDIPAAALHHFPMQVNGQIAIPSIPT---PATLVRA 158
Query: 336 PQSAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPH 395
W Y V + ++++L LPP ++ R + P + P
Sbjct: 159 TMKVS-WRLYAYVSKDA--------DRAQRRELGLPPAPAPAVRRLAERGAPEIQAYDPV 209
Query: 396 VVPK-PSDWGPLVDVVGYCFLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTT 454
P ++W VG + L S+ P E+ WI G PP+YFGFGS P++ P +T
Sbjct: 210 FFPGLAAEWSDRRPFVGPLTMELHSE--PNEELESWIAAGTPPIYFGFGSTPVQTPVQTL 267
Query: 455 DVILEALKDTEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTT 514
+I + +R +I N + D+V + + + P+C AVVHHGGAGTT
Sbjct: 268 AMISDVCAQLGERALIYSPAANSTRIRHA-DHVKRVGLVNYSTILPKCRAVVHHGGAGTT 326
Query: 515 ATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKS 574
A GL+AG PT I+ DQ W + ++G A + + +L ++ +L PE +
Sbjct: 327 AAGLRAGMPTLILWDVADQPIWAGAVQRLKVGSAK-RFTNITRGSLLKELRSILAPECAA 385
Query: 575 RVMLIA-KLVENEDGVAAAVD---AFHRHLPDELPLPTP 609
R I+ ++ V AA D A R P P +P
Sbjct: 386 RAREISTRMTRPTAAVTAAADLLEATARQTPGSTPSSSP 424
>YF26_MYCTU (P64869) Hypothetical protein Rv1526c/MT1577
Length = 426
Score = 132 bits (332), Expect = 3e-30
Identities = 129/459 (28%), Positives = 193/459 (41%), Gaps = 67/459 (14%)
Query: 183 LQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDP--- 239
++ + V GTRGDV+P A+ L+ GH V +A N FV+SAG+ G D
Sbjct: 1 MKFVLAVHGTRGDVEPCAAVGVELRRRGHAVHMAVPPNLIEFVESAGLTGVAYGPDSDEQ 60
Query: 240 -RILAGYM-----ARN--------KGLIPSGPGEVSIQLKQLKDIIDSLLPACTAPDLET 285
+A ++ A+N K L G E+ L L D D ++ T
Sbjct: 61 INTVAAFVRNLTRAQNPLNLARAVKELFVEGWAEMGTTLTTLADGADLVM---------T 111
Query: 286 GVPFRAQAIIANPPAYGHAHVAEALGVP---LHIF-------FTMPWTPTYDFPHPLARV 335
G + A A+VAE +P LH F +P PT P L R
Sbjct: 112 GQTYHGVA----------ANVAEYYDIPAAALHHFPMQVNGQIAIPSIPT---PATLVRA 158
Query: 336 PQSAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPH 395
W Y V + ++++L LPP ++ R + P + P
Sbjct: 159 TMKVS-WRLYAYVSKDA--------DRAQRRELGLPPAPAPAVRRLAERGAPEIQAYDPV 209
Query: 396 VVPK-PSDWGPLVDVVGYCFLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTT 454
P ++W VG + L S+ P E+ WI G PP+YFGFGS P++ P +T
Sbjct: 210 FFPGLAAEWSDRRPFVGPLTMELHSE--PNEELESWIAAGTPPIYFGFGSTPVQTPVQTL 267
Query: 455 DVILEALKDTEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTT 514
+I + +R +I N + D+V + + + P+C AVVHHGGAGTT
Sbjct: 268 AMISDVCAQLGERALIYSPAANSTRIRHA-DHVKRVGLVNYSTILPKCRAVVHHGGAGTT 326
Query: 515 ATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKS 574
A GL+AG PT I+ DQ W + ++G A + + +L ++ +L PE +
Sbjct: 327 AAGLRAGMPTLILWDVADQPIWAGAVQRLKVGSAK-RFTNITRGSLLKELRSILAPECAA 385
Query: 575 RVMLIA-KLVENEDGVAAAVD---AFHRHLPDELPLPTP 609
R I+ ++ V AA D A R P P +P
Sbjct: 386 RAREISTRMTRPTAAVTAAADLLEATARQTPGSTPSSSP 424
>YF51_MYCBO (P64866) Hypothetical glycosyl transferase Mb1551 (EC
2.-.-.-)
Length = 414
Score = 130 bits (326), Expect = 1e-29
Identities = 123/426 (28%), Positives = 189/426 (43%), Gaps = 36/426 (8%)
Query: 183 LQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGG----- 237
++ + GTRGD++P A+ LQ GH V LA N FV++AG+ G
Sbjct: 1 MKFVVASYGTRGDIEPCAAVGLELQRRGHDVCLAVPPNLIGFVETAGLSAVAYGSRDSQE 60
