

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
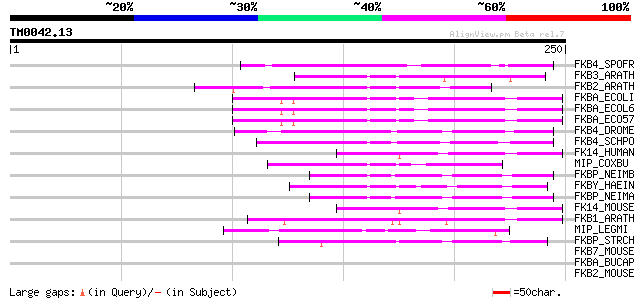
Score E
Sequences producing significant alignments: (bits) Value
FKB4_SPOFR (Q26486) 46 kDa FK506-binding nuclear protein (EC 5.2... 63 8e-10
FKB3_ARATH (Q9SCY2) FKBP-type peptidyl-prolyl cis-trans isomeras... 57 5e-08
FKB2_ARATH (O22870) Probable FKBP-type peptidyl-prolyl cis-trans... 56 9e-08
FKBA_ECOLI (P45523) FKBP-type peptidyl-prolyl cis-trans isomeras... 55 2e-07
FKBA_ECOL6 (P65764) FKBP-type peptidyl-prolyl cis-trans isomeras... 55 2e-07
FKBA_ECO57 (P65765) FKBP-type peptidyl-prolyl cis-trans isomeras... 55 2e-07
FKB4_DROME (P54397) 39 kDa FK506-binding nuclear protein (EC 5.2... 52 1e-06
FKB4_SCHPO (O74191) FK506-binding protein 39 kDa (EC 5.2.1.8) (P... 49 1e-05
FK14_HUMAN (Q9NWM8) FK506 binding protein 14 precursor (EC 5.2.1... 49 1e-05
MIP_COXBU (P51752) Peptidyl-prolyl cis-trans isomerase Mip precu... 48 3e-05
FKBP_NEIMB (P25138) FK506-binding protein (Peptidyl-prolyl cis-t... 48 3e-05
FKBY_HAEIN (P44760) Probable FKBP-type peptidyl-prolyl cis-trans... 47 3e-05
FKBP_NEIMA (P56989) FK506-binding protein (Peptidyl-prolyl cis-t... 47 3e-05
FK14_MOUSE (P59024) FK506 binding protein 14 precursor (EC 5.2.1... 47 4e-05
FKB1_ARATH (Q9LM71) Probable FKBP-type peptidyl-prolyl cis-trans... 46 1e-04
MIP_LEGMI (P31106) Outer membrane protein MIP precursor (Macroph... 45 1e-04
FKBP_STRCH (P28725) FK506-binding protein (Peptidyl-prolyl cis-t... 45 2e-04
FKB7_MOUSE (O54998) FK506-binding protein 7 precursor (EC 5.2.1.... 42 0.001
FKBA_BUCAP (Q8K943) FKBP-type peptidyl-prolyl cis-trans isomeras... 42 0.001
FKB2_MOUSE (P45878) FK506-binding protein 2 precursor (EC 5.2.1.... 42 0.002
>FKB4_SPOFR (Q26486) 46 kDa FK506-binding nuclear protein (EC
5.2.1.8) (Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase)
Length = 412
Score = 62.8 bits (151), Expect = 8e-10
Identities = 45/141 (31%), Positives = 73/141 (50%), Gaps = 13/141 (9%)
Query: 105 KTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGE 164
K + E ++EA VEK+E+ + G+ +LKVG G + G +V++ G+++ +
Sbjct: 284 KKKPEAKKEEA---PVEKKEKKQIAGGVSIEDLKVGSGPVAKAGKVVMVYYEGRLKQNNK 340
Query: 165 VFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADL 224
+F N +G L SK V G + + MK GGKRK++ PP + + GA
Sbjct: 341 MFDNCVKGPGFKFRL------GSKEVISGWDVGIAGMKVGGKRKIVCPPAMAY---GAK- 390
Query: 225 GSGVQIPPLATLEYIVEVDKV 245
GS IPP +TL + V++ V
Sbjct: 391 GSPPVIPPNSTLVFEVDLKNV 411
>FKB3_ARATH (Q9SCY2) FKBP-type peptidyl-prolyl cis-trans isomerase
3, chloroplast precursor (EC 5.2.1.8) (PPIase)
(Rotamase) (AtFKBP13) (FK506 binding protein 1)
Length = 208
Score = 56.6 bits (135), Expect = 5e-08
Identities = 38/116 (32%), Positives = 61/116 (51%), Gaps = 5/116 (4%)
