

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0045.6
(214 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
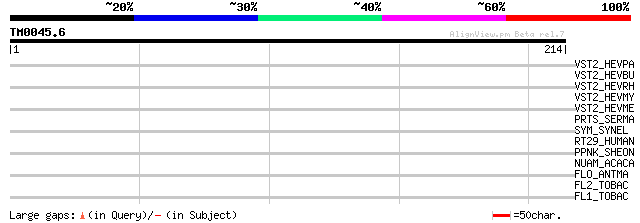
Score E
Sequences producing significant alignments: (bits) Value
VST2_HEVPA (P33426) Structural protein 2 precursor 32 1.1
VST2_HEVBU (P29326) Structural protein 2 precursor 32 1.1
VST2_HEVRH (Q00270) Structural protein 2 (Fragment) 31 1.9
VST2_HEVMY (Q04611) Structural protein 2 precursor 31 1.9
VST2_HEVME (Q03500) Structural protein 2 precursor 30 3.2
PRTS_SERMA (P09489) Extracellular serine protease precursor (EC ... 30 5.5
SYM_SYNEL (P59081) Methionyl-tRNA synthetase (EC 6.1.1.10) (Meth... 29 7.1
RT29_HUMAN (P51398) Mitochondrial 28S ribosomal protein S29 (S29... 29 7.1
PPNK_SHEON (Q8EGS1) Probable inorganic polyphosphate/ATP-NAD kin... 29 9.3
NUAM_ACACA (Q37373) NADH-ubiquinone oxidoreductase 75 kDa subuni... 29 9.3
FLO_ANTMA (P23915) Floricaula protein 29 9.3
FL2_TOBAC (Q40505) Floricaula/leafy homolog 2 (NFL2) 29 9.3
FL1_TOBAC (Q40504) Floricaula/leafy homolog 1 (NFL1) 29 9.3
>VST2_HEVPA (P33426) Structural protein 2 precursor
Length = 660
Score = 32.0 bits (71), Expect = 1.1
Identities = 16/66 (24%), Positives = 36/66 (54%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L ++ V P S ++ ++ F ++ + +L+ W+ G T+ + YN +
Sbjct: 506 GAQAVARSLDWTKVTLDGRPLSTIQQYSKTFFVLPLRGKLSFWEAGTTKAGYPYNYNTTA 565
Query: 135 DEFLVV 140
+ L+V
Sbjct: 566 SDQLLV 571
>VST2_HEVBU (P29326) Structural protein 2 precursor
Length = 660
Score = 32.0 bits (71), Expect = 1.1
Identities = 16/66 (24%), Positives = 36/66 (54%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L ++ V P S ++ ++ F ++ + +L+ W+ G T+ + YN +
Sbjct: 506 GAQAVARSLDWTKVTLDGRPLSTIQQYSKTFFVLPLRGKLSFWEAGTTKAGYPYNYNTTA 565
Query: 135 DEFLVV 140
+ L+V
Sbjct: 566 SDQLLV 571
>VST2_HEVRH (Q00270) Structural protein 2 (Fragment)
Length = 485
Score = 31.2 bits (69), Expect = 1.9
Identities = 16/66 (24%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L ++ V P S ++ + F ++ + +L+ W+ G T+ + YN +
Sbjct: 374 GAQAVARSLDWTKVTLDGRPLSTIQQYPKTFFVLPLRGKLSFWEAGTTKAGYPYNYNTTA 433
Query: 135 DEFLVV 140
+ L+V
Sbjct: 434 SDQLLV 439
>VST2_HEVMY (Q04611) Structural protein 2 precursor
Length = 660
Score = 31.2 bits (69), Expect = 1.9
Identities = 16/66 (24%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L ++ V P S ++ + F ++ + +L+ W+ G T+ + YN +
Sbjct: 506 GAQAVARSLDWTKVTLDGRPLSTIQQYPKTFFVLPLRGKLSFWEAGTTKAGYPYNYNTTA 565
Query: 135 DEFLVV 140
+ L+V
Sbjct: 566 SDQLLV 571
>VST2_HEVME (Q03500) Structural protein 2 precursor
Length = 659
Score = 30.4 bits (67), Expect = 3.2
Identities = 15/66 (22%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L +S V P V+ ++ F ++ + +L+ W+ G T+ + YN +
Sbjct: 505 GAQAVARSLDWSKVTLDGRPLPTVEQYSKTFFVLPLRGKLSFWEAGTTKAGYPYNYNTTA 564
Query: 135 DEFLVV 140
+ +++
Sbjct: 565 SDQILI 570
>PRTS_SERMA (P09489) Extracellular serine protease precursor (EC
3.4.21.-)
Length = 1045
Score = 29.6 bits (65), Expect = 5.5
Identities = 37/143 (25%), Positives = 52/143 (35%), Gaps = 7/143 (4%)
Query: 44 DGSVRDMITLAKYNGGDLDTYFEHPDFEPLVEGPEALAQVLGFSHVWHFAAPSNVKAFAW 103
DGSV + + N G L F G A +G HV H A +
Sbjct: 495 DGSVASNVYIE--NSGTLSGEGTVGAFRAARSGSVAPGNGIGTLHVLHDAIFDRGSQYN- 551
Query: 104 RFLLVKIPSRLNLWKLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNY 163
V++ K+ A R ++ G E L+ L NK TI++ +
Sbjct: 552 ----VEVADNGRSDKIAARRAFLNGGSVNVSLERSQNLLSQNEAQSLLGNKYTILTTTDG 607