Query: 238 --DPRIL--AGYMARNKGLIPSGPGEVSIQLKQLKDIIDSLLPACTAPDLETGVPFRAQA 293
D + L A + L+ V+ +L ++ + A A L TG + Q
Sbjct: 61 QLDEQFLHNAWKLQNPIKLLREAMAPVTEGWAELSAMLTPV--AAGADLLLTGQIY--QE 116
Query: 294 IIANPPAYGHAHVAEALGVPLHIFFTMPWTPTYDFPHPLARVPQSAGYWMSYIIVDLMTW 353
++AN VAE G+PL P + P AR+P A S I +
Sbjct: 117 VVAN--------VAEHHGIPLAALHFYPVRANGEIAFP-ARLP--APLVRSTITAIDWLY 165
Query: 354 WGMRGIINDFRKKKLKLP----PIAYFSMYRGSISHLPTGYMWSPHVVPKPSDWGPLVDV 409
W M + D ++++L LP P RGS+ + P + ++WG
Sbjct: 166 WRMTKGVEDAQRRELGLPKASTPAPRRMAVRGSLEIQAYDALCFPGLA---AEWGGRRPF 222
Query: 410 VGYCFLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTTDVILEALKDTEQRGI 469
VG L++ S ++ WI PP+YFGFGSMP+ +I A + +R +
Sbjct: 223 VG--ALTMESATDADDEVASWIAADTPPIYFGFGSMPIGSLADRVAMISAACAELGERAL 280
Query: 470 IDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPF 529
I G + + + D+V ++ H +FP C AVVHHGGAGTTA GL+AG PT I+
Sbjct: 281 ICSGPSDATGIPQF-DHVKVVRVVSHAAVFPTCRAVVHHGGAGTTAAGLRAGIPTLILWV 339
Query: 530 FGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVMLIA-KLVENEDG 588
DQ W +I + ++G S E+L ++ +L P+ +R IA ++ +
Sbjct: 340 TSDQPIWAAQIKQLKVGRGR-RFSSATKESLIADLRTILAPDYVTRAREIASRMTKPAAS 398
Query: 589 VAAAVD 594
V A D
Sbjct: 399 VTATAD 404
>YF24_MYCTU (P64865) Hypothetical glycosyl transferase Rv1524/MT1575
(EC 2.-.-.-)
Length = 414
Score = 130 bits (326), Expect = 1e-29
Identities = 123/426 (28%), Positives = 189/426 (43%), Gaps = 36/426 (8%)
Query: 183 LQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGG----- 237
++ + GTRGD++P A+ LQ GH V LA N FV++AG+ G
Sbjct: 1 MKFVVASYGTRGDIEPCAAVGLELQRRGHDVCLAVPPNLIGFVETAGLSAVAYGSRDSQE 60
Query: 238 --DPRIL--AGYMARNKGLIPSGPGEVSIQLKQLKDIIDSLLPACTAPDLETGVPFRAQA 293
D + L A + L+ V+ +L ++ + A A L TG + Q
Sbjct: 61 QLDEQFLHNAWKLQNPIKLLREAMAPVTEGWAELSAMLTPV--AAGADLLLTGQIY--QE 116
Query: 294 IIANPPAYGHAHVAEALGVPLHIFFTMPWTPTYDFPHPLARVPQSAGYWMSYIIVDLMTW 353
++AN VAE G+PL P + P AR+P A S I +
Sbjct: 117 VVAN--------VAEHHGIPLAALHFYPVRANGEIAFP-ARLP--APLVRSTITAIDWLY 165
Query: 354 WGMRGIINDFRKKKLKLP----PIAYFSMYRGSISHLPTGYMWSPHVVPKPSDWGPLVDV 409
W M + D ++++L LP P RGS+ + P + ++WG
Sbjct: 166 WRMTKGVEDAQRRELGLPKASTPAPRRMAVRGSLEIQAYDALCFPGLA---AEWGGRRPF 222
Query: 410 VGYCFLSLASKYKPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTTDVILEALKDTEQRGI 469
VG L++ S ++ WI PP+YFGFGSMP+ +I A + +R +
Sbjct: 223 VG--ALTMESATDADDEVASWIAADTPPIYFGFGSMPIGSLADRVAMISAACAELGERAL 280
Query: 470 IDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPF 529
I G + + + D+V ++ H +FP C AVVHHGGAGTTA GL+AG PT I+
Sbjct: 281 ICSGPSDATGIPQF-DHVKVVRVVSHAAVFPTCRAVVHHGGAGTTAAGLRAGIPTLILWV 339
Query: 530 FGDQFFWGDRIHEKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVMLIA-KLVENEDG 588
DQ W +I + ++G S E+L ++ +L P+ +R IA ++ +
Sbjct: 340 TSDQPIWAAQIKQLKVGRGR-RFSSATKESLIADLRTILAPDYVTRAREIASRMTKPAAS 398
Query: 589 VAAAVD 594
V A D
Sbjct: 399 VTATAD 404
>UD16_RABIT (Q28611) UDP-glucuronosyltransferase 1-6 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*6) (UGT1-06)
(UGT1.6) (UGT1A6)
