

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
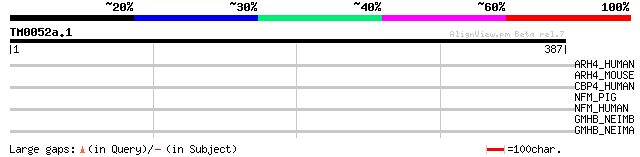
Query= TM0052a.1
(387 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ARH4_HUMAN (Q9NR80) Rho guanine nucleotide exchange factor 4 (AP... 36 0.19
ARH4_MOUSE (Q7TNR9) Rho guanine nucleotide exchange factor 4 (AP... 33 1.2
CBP4_HUMAN (Q9UI42) Carboxypeptidase A4 precursor (EC 3.4.17.-) ... 32 3.6
NFM_PIG (P08552) Neurofilament triplet M protein (160 kDa neurof... 31 6.1
NFM_HUMAN (P07197) Neurofilament triplet M protein (160 kDa neur... 31 6.1
GMHB_NEIMB (Q9JXI5) D,D-heptose 1,7-bisphosphate phosphatase (EC... 31 6.1
GMHB_NEIMA (Q9JWE9) D,D-heptose 1,7-bisphosphate phosphatase (EC... 31 6.1
>ARH4_HUMAN (Q9NR80) Rho guanine nucleotide exchange factor 4
(APC-stimulated guanine nucleotide exchange factor)
(Asef)
Length = 657
Score = 35.8 bits (81), Expect = 0.19
Identities = 25/79 (31%), Positives = 37/79 (46%), Gaps = 6/79 (7%)
Query: 170 GDSEFIST-----VQTIIYAIWEARNQVQFQQRT-FSVTGVLRRVAAMNSASQKSDGGPQ 223
GDS + T Q + A R QVQ Q T FS+T + R+ A +N++ Q+ G P+
Sbjct: 548 GDSHLLCTRKPEQKQRWLKAFAREREQVQLDQETGFSITELQRKQAMLNASKQQVTGKPK 607
Query: 224 TGSRPARWRRPQPGTIKCN 242
RP R + + N
Sbjct: 608 AVGRPCYLTRQKHPALPSN 626
>ARH4_MOUSE (Q7TNR9) Rho guanine nucleotide exchange factor 4
(APC-stimulated guanine nucleotide exchange factor)
(Asef)
Length = 484
Score = 33.1 bits (74), Expect = 1.2
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Query: 179 QTIIYAIWEARNQVQFQQRT-FSVTGVLRRVAAMNSASQKSDGGPQTGSRP 228
Q + A R QV+ Q T FS+T + R+ A +N++ Q++ G P+ RP
Sbjct: 389 QRWLKAFAREREQVRLDQETGFSITELQRKQAMLNASKQQATGKPKAVGRP 439
>CBP4_HUMAN (Q9UI42) Carboxypeptidase A4 precursor (EC 3.4.17.-)
(Carboxypeptidase A3)
Length = 421
Score = 31.6 bits (70), Expect = 3.6
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 11/80 (13%)
Query: 184 AIWEARNQVQFQQRTFSVTGVLRRVAAMNSASQKSDGGPQTGSRPARWRRPQ---PGTIK 240
AIW AR V QR ++T +L ++ DG T ++ WR+ + PG+
Sbjct: 191 AIWTARKIVSDYQRDPAITSILEKMDIFLLPVANPDGYVYTQTQNRLWRKTRSRNPGS-S 249
Query: 241 C-------NFDASFRGGGTA 253
C N++ASF G G +
Sbjct: 250 CIGADPNRNWNASFAGKGAS 269
>NFM_PIG (P08552) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M) (Fragment)
Length = 454
Score = 30.8 bits (68), Expect = 6.1
Identities = 13/41 (31%), Positives = 19/41 (45%)
Query: 206 RRVAAMNSASQKSDGGPQTGSRPARWRRPQPGTIKCNFDAS 246
RRV S+ + G P +G R W R P T+ ++ S
Sbjct: 15 RRVTETRSSFSRVSGSPSSGFRSQSWSRGSPSTVSSSYKRS 55
>NFM_HUMAN (P07197) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M) (Neurofilament 3)
Length = 915
Score = 30.8 bits (68), Expect = 6.1
Identities = 13/41 (31%), Positives = 19/41 (45%)
Query: 206 RRVAAMNSASQKSDGGPQTGSRPARWRRPQPGTIKCNFDAS 246
RRV S+ + G P +G R W R P T+ ++ S
Sbjct: 14 RRVTETRSSFSRVSGSPSSGFRSQSWSRGSPSTVSSSYKRS 54
>GMHB_NEIMB (Q9JXI5) D,D-heptose 1,7-bisphosphate phosphatase (EC
3.1.3.-) (D-glycero-D-manno-heptose 1,7-bisphosphate
phosphatase)
Length = 187
Score = 30.8 bits (68), Expect = 6.1
Identities = 21/94 (22%), Positives = 41/94 (43%), Gaps = 9/94 (9%)
Query: 154 DSFIAFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMNS 213
D F+ D W+ +G + ++ + Y + A NQ ++ F+V + A M+
Sbjct: 17 DDFVKSVD--EWIPVEGSMDAVAFLTQAGYTVAVATNQSGIGRKYFTVQNLTEMHAKMHR 74
Query: 214 ASQKSDGG-------PQTGSRPARWRRPQPGTIK 240
+++ G P T + R+P+PG I+
Sbjct: 75 LVRQAGGEINGIWFCPHTDADNCNCRKPKPGMIE 108
>GMHB_NEIMA (Q9JWE9) D,D-heptose 1,7-bisphosphate phosphatase (EC
3.1.3.-) (D-glycero-D-manno-heptose 1,7-bisphosphate
phosphatase)
Length = 187
Score = 30.8 bits (68), Expect = 6.1
Identities = 21/94 (22%), Positives = 41/94 (43%), Gaps = 9/94 (9%)
Query: 154 DSFIAFADFWAWVVEQGDSEFISTVQTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMNS 213
D F+ D W+ +G + ++ + Y + A NQ ++ F+V + A M+
Sbjct: 17 DDFVKSVD--EWIPVEGSMDAVAFLTQAGYTVAVATNQSGIGRKYFTVQNLTEMHAKMHR 74
Query: 214 ASQKSDGG-------PQTGSRPARWRRPQPGTIK 240
+++ G P T + R+P+PG I+
Sbjct: 75 LVRQAGGEINGIWFCPHTDADNCNCRKPKPGMIE 108
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.138 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,437,225
Number of Sequences: 164201
Number of extensions: 1775311
Number of successful extensions: 4215
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4213
Number of HSP's gapped (non-prelim): 7
length of query: 387
length of database: 59,974,054
effective HSP length: 112
effective length of query: 275
effective length of database: 41,583,542
effective search space: 11435474050
effective search space used: 11435474050
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0052a.1


