

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
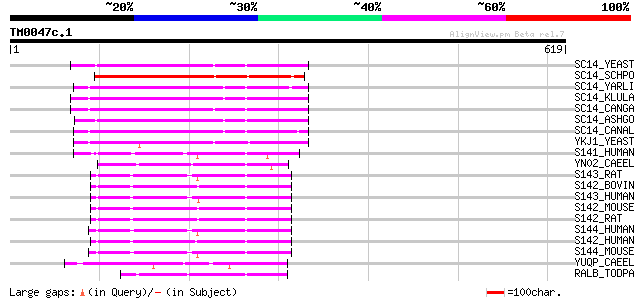
Query= TM0047c.1
(619 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SC14_YEAST (P24280) SEC14 cytosolic factor (Phosphatidylinositol... 180 1e-44
SC14_SCHPO (Q10137) Sec14 cytosolic factor (Phosphatidylinositol... 179 2e-44
SC14_YARLI (P45816) SEC14 cytosolic factor (Phosphatidylinositol... 178 3e-44
SC14_KLULA (P24859) SEC14 cytosolic factor (Phosphatidylinositol... 174 6e-43
SC14_CANGA (P53989) SEC14 cytosolic factor (Phosphatidylinositol... 174 8e-43
SC14_ASHGO (Q75DK1) SEC14 cytosolic factor (Phosphatidylinositol... 166 2e-40
SC14_CANAL (P46250) SEC14 cytosolic factor (Phosphatidylinositol... 165 3e-40
YKJ1_YEAST (P33324) 36.1 kDa protein in BUD2-MIF2 intergenic region 151 4e-36
S141_HUMAN (Q92503) SEC14-like protein 1 123 2e-27
YN02_CAEEL (Q03606) Hypothetical protein T23G5.2 in chromosome III 113 1e-24
S143_RAT (Q9Z1J8) SEC14-like protein 3 (45 kDa secretory protein... 97 1e-19
S142_BOVIN (P58875) SEC14-like protein 2 (Alpha-tocopherol assoc... 97 2e-19
S143_HUMAN (Q9UDX4) SEC14-like protein 3 (Tocopherol-associated ... 95 5e-19
S142_MOUSE (Q99J08) SEC14-like protein 2 (Alpha-tocopherol assoc... 88 8e-17
S142_RAT (Q99MS0) SEC14-like protein 2 (Alpha-tocopherol associa... 87 1e-16
S144_HUMAN (Q9UDX3) SEC14-like protein 4 (Tocopherol-associated ... 86 2e-16
S142_HUMAN (O76054) SEC14-like protein 2 (Alpha-tocopherol assoc... 86 2e-16
S144_MOUSE (Q8R0F9) SEC14-like protein 4 84 1e-15
YUQP_CAEEL (Q19895) Hypothetical protein F28H7.8 in chromosome V 69 3e-11
RALB_TODPA (P49193) Retinal-binding protein (RALBP) 57 1e-07
>SC14_YEAST (P24280) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 303
Score = 180 bits (456), Expect = 1e-44
Identities = 101/266 (37%), Positives = 156/266 (57%), Gaps = 5/266 (1%)
Query: 69 DEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYG 128
D +E + ELR+ L + G + D TLLRFL+AR F+++ +M+E WRK+YG
Sbjct: 29 DSAQEKALAELRKLLEDAGFIERLDDS--TLLRFLRARKFDVQLAKEMFENCEKWRKDYG 86
Query: 129 TDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYH 188
TDTIL++F ++E + ++YPQ YH DK+GRPVY E LG + + ++T+ +R LK
Sbjct: 87 TDTILQDFHYDEKPLIAKFYPQYYHKTDKDGRPVYFEELGAVNLHEMNKVTSEERMLKNL 146
Query: 189 VQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYP 248
V E+E +Q + PACS AA + ++ TI+D++G+ + + + + + + I NYYP
Sbjct: 147 VWEYESVVQYRLPACSRAAGHLVETSCTIMDLKGISISS-AYSVMSYVREASYISQNYYP 205
Query: 249 ETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLG 308
E + + YI+NA GF + + F DP T++KI IL S +LL+ I + LP G
Sbjct: 206 ERMGKFYIINAPFGF-STAFRLFKPFLDPVTVSKIFILGSSYQKELLKQIPAENLPVKFG 264
Query: 309 G-SCTCPTEGGCLKTSKGPWNDPDII 333
G S ++GG + GPW DP I
Sbjct: 265 GKSEVDESKGGLYLSDIGPWRDPKYI 290
>SC14_SCHPO (Q10137) Sec14 cytosolic factor
(Phosphatidylinositol/phosphatidyl-choline transfer