Length = 531
Score = 48.9 bits (115), Expect = 4e-05
Identities = 39/147 (26%), Positives = 69/147 (46%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W GS + + N +L++ P + L
Sbjct: 304 FSLGSMVSEIPEKKAMEIADALGKIPQTVL----WRYTGSRPSNLAKNTYLVKWLPQNVL 359
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P+ A + H G+ G+ G P ++P FGDQ RI + G + + ++
Sbjct: 360 LGHPKTRAFITHSGSHGIYEGICNGVPMVMLPLFGDQMDNAKRIETRGAG-VTLNVLEMT 418
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
++L+NA+K ++ + K +M ++ L
Sbjct: 419 SDDLANALKTVINDKSYKENIMRLSSL 445
>UD14_RABIT (Q28612) UDP-glucuronosyltransferase 1-4 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*4) (UGT1-04)
(UGT1.4) (UGT1A4)
Length = 532
Score = 48.9 bits (115), Expect = 4e-05
Identities = 39/147 (26%), Positives = 69/147 (46%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W GS + + N +L++ P + L
Sbjct: 305 FSLGSMVSEIPEKKAMEIADALGKIPQTVL----WRYTGSRPSNLAKNTYLVKWLPQNVL 360
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P+ A + H G+ G+ G P ++P FGDQ RI + G + + ++
Sbjct: 361 LGHPKTRAFITHSGSHGIYEGICNGVPMVMLPLFGDQMDNAKRIETRGAG-VTLNVLEMT 419
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
++L+NA+K ++ + K +M ++ L
Sbjct: 420 SDDLANALKTVINDKSYKENIMRLSSL 446
>UD1A_HUMAN (Q9HAW8) UDP-glucuronosyltransferase 1-10 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*10) (UGT1-10)
(UGT1.10) (UGT-1J) (UGT1J)
Length = 530
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 302 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 357
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 358 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 416
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 417 SEDLENALKAVINDKSYKENIMRLSSL 443
>UD19_HUMAN (O60656) UDP-glucuronosyltransferase 1-9 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A9) (UDPGT) (UGT1*9) (UGT1-9) (UGT1.9) (UGT-1I) (UGT1I)
Length = 530
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 302 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 357
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 358 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 416
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 417 SEDLENALKAVINDKSYKENIMRLSSL 443
>UD18_HUMAN (Q9HAW9) UDP-glucuronosyltransferase 1-8 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A8) (UDPGT) (UGT1*8) (UGT1-08) (UGT1.8) (UGT-1H)
(UGT1H)
Length = 530
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 302 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 357
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 358 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 416
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 417 SEDLENALKAVINDKSYKENIMRLSSL 443
>UD17_HUMAN (Q9HAW7) UDP-glucuronosyltransferase 1-7 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A7) (UDPGT) (UGT1*7) (UGT1-07) (UGT1.7) (UGT-1G)
(UGT1G)
Length = 530
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 302 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 357