Query: 129 PNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSK 188
P+G+ +C+ VG G G L+ +GK+E+ G+VF +++ K PL +G K
Sbjct: 90 PSGLAFCDKVVGYGPEAVKGQLIKAHYVGKLEN-GKVFDSSYNRGK-PLTFRIGVGEVIK 147
Query: 189 GVCEGI--EYVLKTMKAGGKRKVIVPPQLGFRENGADL-GSGVQIPPLATLEYIVE 241
G +GI + M GGKR + +PP+L + + GA G IPP + L + +E
Sbjct: 148 GWDQGILGSDGIPPMLTGGKRTLRIPPELAYGDRGAGCKGGSCLIPPASVLLFDIE 203
>FKB2_ARATH (O22870) Probable FKBP-type peptidyl-prolyl cis-trans
isomerase 2, chloroplast precursor (EC 5.2.1.8) (PPIase)
(Rotamase)
Length = 223
Score = 55.8 bits (133), Expect = 9e-08
Identities = 44/135 (32%), Positives = 71/135 (52%), Gaps = 10/135 (7%)
Query: 84 GLGAGLAWVGFLAFGV-VSEQIKTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGG 142
G+ GL+ A+ + + K RL ++ E +++E VT +G++Y ++KVG G
Sbjct: 62 GVSGGLSMSSLAAYAAGLPPEDKPRLCEAECE---KELENVPMVTTESGLQYKDIKVGRG 118
Query: 143 ASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMK 202
SP G V + + V S G++F ++ E P +GS KG+ EGI +MK
Sbjct: 119 PSPPVGFQVAANYVAMVPS-GQIFDSSLEKGL-PYLFRVGSGQVIKGLDEGI----LSMK 172
Query: 203 AGGKRKVIVPPQLGF 217
AGGKR++ +P L F
Sbjct: 173 AGGKRRLYIPGPLAF 187
>FKBA_ECOLI (P45523) FKBP-type peptidyl-prolyl cis-trans isomerase
fkpA precursor (EC 5.2.1.8) (PPIase) (Rotamase)
Length = 270
Score = 55.1 bits (131), Expect = 2e-07
Identities = 47/154 (30%), Positives = 76/154 (48%), Gaps = 18/154 (11%)
Query: 101 SEQIKTRLEVSQQEANTRDVE----KEEEV-TLPNGIRYCELKVGGGASPRPGDLVVIDI 155
S Q K + + EA ++ KE+ V T G+ Y ++ G G +P+ D VV++
Sbjct: 112 SAQAKMEKDAADNEAKGKEYREKFAKEKGVKTSSTGLVYQVVEAGKGEAPKDSDTVVVNY 171
Query: 156 MGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQL 215
G + G+ F N++ + PL+ + GV G LK +K GGK K+++PP+L
Sbjct: 172 KGTLID-GKEFDNSYTRGE-PLSFRLD------GVIPGWTEGLKNIKKGGKIKLVIPPEL 223
Query: 216 GFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
+ + G IPP +TL + VE+ V AP
Sbjct: 224 AYGKAGVP-----GIPPNSTLVFDVELLDVKPAP 252
>FKBA_ECOL6 (P65764) FKBP-type peptidyl-prolyl cis-trans isomerase
fkpA precursor (EC 5.2.1.8) (PPIase) (Rotamase)
Length = 270
Score = 55.1 bits (131), Expect = 2e-07
Identities = 47/154 (30%), Positives = 76/154 (48%), Gaps = 18/154 (11%)
Query: 101 SEQIKTRLEVSQQEANTRDVE----KEEEV-TLPNGIRYCELKVGGGASPRPGDLVVIDI 155
S Q K + + EA ++ KE+ V T G+ Y ++ G G +P+ D VV++
Sbjct: 112 SAQAKMEKDAADNEAKGKEYREKFAKEKGVKTSSTGLVYQVVEAGKGEAPKDSDTVVVNY 171
Query: 156 MGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQL 215
G + G+ F N++ + PL+ + GV G LK +K GGK K+++PP+L
Sbjct: 172 KGTLID-GKEFDNSYTRGE-PLSFRLD------GVIPGWTEGLKNIKKGGKIKLVIPPEL 223
Query: 216 GFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
+ + G IPP +TL + VE+ V AP
Sbjct: 224 AYGKAGVP-----GIPPNSTLVFDVELLDVKPAP 252