Query: 164 VRDYFDIADTEYYFYKFNLDYIG 186
V F+ A+ Y F K LDY G
Sbjct: 608 VTGRFENANPSYPFVKVALDYRG 630
>SYM_SYNEL (P59081) Methionyl-tRNA synthetase (EC 6.1.1.10)
(Methionine--tRNA ligase) (MetRS)
Length = 529
Score = 29.3 bits (64), Expect = 7.1
Identities = 16/34 (47%), Positives = 19/34 (55%), Gaps = 3/34 (8%)
Query: 53 LAKYNGGDLDTYFEHPDFEPLVEGPEALAQVLGF 86
L+KY LD Y EHPDF V+ P +VL F
Sbjct: 165 LSKYQQALLDHYAEHPDF---VQPPSRRNEVLSF 195
>RT29_HUMAN (P51398) Mitochondrial 28S ribosomal protein S29 (S29mt)
(MRP-S29) (Death-associated protein 3) (DAP-3) (Ionizing
radiation resistance conferring protein)
Length = 398
Score = 29.3 bits (64), Expect = 7.1
Identities = 20/73 (27%), Positives = 34/73 (46%), Gaps = 9/73 (12%)
Query: 62 DTYFEHPDFEPLVEGPEALAQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGA 121
+T F +P L+ G + + L HV HF A + +L++ IP +LW +
Sbjct: 114 NTSFAYPAIRYLLYGEKGTGKTLSLCHVIHFCAKQD-------WLILHIPD-AHLW-VKN 164
Query: 122 TRGLISFGYNRSK 134
R L+ YN+ +
Sbjct: 165 CRDLLQSSYNKQR 177
>PPNK_SHEON (Q8EGS1) Probable inorganic polyphosphate/ATP-NAD kinase
(EC 2.7.1.23) (Poly(P)/ATP NAD kinase)
Length = 309
Score = 28.9 bits (63), Expect = 9.3
Identities = 16/48 (33%), Positives = 19/48 (39%)
Query: 9 DIDYDTFHIFLLEVMAIAHGYQNFQNMWRNIGAYEDGSVRDMITLAKY 56
D ++DT H FLLE HG N N G + MI Y
Sbjct: 134 DGEFDTEHRFLLEAEVYRHGQLKASNTAVNEAVLHPGKIAHMIEFEVY 181
>NUAM_ACACA (Q37373) NADH-ubiquinone oxidoreductase 75 kDa subunit
(EC 1.6.5.3) (EC 1.6.99.3) (Complex I-75KD) (CI-75KD)
(NADH dehydrogenase subunit 11)
Length = 675
Score = 28.9 bits (63), Expect = 9.3
Identities = 19/49 (38%), Positives = 24/49 (48%), Gaps = 3/49 (6%)
Query: 134 KDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNL 182
K L + CN L N + I LR Y+ DY D+ +T Y F KF L
Sbjct: 291 KKAMLFIKKFFCNFLGF--NHSSFIPLRGYIGDYLDL-ETIYTFKKFLL 336
>FLO_ANTMA (P23915) Floricaula protein
Length = 396
Score = 28.9 bits (63), Expect = 9.3
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Query: 32 FQNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFE-HP 68
F+ N+GA+ + ++ +A G D+DT F HP
Sbjct: 318 FKERGENVGAWRQACYKPLVAIAARQGWDIDTIFNAHP 355
>FL2_TOBAC (Q40505) Floricaula/leafy homolog 2 (NFL2)
Length = 416
Score = 28.9 bits (63), Expect = 9.3
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Query: 32 FQNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFE-HP 68
F+ N+GA+ + ++ +A G D+DT F HP
Sbjct: 335 FKERGENVGAWRQACYKPLVAIAARQGWDIDTIFNAHP 372
>FL1_TOBAC (Q40504) Floricaula/leafy homolog 1 (NFL1)
Length = 413
Score = 28.9 bits (63), Expect = 9.3
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Query: 32 FQNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFE-HP 68
F+ N+GA+ + ++ +A G D+DT F HP
Sbjct: 335 FKERGENVGAWRQACYKPLVAIAARQGWDIDTIFNAHP 372
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.144 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,922,218
Number of Sequences: 164201
Number of extensions: 1085741
Number of successful extensions: 2770
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2769
Number of HSP's gapped (non-prelim): 13
length of query: 214
length of database: 59,974,054
effective HSP length: 106
effective length of query: 108
effective length of database: 42,568,748
effective search space: 4597424784
effective search space used: 4597424784
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0045.6