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 358 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 416
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 417 SEDLENALKAVINDKSYKENIMRLSSL 443
>UD16_HUMAN (P19224) UDP-glucuronosyltransferase 1-6 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A6) (UDPGT) (UGT1*6) (UGT1-06) (UGT1.6) (UGT-1F)
(UGT1F) (Phenol-metabolizing
UDP-glucuronosyltransferase)
Length = 532
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 304 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 359
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 360 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 418
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 419 SEDLENALKAVINDKSYKENIMRLSSL 445
>UD15_HUMAN (P35504) UDP-glucuronosyltransferase 1-5 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A5) (UDPGT) (UGT1*5) (UGT1-05) (UGT1.5) (UGT-1E)
(UGT1E)
Length = 534
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 306 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 361
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 362 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 420
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 421 SEDLENALKAVINDKSYKENIMRLSSL 447
>UD14_HUMAN (P22310) UDP-glucuronosyltransferase 1-4 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A4) (UDPGT) (UGT1*4) (UGT1-04) (UGT1.4) (UGT-1D)
(UGT1D) (Bilirubin specific UDPGT isozyme 2) (HUG-BR2)
Length = 534
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 306 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 361
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 362 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 420
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 421 SEDLENALKAVINDKSYKENIMRLSSL 447
>UD13_HUMAN (P35503) UDP-glucuronosyltransferase 1-3 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A3) (UDPGT) (UGT1*3) (UGT1-03) (UGT1.3) (UGT-1C)
(UGT1C)
Length = 534
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 306 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 361
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 362 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 420
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 421 SEDLENALKAVINDKSYKENIMRLSSL 447
>UD12_HUMAN (P36509) UDP-glucuronosyltransferase 1-2 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A2) (UDPGT) (UGT1*2) (UGT1-02) (UGT1.2) (UGT-1B)
(UGT1B) (HLUGP4)
Length = 530
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 302 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 357
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 358 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 416
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 417 SEDLENALKAVINDKSYKENIMRLSSL 443
>UD11_RAT (Q64550) UDP-glucuronosyltransferase 1-1 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*1) (UGT1-01)
(UGT1.1) (UGT1A1) (B1)
Length = 535
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I EAL Q + W G+ + + N L++ P + L