protein) (PI/PC TP) (Sporulation-specific protein 20)
Length = 286
Score = 179 bits (454), Expect = 2e-44
Identities = 93/235 (39%), Positives = 145/235 (61%), Gaps = 4/235 (1%)
Query: 95 DYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHG 154
D TLLRFL+AR FN++++++M+ + WRKE+G D +++ F ++E E V +YYPQ YH
Sbjct: 49 DDATLLRFLRARKFNLQQSLEMFIKCEKWRKEFGVDDLIKNFHYDEKEAVSKYYPQFYHK 108
Query: 155 VDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFST 214
D +GRPVY+E+LG +L +ITT +R ++ V E+E ++FPACS A I ++
Sbjct: 109 TDIDGRPVYVEQLGNIDLKKLYQITTPERMMQNLVYEYEMLALKRFPACSRKAGGLIETS 168
Query: 215 TTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKF 274
TI+D++G+G+ + + + + + I +YYPE + + Y++NA GF + + F
Sbjct: 169 CTIMDLKGVGITSI-HSVYSYIRQASSISQDYYPERMGKFYVINAPWGFSS-AFNLIKGF 226
Query: 275 CDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWND 329
D T+ KI IL S LLE I + LP LGG+C CP GGC + GPW++
Sbjct: 227 LDEATVKKIHILGSNYKSALLEQIPADNLPAKLGGNCQCP--GGCELSDAGPWHE 279
>SC14_YARLI (P45816) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 492
Score = 178 bits (452), Expect = 3e-44
Identities = 100/262 (38%), Positives = 156/262 (59%), Gaps = 6/262 (2%)
Query: 72 EETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDT 131
+E + EL+ L+ +G R DD TLLRFL+AR F++ +MWE WRKE+GT+T
Sbjct: 32 QEQKLGELKMILLTKGY-EDRTDDA-TLLRFLRARKFDVPLAQEMWENCEKWRKEFGTNT 89
Query: 132 ILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQE 191
ILE+F ++E +EV + YPQ YH DK+GRPVY+E +GK + + +ITT +R L+ V E
Sbjct: 90 ILEDFWYKEKKEVAKLYPQYYHKTDKDGRPVYVENVGKVNIHEMYKITTQERMLRNLVWE 149
Query: 192 FERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETL 251
+E ++ + PACS I ++ TILD++G+ + + ++ L A + I NYYPE +
Sbjct: 150 YESFVRHRLPACSRVVGHLIETSCTILDLKGVSLSSASQVYGFLKDA-SNIGQNYYPERM 208
Query: 252 HQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSC 311
+ Y++NA GF ++ ++F DP T++KI + S KLL + + LP GG
Sbjct: 209 GKFYLINAPFGF-STVFSVIKRFLDPVTVSKIHVYGSNYKEKLLAQVPAYNLPIKFGGQS 267
Query: 312 TCPTEGGCLKTSKGPWNDPDII 333
+ ++ G + GPW DP +
Sbjct: 268 S--SKIGVELSDDGPWRDPQFV 287
>SC14_KLULA (P24859) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 301
Score = 174 bits (441), Expect = 6e-43
Identities = 105/266 (39%), Positives = 153/266 (57%), Gaps = 5/266 (1%)
Query: 69 DEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYG 128
D +E + E R+ L G R DD TLLRFL+AR F++E + M+E WRKE+G
Sbjct: 28 DSEQEAKLKEFRELLESLGY-KERLDDS-TLLRFLRARKFDLEASKIMYENCEKWRKEFG 85
Query: 129 TDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYH 188
DTI E+F +EE V +YYPQ YH D +GRPVYIE LG + +++ +ITT +R LK
Sbjct: 86 VDTIFEDFHYEEKPLVAKYYPQYYHKTDNDGRPVYIEELGSVNLTQMYKITTQERMLKNL 145
Query: 189 VQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYP 248
V E+E ++ + PACS A + ++ TILD++G+ + + + + + A + I NYYP
