

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
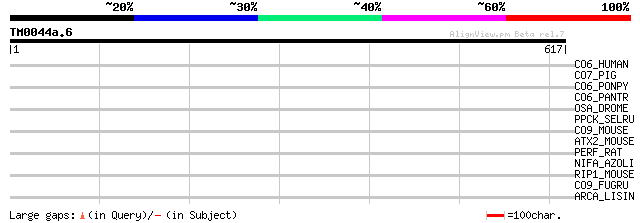
Query= TM0044a.6
(617 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CO6_HUMAN (P13671) Complement component C6 precursor 40 0.024
CO7_PIG (Q9TUQ3) Complement component C7 precursor 38 0.069
CO6_PONPY (P61135) Complement component C6 precursor 37 0.12
CO6_PANTR (P61134) Complement component C6 precursor 37 0.15
OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein) 33 1.7
PPCK_SELRU (O83023) Phosphoenolpyruvate carboxykinase [ATP] (EC ... 33 2.2
CO9_MOUSE (P06683) Complement component C9 precursor 33 2.2
ATX2_MOUSE (O70305) Ataxin-2 (Spinocerebellar ataxia type 2 prot... 33 2.9
PERF_RAT (P35763) Perforin 1 precursor (P1) (Lymphocyte pore for... 32 3.8
NIFA_AZOLI (P54929) Nif-specific regulatory protein 32 6.4
RIP1_MOUSE (Q7TNF8) Peripheral-type benzodiazepine receptor-asso... 31 8.4
CO9_FUGRU (P79755) Complement component C9 precursor 31 8.4
ARCA_LISIN (Q92FR7) Arginine deiminase (EC 3.5.3.6) (ADI) (Argin... 31 8.4
>CO6_HUMAN (P13671) Complement component C6 precursor
Length = 934
Score = 39.7 bits (91), Expect = 0.024
Identities = 37/160 (23%), Positives = 68/160 (42%), Gaps = 14/160 (8%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF 222
+P +++ A SR +GTH ++GG + + Q S ++ L E+ +
Sbjct: 345 LPLEYNSALYSRIFDDFGTHYFTSGSLGGVYDL-LYQFSSEELKNSGLTE--EEAKHCVR 401
Query: 223 SDVRSPPLLQRKAADGKQKVPEVFNRVMQSNTMSFTSISETS--------SKDGLTIICS 274
+ + L +K K + N++ + + SF +E S S+ G +
Sbjct: 402 IETKKRVLFAKKT---KVEHRCTTNKLSEKHEGSFIQGAEKSISLIRGGRSEYGAALAWE 458
Query: 275 KRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIP 314
K + + + S WL++V NP I F+ PI L+ IP
Sbjct: 459 KGSSGLEEKTFSEWLESVKENPAVIDFELAPIVDLVRNIP 498
>CO7_PIG (Q9TUQ3) Complement component C7 precursor
Length = 843
Score = 38.1 bits (87), Expect = 0.069
Identities = 47/182 (25%), Positives = 72/182 (38%), Gaps = 32/182 (17%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDL-----RRHLEDL 217
+P +D +A R I YGTH + ++GG+ + + S K+ DL ++
Sbjct: 283 LPPLYDYSAYRRLIDQYGTHYLQSGSLGGEYKV-LFYVDSEKVAESDLGSEDKKKCASSH 341
Query: 218 GDFLFSDVRSPPLLQR---KAADGKQK-----VPEVFNRVMQSNTMSFTSISETSSKDGL 269
FLF + K+A G Q VP F R + +S S E + DG
Sbjct: 342 ISFLFKSSKHKCKAMEEALKSASGTQSNVLRGVP--FVRGGRPGFVSGLSYLELDNPDG- 398
Query: 270 TIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIPGSG----YLSHAINL 325
K +SSW +V P+ I K P+ L+ +P + YL A+
Sbjct: 399 -----------NKQRYSSWAGSVTDLPQVIKQKLTPLYELVKEVPCASVKRLYLKRALEE 447
