

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047b.8
(171 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
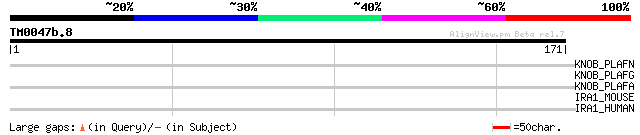
Score E
Sequences producing significant alignments: (bits) Value
KNOB_PLAFN (P06719) Knob-associated histidine-rich protein precu... 30 3.5
KNOB_PLAFG (P09346) Knob-associated histidine-rich protein precu... 30 3.5
KNOB_PLAFA (P13817) Knob-associated histidine-rich protein precu... 30 3.5
IRA1_MOUSE (Q62406) Interleukin-1 receptor-associated kinase 1 (... 29 4.6
IRA1_HUMAN (P51617) Interleukin-1 receptor-associated kinase 1 (... 28 7.9
>KNOB_PLAFN (P06719) Knob-associated histidine-rich protein
precursor (KAHRP)
Length = 657
Score = 29.6 bits (65), Expect = 3.5
Identities = 30/105 (28%), Positives = 44/105 (41%), Gaps = 27/105 (25%)
Query: 48 NLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCAST 107
N++ KK C +E NY AF E CPY +ND + +
Sbjct: 172 NVVDGKKDCDEKYEAANY-------------------AFSE-ECPY--TVNDYSQENGPN 209
Query: 108 MFSYINLYGKYPPGLFAHECREGKEGLACPALPPSALADDTSSQV 152
+F+ L ++P G+ E EGKE LA P L D+ +Q+
Sbjct: 210 IFA---LRKRFPLGM-NDEDEEGKEALAIKDKLPGGL-DEYQNQL 249
>KNOB_PLAFG (P09346) Knob-associated histidine-rich protein
precursor (KAHRP) (KP)
Length = 634
Score = 29.6 bits (65), Expect = 3.5
Identities = 30/105 (28%), Positives = 44/105 (41%), Gaps = 27/105 (25%)
Query: 48 NLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCAST 107
N++ KK C +E NY AF E CPY +ND + +
Sbjct: 168 NVVDGKKDCDEKYEAANY-------------------AFSE-ECPY--TVNDYSQENGPN 205
Query: 108 MFSYINLYGKYPPGLFAHECREGKEGLACPALPPSALADDTSSQV 152
+F+ L ++P G+ E EGKE LA P L D+ +Q+
Sbjct: 206 IFA---LRKRFPLGM-NDEDEEGKEALAIKDKLPGGL-DEYQNQL 245
>KNOB_PLAFA (P13817) Knob-associated histidine-rich protein
precursor (KAHRP) (HRPI) (Fragment)
Length = 473
Score = 29.6 bits (65), Expect = 3.5
Identities = 30/105 (28%), Positives = 44/105 (41%), Gaps = 27/105 (25%)
Query: 48 NLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCAST 107
N++ KK C +E NY AF E CPY +ND + +
Sbjct: 168 NVVDGKKDCDEKYEAANY-------------------AFSE-ECPY--TVNDYSQENGPN 205
Query: 108 MFSYINLYGKYPPGLFAHECREGKEGLACPALPPSALADDTSSQV 152
+F+ L ++P G+ E EGKE LA P L D+ +Q+
Sbjct: 206 IFA---LRKRFPLGM-NDEDEEGKEALAIKDKLPGGL-DEYQNQL 245
>IRA1_MOUSE (Q62406) Interleukin-1 receptor-associated kinase 1 (EC
2.7.1.37) (IRAK-1) (IRAK) (Pelle-like protein kinase)
(mPLK)
Length = 710
Score = 29.3 bits (64), Expect = 4.6
Identities = 15/24 (62%), Positives = 17/24 (70%)
Query: 28 SSSSTFLSDAVFGSQAHTGRNLLQ 51
SS+STFLS A GSQ H+ LLQ
Sbjct: 137 SSASTFLSPAFPGSQTHSESELLQ 160
>IRA1_HUMAN (P51617) Interleukin-1 receptor-associated kinase 1 (EC
2.7.1.37) (IRAK-1)
Length = 712
Score = 28.5 bits (62), Expect = 7.9
Identities = 14/22 (63%), Positives = 16/22 (72%)
Query: 28 SSSSTFLSDAVFGSQAHTGRNL 49
SS+STFLS A GSQ H+G L
Sbjct: 137 SSASTFLSPAFPGSQTHSGPEL 158
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.137 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,825,737
Number of Sequences: 164201
Number of extensions: 737657
Number of successful extensions: 1792
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1790
Number of HSP's gapped (non-prelim): 5
length of query: 171
length of database: 59,974,054
effective HSP length: 102
effective length of query: 69
effective length of database: 43,225,552
effective search space: 2982563088
effective search space used: 2982563088
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0047b.8