Sbjct: 146 VWEYEAFVRYRLPACSRKAGYLVETSCTILDLKGISISSAAQVLSYVREA-SNIGQNYYP 204
Query: 249 ETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLG 308
E + + Y++NA GF + + F DP T++KI IL S LL+ I + LP G
Sbjct: 205 ERMGKFYLINAPFGF-STAFRLFKPFLDPVTVSKIFILGSSYQKDLLKQIPAENLPKKFG 263
Query: 309 G-SCTCPTEGGCLKTSKGPWNDPDII 333
G S EGG + GPW + + I
Sbjct: 264 GQSEVSEAEGGLYLSDIGPWREEEYI 289
>SC14_CANGA (P53989) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 302
Score = 174 bits (440), Expect = 8e-43
Identities = 102/266 (38%), Positives = 152/266 (56%), Gaps = 5/266 (1%)
Query: 69 DEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYG 128
DE +E + +LR L G R DD TLLRFL+AR F++ +M+E WRKEYG
Sbjct: 28 DEAQEGALKQLRSELEAAGF-KERLDDS-TLLRFLRARKFDVALAKEMFENCEKWRKEYG 85
Query: 129 TDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYH 188
T+TI+++F ++E V +YYPQ YH DK+GRPVY E LG + + + +ITT +R LK
Sbjct: 86 TNTIMQDFHYDEKPLVAKYYPQYYHKTDKDGRPVYFEELGAVNLTEMEKITTQERMLKNL 145
Query: 189 VQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYP 248
V E+E + + PACS AA + ++ T++D++G+ + + + + + + I NYYP
Sbjct: 146 VWEYESVVNYRLPACSRAAGYLVETSCTVMDLKGISISS-AYSVLSYVREASYISQNYYP 204
Query: 249 ETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLG 308
E + + Y++NA GF + + F DP T++KI IL S +LL+ I + LP G
Sbjct: 205 ERMGKFYLINAPFGF-STAFRLFKPFLDPVTVSKIFILGSSYQSELLKQIPAENLPSKFG 263
Query: 309 G-SCTCPTEGGCLKTSKGPWNDPDII 333
G S GG + GPW D I
Sbjct: 264 GKSEVDEAAGGLYLSDIGPWRDAKYI 289
>SC14_ASHGO (Q75DK1) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 311
Score = 166 bits (420), Expect = 2e-40
Identities = 101/262 (38%), Positives = 146/262 (55%), Gaps = 5/262 (1%)
Query: 73 ETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTI 132
E + ELR+ L + G D TLLRFL+AR F++ M+E WRKE G DTI
Sbjct: 35 EAALEELRKVLKQAGFTKRLDDS--TLLRFLRARKFDVAAARAMFENCEKWRKENGVDTI 92
Query: 133 LEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEF 192
E+F +EE V ++YPQ YH DK+GRPVYIE LG + + + +ITT +R LK + E+
Sbjct: 93 FEDFHYEEKPLVAKFYPQYYHKTDKDGRPVYIEELGAVNLTEMYKITTQERMLKNLIWEY 152
Query: 193 ERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLH 252
E + + PA S A + ++ TILD++G+ + + + + A + I NYYPE +
Sbjct: 153 ESFSRYRLPASSRQADCLVETSCTILDLKGISISAAAQVLSYVREA-SNIGQNYYPERMG 211
Query: 253 QMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGG-SC 311
+ Y++NA GF + + F DP T++KI IL S +LL+ I + LP GG S
Sbjct: 212 KFYMINAPFGF-SAAFRLFKPFLDPVTVSKIFILGSSYQKELLKQIPAENLPVKFGGQSD 270
Query: 312 TCPTEGGCLKTSKGPWNDPDII 333
EGG + GPW +P I
Sbjct: 271 VSEAEGGLYLSDIGPWRNPKYI 292
>SC14_CANAL (P46250) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 301
Score = 165 bits (418), Expect = 3e-40
Identities = 100/262 (38%), Positives = 148/262 (56%), Gaps = 5/262 (1%)
Query: 72 EETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDT 131
++T + RQ+L E G R DD +LLRFL+AR F+I+K I M+ WR+++G +T