Query: 326 YL 327
YL
Sbjct: 448 YL 449
>CO6_PONPY (P61135) Complement component C6 precursor
Length = 934
Score = 37.4 bits (85), Expect = 0.12
Identities = 36/160 (22%), Positives = 67/160 (41%), Gaps = 14/160 (8%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF 222
+P +++ A SR +GTH ++GG + + Q S ++ L E+ +
Sbjct: 345 LPLEYNSALYSRIFDDFGTHYFTSGSLGGVYDL-LYQFSSEELKNSGLTE--EEAKHCVR 401
Query: 223 SDVRSPPLLQRKAADGKQKVPEVFNRVMQSNTMSFTSISETS--------SKDGLTIICS 274
+ + L +K K + N++ + + SF +E S S+ +
Sbjct: 402 IETKKRVLFAKKT---KVEHRCTTNKLSEKHEGSFIEGAEKSISLIRGGRSEYAAALAWE 458
Query: 275 KRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIP 314
K + + + S WL++V NP I F+ PI L+ IP
Sbjct: 459 KGSSGLEEKTFSEWLESVKENPAVIDFELAPIVDLVRNIP 498
>CO6_PANTR (P61134) Complement component C6 precursor
Length = 934
Score = 37.0 bits (84), Expect = 0.15
Identities = 36/160 (22%), Positives = 67/160 (41%), Gaps = 14/160 (8%)
Query: 163 VPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF 222
+P +++ A SR +GTH ++GG + + Q S ++ L E+ +
Sbjct: 345 LPLEYNSALYSRIFDDFGTHYFTSGSLGGVYDL-LYQFSSEELKNSGLTE--EEAKHCVR 401
Query: 223 SDVRSPPLLQRKAADGKQKVPEVFNRVMQSNTMSFTSISETS--------SKDGLTIICS 274
+ + L +K K + N++ + + SF +E S S+ +
Sbjct: 402 IETKKRVLFVKKT---KVEHRCTTNKLSEKHEGSFIQGAEKSISLIRGGRSEYAAALAWE 458
Query: 275 KRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLAGIP 314
K + + + S WL++V NP I F+ PI L+ IP
Sbjct: 459 KGSSGLEEKTFSEWLESVKENPAVIDFELAPIVDLVRNIP 498
>OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein)
Length = 2716
Score = 33.5 bits (75), Expect = 1.7
Identities = 34/110 (30%), Positives = 44/110 (39%), Gaps = 8/110 (7%)
Query: 508 SIRKTEWAAA---PEASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGI--YPDGP 562
S +KT AA+ P + TN +T S Q + PP AP G YP G
Sbjct: 1108 SKKKTAKAASVPSPGGGHLDAGTTN--STGSSNSQDSFPAPPGSAPNAAIDGYPGYPGGS 1165
Query: 563 PVPVRAGKLLKYVEAAEVVRGP-HDTPGHWLVTAAKLVTEGGKIGLQVKF 611
P PV +G Y A ++ R P + P AA V G I + F
Sbjct: 1166 PYPVASGPQPDYATAGQMQRPPSQNNPQTPHPGAAAAVAAGDNISVSNPF 1215
>PPCK_SELRU (O83023) Phosphoenolpyruvate carboxykinase [ATP] (EC
4.1.1.49) (PEP carboxykinase) (Phosphoenolpyruvate
carboxylase) (PEPCK)
Length = 539
Score = 33.1 bits (74), Expect = 2.2
Identities = 23/84 (27%), Positives = 38/84 (44%), Gaps = 4/84 (4%)
Query: 379 LHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALH----IHHLSSLPNKMILSSETSTPSMW 434
LH S + L E++ GL Y +G+ + A++ I S +K I+ ETS ++W
Sbjct: 18 LHNPSYKTLFEEETKEGLTGYEQGQVSELGAVNVKTGIFTGRSPKDKFIVDDETSHDTVW 77