Sbjct: 307 FSLGSMVSEIPEKKAMEIAEALGRIPQTVL----WRYTGTRPSNLAKNTILVKWLPQNDL 362
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P+ A + H G+ G+ G P ++P FGDQ R+ + G + + ++
Sbjct: 363 LGHPKARAFITHSGSHGIYEGICNGVPMVMMPLFGDQMDNAKRMETRGAG-VTLNVLEMT 421
Query: 557 VENLSNAIKFML-QPEVKSRVMLIAKL 582
++L NA+K ++ K +M ++ L
Sbjct: 422 ADDLENALKTVINNKSYKENIMRLSSL 448
>UD11_HUMAN (P22309) UDP-glucuronosyltransferase 1-1 precursor,
microsomal (EC 2.4.1.17) (UDP-glucuronosyltransferase
1A1) (UDPGT) (UGT1*1) (UGT1-01) (UGT1.1) (UGT-1A)
(UGT1A) (Bilirubin specific UDPGT isozyme 1) (HUG-BR1)
Length = 533
Score = 46.2 bits (108), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I +AL Q + W G+ + + +N L++ P + L
Sbjct: 305 FSLGSMVSEIPEKKAMAIADALGKIPQTVL----WRYTGTRPSNLANNTILVKWLPQNDL 360
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P A + H G+ + G P ++P FGDQ R+ K G + + ++
Sbjct: 361 LGHPMTRAFITHAGSHGVYESICNGVPMVMMPLFGDQMDNAKRMETKGAG-VTLNVLEMT 419
Query: 557 VENLSNAIKFMLQPE-VKSRVMLIAKL 582
E+L NA+K ++ + K +M ++ L
Sbjct: 420 SEDLENALKAVINDKSYKENIMRLSSL 446
>UD18_RAT (Q64634) UDP-glucuronosyltransferase 1-8 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*8) (UGT1-08)
(UGT1.8) (UGT1A8) (A3)
Length = 530
Score = 45.8 bits (107), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I EAL Q + W G+ + + N L++ P + L
Sbjct: 302 FSLGSMVSEIPEKKAMEIAEALGRIPQTLL----WRYTGTRPSNLAKNTILVKWLPQNDL 357
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P+ A + H G+ G+ G P ++P FGDQ R+ + G + + ++
Sbjct: 358 LGHPKARAFITHSGSHGIYEGICNGVPMVMMPLFGDQMDNAKRMETRGAG-VTLNVLEMT 416
Query: 557 VENLSNAIKFML-QPEVKSRVMLIAKL 582
++L NA+K ++ K +M ++ L
Sbjct: 417 ADDLENALKTVINNKSYKENIMRLSSL 443
>UD17_RAT (Q64633) UDP-glucuronosyltransferase 1-7 precursor,
microsomal (EC 2.4.1.17) (UDPGT) (UGT1*7) (UGT1-07)
(UGT1.7) (UGT1A7) (A2)
Length = 531
Score = 45.8 bits (107), Expect = 3e-04
Identities = 38/147 (25%), Positives = 66/147 (44%), Gaps = 9/147 (6%)
Query: 440 FGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSL-AEVPDNVFLLEECPHDWL 498
F GSM E P + I EAL Q + W G+ + + N L++ P + L
Sbjct: 303 FSLGSMVSEIPEKKAMEIAEALGRIPQTLL----WRYTGTRPSNLAKNTILVKWLPQNDL 358
Query: 499 F--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLN 556
P+ A + H G+ G+ G P ++P FGDQ R+ + G + + ++
Sbjct: 359 LGHPKARAFITHSGSHGIYEGICNGVPMVMMPLFGDQMDNAKRMETRGAG-VTLNVLEMT 417
Query: 557 VENLSNAIKFML-QPEVKSRVMLIAKL 582
++L NA+K ++ K +M ++ L
Sbjct: 418 ADDLENALKTVINNKSYKENIMRLSSL 444
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 82,568,671
Number of Sequences: 164201
Number of extensions: 3815275
Number of successful extensions: 9208
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 9173
Number of HSP's gapped (non-prelim): 85
length of query: 638
length of database: 59,974,054
effective HSP length: 117
effective length of query: 521
effective length of database: 40,762,537
effective search space: 21237281777
effective search space used: 21237281777
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0052a.5