Sbjct: 33 QKTTLDIFRQQLTELGY-KDRLDDA-SLLRFLRARKFDIQKAIDMFVACEKWREDFGVNT 90
Query: 132 ILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQE 191
IL++F +EE V + YP YH DK+GRPVY E LGK ++++ITT +R LK V E
Sbjct: 91 ILKDFHYEEKPIVAKMYPTYYHKTDKDGRPVYFEELGKVDLVKMLKITTQERMLKNLVWE 150
Query: 192 FERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETL 251
+E Q + PACS A + ++ T+LD+ G+ + + + A +KI +YYPE +
Sbjct: 151 YEAMCQYRLPACSRKAGYLVETSCTVLDLSGISVTSAYNVIGYVREA-SKIGQDYYPERM 209
Query: 252 HQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSC 311
+ Y++NA GF + + F DP T++KI IL +LL+ I LP GG
Sbjct: 210 GKFYLINAPFGF-STAFKLFKPFLDPVTVSKIHILGYSYKKELLKQIPPQNLPVKFGGMS 268
Query: 312 TCPTEGGCLKTSKGPWNDPDII 333
+ LK GPW DP+ I
Sbjct: 269 DVSDDDLLLK-DVGPWRDPEFI 289
>YKJ1_YEAST (P33324) 36.1 kDa protein in BUD2-MIF2 intergenic region
Length = 310
Score = 151 bits (382), Expect = 4e-36
Identities = 96/269 (35%), Positives = 148/269 (54%), Gaps = 11/269 (4%)
Query: 72 EETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDT 131
+E + + R L+E+ R DD TLLRFL+AR F+I +++M+ E WR+EYG +T
Sbjct: 29 QEEALLQFRSILLEKNY-KERLDDS-TLLRFLRARKFDINASVEMFVETERWREEYGANT 86
Query: 132 ILEEFEFEELEE------VLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYL 185
I+E++E + E + + YPQ YH VDK+GRP+Y E LG + ++ +ITT + L
Sbjct: 87 IIEDYENNKEAEDKERIKLAKMYPQYYHHVDKDGRPLYFEELGGINLKKMYKITTEKQML 146
Query: 186 KYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTN 245
+ V+E+E + PACS A I ++ T+LD++G+ + N + + + I N
Sbjct: 147 RNLVKEYELFATYRVPACSRRAGYLIETSCTVLDLKGISLSN-AYHVLSYIKDVADISQN 205
Query: 246 YYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPD 305
YYPE + + YI+++ GF M + + F DP T++KI IL S +LL+ I LP
Sbjct: 206 YYPERMGKFYIIHSPFGFSTM-FKMVKPFLDPVTVSKIFILGSSYKKELLKQIPIENLPV 264
Query: 306 FLGGSCTCPTEGGCLKTSK-GPWNDPDII 333
GG+ S GPW DP I
Sbjct: 265 KYGGTSVLHNPNDKFYYSDIGPWRDPRYI 293
>S141_HUMAN (Q92503) SEC14-like protein 1
Length = 715
Score = 123 bits (308), Expect = 2e-27
Identities = 93/262 (35%), Positives = 137/262 (51%), Gaps = 22/262 (8%)
Query: 72 EETVVHELRQRLIE--RGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGT 129
+E+ + LRQ L E +G +P D++ +LRFL+ARDFNI+K ++ + L WRK++
Sbjct: 255 QESCLIRLRQWLQETHKGKIPK--DEH--ILRFLRARDFNIDKAREIMCQSLTWRKQHQV 310
Query: 130 DTILEEFEFEELEEVLQ-YYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYH 188
D ILE + +VLQ YY G+H DK+GRP+Y+ RLG+ L+R + L+Y
Sbjct: 311 DYILETW---TPPQVLQDYYAGGWHHHDKDGRPLYVLRLGQMDTKGLVRALGEEALLRYV 367
Query: 189 VQEFERALQEKFPACSIAAK---RRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTN 245
+ E E+ C K R I S T ++D++GL M++ R L + ++
Sbjct: 368 LSVNE----ERLRRCEENTKVFGRPISSWTCLVDLEGLNMRHLWRPGVKALLRIIEVVEA 423
Query: 246 YYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQIL---DSKSLYKLLEVIDSSQ 302
YPETL ++ I+ A F +LW F D T K I D + LL+ ID