Query: 435 RGSDDNESSNQFLEPVRWKRFSNV 458
S+D + N P W +
Sbjct: 78 WDSEDYHNDNHRATPETWNALKEI 101
>CO9_MOUSE (P06683) Complement component C9 precursor
Length = 548
Score = 33.1 bits (74), Expect = 2.2
Identities = 38/162 (23%), Positives = 68/162 (41%), Gaps = 12/162 (7%)
Query: 161 KSVPAQWDPAALSRFIQTYGTHLIVGMAVGGQ-DVICVKQKHSSKIPPGDLR--RHL--- 214
K++P ++ F++TYGTH ++GGQ +++ V K S K DL +H
Sbjct: 322 KALPTSYEKGEYFGFLETYGTHYSTSGSLGGQYEIVYVLDKASMKEKGVDLNDVKHCLGF 381
Query: 215 -EDLGDFLFSDVRSPPLLQRKAADG--KQKVPEVFNRVMQSNTMSFTSISETSSKDGLTI 271
DL L D++ + ADG K + N + S +++ +
Sbjct: 382 NMDLRIPLQDDLKDASVTASVNADGCIKTDNGKTVNITRDNIIDDVISFIRGGTREQAIL 441
Query: 272 ICSK--RGGDML-KHSHSSWLQTVPSNPEAILFKFVPISSLL 310
+ K RG K ++W ++ + P I + PI +L+
Sbjct: 442 LKEKILRGDKTFDKTDFANWASSLANAPALISQRMSPIYNLI 483
>ATX2_MOUSE (O70305) Ataxin-2 (Spinocerebellar ataxia type 2 protein
homolog)
Length = 1285
Score = 32.7 bits (73), Expect = 2.9
Identities = 16/46 (34%), Positives = 28/46 (60%)
Query: 519 EASRKSSFLTNLSTTFSFTQQSATSGPPKQAPAMLNSGIYPDGPPV 564
EAS K SF+ + S++ + T S+ + P +P+ML++ + GP V
Sbjct: 806 EASAKDSFIDSSSSSSNCTSGSSKTNSPSISPSMLSNAEHKRGPEV 851
>PERF_RAT (P35763) Perforin 1 precursor (P1) (Lymphocyte pore
forming protein) (Cytolysin)
Length = 554
Score = 32.3 bits (72), Expect = 3.8
Identities = 36/167 (21%), Positives = 68/167 (40%), Gaps = 19/167 (11%)
Query: 171 ALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSD------ 224
A R I +YGTH I + +GG+ + + G +++GD L +
Sbjct: 210 AYRRLISSYGTHFITAVDLGGRVSVLTALRTCQLTLDG---LTADEVGDCLSVEAQVSIG 266
Query: 225 VRSPPLLQRKAADGKQKVPEV---FNRVMQSNTMSFTSISETSSKDGLTIICSKRGGDML 281
++ + KA + K+K ++ F++ + + SS D L G
Sbjct: 267 AQASVSSEYKACEEKKKQHKIATSFHQTYRERHVEVLGGPLDSSNDLLF------GNQAT 320
Query: 282 KHSHSSWLQTVPSNPEAILFKFVPISSLLA-GIPGSGYLSHAINLYL 327
S+W+ ++P+ P+ + + P+ LL P L AI+ Y+
Sbjct: 321 PEHFSTWIASLPTRPDVVDYSLEPLHILLEDSDPKREALRQAISHYV 367
>NIFA_AZOLI (P54929) Nif-specific regulatory protein
Length = 624
Score = 31.6 bits (70), Expect = 6.4
Identities = 28/100 (28%), Positives = 41/100 (41%), Gaps = 6/100 (6%)
Query: 471 NSSTGVYIVTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNL 530
N T V ++++RC+WP NV R L I + + + E+ S L N
Sbjct: 396 NGMTLVMEDEALEVLNRCTWPGNV---RELENCIERAATQSRDGIIRTESLSCSLNLCNS 452
Query: 531 STTFSFTQQSATSG--PPKQAPAMLNSGIYPDGPPVPVRA 568
S F + A+ G P P +N + P P VP A
Sbjct: 453 SVLFQYRTLGASVGGLAPSMGPGSVNR-VPPGRPGVPAPA 491
>RIP1_MOUSE (Q7TNF8) Peripheral-type benzodiazepine