Sbjct: 424 NYPETLGRLLILRAPRVF-PVLWTLVSPFIDDNTRRKFLIYAGNDYQGPGGLLDYIDKEI 482
Query: 303 LPDFLGGSCTCPT-EGGCLKTS 323
+PDFL G C C EGG + S
Sbjct: 483 IPDFLSGECMCEVPEGGLVPKS 504
>YN02_CAEEL (Q03606) Hypothetical protein T23G5.2 in chromosome III
Length = 743
Score = 113 bits (283), Expect = 1e-24
Identities = 72/216 (33%), Positives = 110/216 (50%), Gaps = 7/216 (3%)
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKE 158
LLRFL+ARDF++ K M ++WRK++ D ILEE+ + + QY+P +H DK
Sbjct: 304 LLRFLRARDFDVAKAKDMVHASIIWRKQHNVDKILEEWTRPTV--IKQYFPGCWHNSDKA 361
Query: 159 GRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTIL 218
GRP+YI R G+ ++R ++ +K + E LQ A + I S + ++
Sbjct: 362 GRPMYILRFGQLDTKGMLRSCGVENLVKLTLSICEDGLQRAAEA-TRKLGTPISSWSLVV 420
Query: 219 DVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPK 278
D+ GL M++ R L + +I YPET+ Q+ +V A F +LW F D K
Sbjct: 421 DLDGLSMRHLWRPGVQCLLKIIEIVEANYPETMGQVLVVRAPRVF-PVLWTLISPFIDEK 479
Query: 279 TIAKIQILDSKS---LYKLLEVIDSSQLPDFLGGSC 311
T K + +L + I+ +PDFLGGSC
Sbjct: 480 TRKKFMVSGGSGGDLKEELRKHIEEKFIPDFLGGSC 515
>S143_RAT (Q9Z1J8) SEC14-like protein 3 (45 kDa secretory protein)
(rsec45)
Length = 400
Score = 97.4 bits (241), Expect = 1e-19
Identities = 71/227 (31%), Positives = 115/227 (50%), Gaps = 12/227 (5%)
Query: 91 PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQ 150
P DDY+ LLR+L+AR+F+++K+ M + + +RK D IL+ ++ E + +Y P
Sbjct: 31 PNPDDYF-LLRWLRARNFDLQKSEAMLRKYMEFRKTMDIDHILD---WQPPEVIQKYMPG 86
Query: 151 GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAK-- 208
G G D++G PV+ + +G P L+ T LK +++ ER L E C + +
Sbjct: 87 GLCGYDRDGCPVWYDIIGPLDPKGLLFSVTKQDLLKTKMRDCERILHE----CDLQTERL 142
Query: 209 -RRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKML 267
R+I + I D +GLG+K+F + + + YPETL M IV A F +
Sbjct: 143 GRKIETIVMIFDCEGLGLKHFWKPLVEVYQEFFGLLEENYPETLKFMLIVKATKLF-PVG 201
Query: 268 WPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
+ + F T KI +L + LL++I +LP GG+ T P
Sbjct: 202 YNLMKPFLSEDTRRKIVVLGNSWKEGLLKLISPEELPAHFGGTLTDP 248
>S142_BOVIN (P58875) SEC14-like protein 2 (Alpha-tocopherol
associated protein) (TAP) (bTAP) (Fragment)
Length = 387
Score = 96.7 bits (239), Expect = 2e-19
Identities = 71/224 (31%), Positives = 111/224 (48%), Gaps = 6/224 (2%)
Query: 91 PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQ 150
P DDY+ LLR+L+AR+FN++K+ M + + +RK+ D I+ ++ E V QY
Sbjct: 31 PNPDDYF-LLRWLRARNFNLQKSEAMLRKHVEFRKQKDIDNIMS---WQPPEVVQQYLSG 86
Query: 151 GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRR 210
G G D EG P++ + +G L+ + K +++ E LQE K+
Sbjct: 87 GMCGYDLEGSPIWYDIIGPLDAKGLLLSASKQDLFKTKMRDCELLLQECVRQTEKMGKK- 145
Query: 211 IFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPA 270
I +TT I D +GLG+K+ + A + YPETL +++IV A F + +
Sbjct: 146 IEATTLIYDCEGLGLKHLWKPAVEAYGEFLCMFEENYPETLKRLFIVKAPKLF-PVAYNL 204