receptor-associated protein 1 (PRAX-1) (Peripheral
benzodiazepine receptor interacting protein) (PBR-IP)
(RIM binding protein 1) (RIM-BP1)
Length = 1846
Score = 31.2 bits (69), Expect = 8.4
Identities = 30/114 (26%), Positives = 47/114 (40%), Gaps = 25/114 (21%)
Query: 300 LFKFVPISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYF---------------LEFQI 344
+ + S+ +A +PG+ L+HAI L PPA Y+ +E QI
Sbjct: 886 IHRLTATSAEIAWVPGNSNLAHAIYLNGEECPPARPSTYWATFCNLRPGTLYQARVEAQI 945
Query: 345 PRQ--WAPMFCELPLRHHQRRKSSSLSLQFNFLGPKLHVSSTQVLSEQKPVVGL 396
P Q W P + +R + + +LQF L L + V +E P G+
Sbjct: 946 PSQGPWEPGW--------ERPEQRAATLQFTTLPAGLPDAPLDVQAEPGPSPGI 991
>CO9_FUGRU (P79755) Complement component C9 precursor
Length = 586
Score = 31.2 bits (69), Expect = 8.4
Identities = 38/167 (22%), Positives = 72/167 (42%), Gaps = 13/167 (7%)
Query: 161 KSVPAQWDPAALSRFIQTYGTHLIVGMAVGGQ-DVICVKQKHSSKIPPGDLRRHLEDLGD 219
KS+P +++ F++ YGTH GG+ +++ V + + K R+ E L
Sbjct: 307 KSLPLEYEKGIYYAFLEDYGTHYTKNGKSGGEYELVYVLNQDTIKAKNLTERKIQECLKI 366
Query: 220 FLFSDVRSPPLLQRKAADGKQKVPEVF--------NRVMQSNTMSFTSISETSSKDGLTI 271
+ ++ + + KA K +V + + N M TS+ S + +T+
Sbjct: 367 GIEAEFATTSVQDGKAHAKLNKCDDVTTKSQGDVEGKAVVDNVM--TSVKGGSLESAVTM 424
Query: 272 ICS-KRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLL-AGIPGS 316
+ G M ++ +W +T+ S P I + PI L+ IPG+
Sbjct: 425 RAKLNKEGVMDIATYQNWARTIASAPALINSEPEPIYMLIPTDIPGA 471
>ARCA_LISIN (Q92FR7) Arginine deiminase (EC 3.5.3.6) (ADI) (Arginine
dihydrolase) (AD)
Length = 408
Score = 31.2 bits (69), Expect = 8.4
Identities = 27/96 (28%), Positives = 41/96 (42%), Gaps = 4/96 (4%)
Query: 318 YLSHAINLYLRYKPPA---EDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNF 374
+L NLY P A + FQ R+ +F EL L+HH R S + +
Sbjct: 148 FLDPLPNLYFTRDPAAVIGSGVTINKMFQPARRRESLFIELILKHHPRFSSQEIPIWSGR 207
Query: 375 LGP-KLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLA 409
P L +L+E+ +VG+ + R +RLA
Sbjct: 208 EEPFPLEGGDELILNEETVLVGVSERTDARAVERLA 243
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,447,880
Number of Sequences: 164201
Number of extensions: 3200253
Number of successful extensions: 7268
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 7255
Number of HSP's gapped (non-prelim): 19
length of query: 617
length of database: 59,974,054
effective HSP length: 116
effective length of query: 501
effective length of database: 40,926,738
effective search space: 20504295738
effective search space used: 20504295738
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0044a.6