Query: 271 AQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
+ F T KIQ+L + LL+ I QLP GG+ T P
Sbjct: 205 VKPFLSEDTRKKIQVLGANWKEVLLKYISPDQLPVEYGGTMTDP 248
>S143_HUMAN (Q9UDX4) SEC14-like protein 3 (Tocopherol-associated
protein 2)
Length = 400
Score = 95.1 bits (235), Expect = 5e-19
Identities = 70/227 (30%), Positives = 115/227 (49%), Gaps = 12/227 (5%)
Query: 91 PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQ 150
P DDY+ LLR+L+AR+F+++K+ + + + +RK D IL+ ++ E + +Y P
Sbjct: 31 PNPDDYF-LLRWLRARNFDLQKSEALLRKYMEFRKTMDIDHILD---WQPPEVIQKYMPG 86
Query: 151 GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKR- 209
G G D++G PV+ + +G P L+ T LK +++ ER L E C + +R
Sbjct: 87 GLCGYDRDGCPVWYDIIGPLDPKGLLFSVTKQDLLKTKMRDCERILHE----CDLQTERL 142
Query: 210 --RIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKML 267
+I + I D +GLG+K+F + + + YPETL M IV A F +
Sbjct: 143 GKKIETIVMIFDCEGLGLKHFWKPLVEVYQEFFGLLEENYPETLKFMLIVKATKLF-PVG 201
Query: 268 WPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
+ + F T KI +L + LL++I +LP GG+ T P
Sbjct: 202 YNLMKPFLSEDTRRKIIVLGNNWKEGLLKLISPEELPAQFGGTLTDP 248
>S142_MOUSE (Q99J08) SEC14-like protein 2 (Alpha-tocopherol
associated protein) (TAP)
Length = 403
Score = 87.8 bits (216), Expect = 8e-17
Identities = 66/224 (29%), Positives = 110/224 (48%), Gaps = 6/224 (2%)
Query: 91 PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQ 150
P DDY+ LLR+L+AR F+++K+ M + + +RK+ D I+ ++ E + QY
Sbjct: 31 PNPDDYF-LLRWLRARSFDLQKSEAMLRKHVEFRKQKDIDKIIS---WQPPEVIQQYLSG 86
Query: 151 GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRR 210
G G D +G PV+ + +G L+ + L+ +++ E LQE + K+
Sbjct: 87 GRCGYDLDGCPVWYDIIGPLDAKGLLFSASKQDLLRTKMRDCELLLQECIQQTTKLGKK- 145
Query: 211 IFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPA 270
I + T I D +GLG+K+ + A + YPETL ++++V A F + +
Sbjct: 146 IETITMIYDCEGLGLKHLWKPAVEAYGEFLTMFEENYPETLKRLFVVKAPKLF-PVAYNL 204
Query: 271 AQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
+ F T KI +L + LL+ I QLP GG+ T P
Sbjct: 205 IKPFLSEDTRRKIMVLGANWKEVLLKHISPDQLPVEYGGTMTDP 248
>S142_RAT (Q99MS0) SEC14-like protein 2 (Alpha-tocopherol associated
protein) (TAP) (Supernatant protein factor) (SPF)
(Squalene transfer protein)
Length = 403
Score = 87.0 bits (214), Expect = 1e-16
Identities = 66/224 (29%), Positives = 110/224 (48%), Gaps = 6/224 (2%)
Query: 91 PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQ 150
P DDY+ LLR+L+AR F+++K+ M + + +RK+ D I+ ++ E + QY
Sbjct: 31 PNPDDYF-LLRWLRARSFDLQKSEAMLRKHVEFRKQKDIDKIIS---WQPPEVIQQYLSG 86
Query: 151 GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRR 210
G G D +G PV+ + +G L+ + L+ +++ E LQE + K+
Sbjct: 87 GRCGYDLDGCPVWYDIIGPLDAKGLLFSASKQDLLRTKMRDCELLLQECTQQTAKLGKK- 145
Query: 211 IFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPA 270
I + T I D +GLG+K+ + A + YPETL ++++V A F + +
Sbjct: 146 IETITMIYDCEGLGLKHLWKPAVEAYGEFLTMFEENYPETLKRLFVVKAPKLF-PVAYNL 204
Query: 271 AQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
+ F T KI +L + LL+ I QLP GG+ T P
Sbjct: 205 IKPFLSEDTRKKIMVLGANWKEVLLKHISPDQLPVEYGGTMTDP 248
>S144_HUMAN (Q9UDX3) SEC14-like protein 4 (Tocopherol-associated
protein 3)
Length = 406
Score = 86.3 bits (212), Expect = 2e-16
Identities = 69/233 (29%), Positives = 110/233 (46%), Gaps = 16/233 (6%)
Query: 88 LLP--PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVL 145
LLP P DDY+ LLR+L+AR+F+++K+ M + +RK+ D I+ + EV+
Sbjct: 26 LLPILPNADDYF-LLRWLRARNFDLQKSEDMLRRHMEFRKQQDLDNIVT----WQPPEVI 80
Query: 146 QYYPQ-GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACS 204
Q Y G G D EG PVY +G P L+ + ++ ++ E L E C
Sbjct: 81 QLYDSGGLCGYDYEGCPVYFNIIGSLDPKGLLLSASKQDMIRKRIKVCELLLHE----CE 136
Query: 205 IAAK---RRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGS 261
+ + R+I + D++GL +K+ + A + I YPETL + ++ A
Sbjct: 137 LQTQKLGRKIEMALMVFDMEGLSLKHLWKPAVEVYQQFFSILEANYPETLKNLIVIRAPK 196
Query: 262 GFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
F + + + F +T KI IL +L + I QLP GG+ T P
Sbjct: 197 LF-PVAFNLVKSFMSEETRRKIVILGDNWKQELTKFISPDQLPVEFGGTMTDP 248
>S142_HUMAN (O76054) SEC14-like protein 2 (Alpha-tocopherol
associated protein) (TAP) (hTAP) (Supernatant protein
factor) (SPF) (Squalene transfer protein)
Length = 403
Score = 86.3 bits (212), Expect = 2e-16
Identities = 65/224 (29%), Positives = 110/224 (49%), Gaps = 6/224 (2%)
Query: 91 PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQ 150
P DDY+ LLR+L+AR F+++K+ M + + +RK+ D I+ ++ E + QY
Sbjct: 31 PNPDDYF-LLRWLRARSFDLQKSEAMLRKHVEFRKQKDIDNIIS---WQPPEVIQQYLSG 86
Query: 151 GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRR 210
G G D +G PV+ + +G L+ + L+ ++E E LQE + R+
Sbjct: 87 GMCGYDLDGCPVWYDIIGPLDAKGLLFSASKQDLLRTKMRECELLLQECAHQTTKLG-RK 145
Query: 211 IFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPA 270
+ + T I D +GLG+K+ + A + YPETL ++++V A F + +
Sbjct: 146 VETITIIYDCEGLGLKHLWKPAVEAYGEFLCMFEENYPETLKRLFVVKAPKLF-PVAYNL 204
Query: 271 AQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
+ F T KI +L + LL+ I Q+P GG+ T P
Sbjct: 205 IKPFLSEDTRKKIMVLGANWKEVLLKHISPDQVPVEYGGTMTDP 248
>S144_MOUSE (Q8R0F9) SEC14-like protein 4
Length = 403
Score = 84.0 bits (206), Expect = 1e-15
Identities = 66/233 (28%), Positives = 113/233 (48%), Gaps = 16/233 (6%)
Query: 88 LLP--PRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVL 145
LLP P+ DDY+ LLR+L+AR+F+++K+ M + + +R + D IL + EV+
Sbjct: 26 LLPTLPKADDYF-LLRWLRARNFDLKKSEDMLRKHVEFRNQQNLDQILT----WQAPEVI 80
Query: 146 QYYPQ-GYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACS 204
Q Y G G D EG PV+ + +G P L + ++ ++ E L E C
Sbjct: 81 QLYDSGGLSGYDYEGCPVWFDIIGTMDPKGLFMSASKQDMIRKRIKVCEMLLHE----CE 136
Query: 205 IAAK---RRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGS 261
+ ++ R+I + D++GL +++ + A + I YPET+ + I+ A
Sbjct: 137 LQSQKLGRKIERMVMVFDMEGLSLRHLWKPAVEVYQQFFAILEANYPETVKNLIIIRAPK 196
Query: 262 GFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCP 314
F + + + F +T KI IL +L++ + QLP GG+ T P
Sbjct: 197 LF-PVAFNLVKSFMGEETQKKIVILGGNWKQELVKFVSPDQLPVEFGGTMTDP 248
>YUQP_CAEEL (Q19895) Hypothetical protein F28H7.8 in chromosome V
Length = 453
Score = 69.3 bits (168), Expect = 3e-11
Identities = 62/255 (24%), Positives = 115/255 (44%), Gaps = 19/255 (7%)
Query: 62 VSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEML 121
V IE++ D E V RL ++ PR+D + +LR+L++ DFNI KT+ + ++ L
Sbjct: 9 VDIENMNDAAIEQV------RLQVSDVIDPRYDTKWNMLRWLQSNDFNIPKTVHLLKKHL 62
Query: 122 LWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKE---GRPVYIERLGKAHPSRLMRI 178
WRK+ D + + + ++ P G ++ R V ++R G+ S LM+
Sbjct: 63 KWRKDRKLDEPESQSLLQFSDARRKHAPIDIIGPQRKEDGDRLVVVDRAGRIDVSGLMKS 122
Query: 179 TTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAA 238
YL + FE +Q + + + I D++ L NF T ++
Sbjct: 123 VQPTEYLHEMFRSFEE-IQRRLMKMEAETGVQCY-MHYIFDLEAL---NFDPTLLGVVNG 177
Query: 239 MTKID----TNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKL 294
++ +Y E + + ++N+ S + +LW A F ++ +I S +L
Sbjct: 178 PFRVSWQLVGQHYREFIDKFIVINSPS-YINVLWSALSPFIPEQSKQRIVFAGSNWKEEL 236
Query: 295 LEVIDSSQLPDFLGG 309
L+++D LP+ GG
Sbjct: 237 LDIVDKECLPERYGG 251
>RALB_TODPA (P49193) Retinal-binding protein (RALBP)
Length = 342
Score = 57.0 bits (136), Expect = 1e-07
Identities = 43/187 (22%), Positives = 89/187 (46%), Gaps = 4/187 (2%)
Query: 124 RKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDR 183
R++ G DT++ E+ ++ + ++ G G DK+G + IE G +M
Sbjct: 3 REQMGADTLIAEYTPPDV--IQKFMTGGDVGHDKDGSVLRIEPWGYLDMKGIMYSCKKSD 60
Query: 184 YLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKID 243
K + + E+ L++ A S + T + D++ +G K+ + ++ + ++
Sbjct: 61 LEKSKLLQCEKHLKD-LEAQSEKVGKPCTGLTVVFDMENVGSKHMWKPGLDMYLYLVQVL 119
Query: 244 TNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQL 303
+ YPE + +++++NA + F +L+ + KI +L LLE ID+ +L
Sbjct: 120 EDNYPEMMKRLFVINAPTLF-PVLYKLVKPLLSEDMKNKIFVLGGDYKDTLLEYIDAEEL 178
Query: 304 PDFLGGS 310
P +LGG+
Sbjct: 179 PAYLGGT 185
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,946,839
Number of Sequences: 164201
Number of extensions: 2994735
Number of successful extensions: 9681
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 42
Number of HSP's that attempted gapping in prelim test: 9581
Number of HSP's gapped (non-prelim): 87
length of query: 619
length of database: 59,974,054
effective HSP length: 116
effective length of query: 503
effective length of database: 40,926,738
effective search space: 20586149214
effective search space used: 20586149214
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0047c.1