>FKBA_ECO57 (P65765) FKBP-type peptidyl-prolyl cis-trans isomerase
fkpA precursor (EC 5.2.1.8) (PPIase) (Rotamase)
Length = 270
Score = 55.1 bits (131), Expect = 2e-07
Identities = 47/154 (30%), Positives = 76/154 (48%), Gaps = 18/154 (11%)
Query: 101 SEQIKTRLEVSQQEANTRDVE----KEEEV-TLPNGIRYCELKVGGGASPRPGDLVVIDI 155
S Q K + + EA ++ KE+ V T G+ Y ++ G G +P+ D VV++
Sbjct: 112 SAQAKMEKDAADNEAKGKEYREKFAKEKGVKTSSTGLVYQVVEAGKGEAPKDSDTVVVNY 171
Query: 156 MGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQL 215
G + G+ F N++ + PL+ + GV G LK +K GGK K+++PP+L
Sbjct: 172 KGTLID-GKEFDNSYTRGE-PLSFRLD------GVIPGWTEGLKNIKKGGKIKLVIPPEL 223
Query: 216 GFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
+ + G IPP +TL + VE+ V AP
Sbjct: 224 AYGKAGVP-----GIPPNSTLVFDVELLDVKPAP 252
>FKB4_DROME (P54397) 39 kDa FK506-binding nuclear protein (EC
5.2.1.8) (Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase)
Length = 357
Score = 52.0 bits (123), Expect = 1e-06
Identities = 40/144 (27%), Positives = 66/144 (45%), Gaps = 16/144 (11%)
Query: 102 EQIKTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVES 161
EQ K + QQ A+ ++ T+ G++ + VG G + G V + +G+++S
Sbjct: 229 EQPKAKEPAKQQPAS------KDPRTITGGVKIVDQVVGKGEEAKQGKRVSVYYIGRLQS 282
Query: 162 TGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENG 221
+ F + +G KP +G KG G+ MK GGKR + PP + + G
Sbjct: 283 NNKTFDSLLKG--KPFKFALGGGEVIKGWDVGVA----GMKVGGKRVITCPPHMAYGARG 336
Query: 222 ADLGSGVQIPPLATLEYIVEVDKV 245
A +I P +TL + VE+ V
Sbjct: 337 AP----PKIGPNSTLVFEVELKAV 356
>FKB4_SCHPO (O74191) FK506-binding protein 39 kDa (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase)
Length = 361
Score = 48.5 bits (114), Expect = 1e-05
Identities = 38/134 (28%), Positives = 67/134 (49%), Gaps = 12/134 (8%)
Query: 112 QQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFE 171
+Q+A++ + TL G+ ++K G GAS G V + +GK+E+ G+VF +
Sbjct: 239 KQQASSNAPSSPKTRTLKGGVVVTDVKTGSGASATNGKKVEMRYIGKLEN-GKVFDKNTK 297
Query: 172 GDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIP 231
G KP A ++G +G G+ M+ GG+RK+ +P + + S IP
Sbjct: 298 G--KPFAFILGRGEVIRGWDVGV----AGMQEGGERKITIPAPMAYGNQ-----SIPGIP 346
Query: 232 PLATLEYIVEVDKV 245
+TL + V++ +V
Sbjct: 347 KNSTLVFEVKLVRV 360
>FK14_HUMAN (Q9NWM8) FK506 binding protein 14 precursor (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) (22 kDa FK506-binding protein) (FKBP-22)
(UNQ322/PRO381)
Length = 211
Score = 48.5 bits (114), Expect = 1e-05
Identities = 32/103 (31%), Positives = 56/103 (54%), Gaps = 10/103 (9%)
Query: 148 GDLVVIDIMGKVESTGEVFVNTFEGDK-KPLALVMGSRPYSKGVCEGIEYVLKTMKAGGK 206
GDL+++ G +E G +F +T + + +P+ +G KG +G LK M G K
Sbjct: 45 GDLMLVHYEGYLEKDGSLFHSTHKHNNGQPIWFTLGILEALKGWDQG----LKGMCVGEK 100
Query: 207 RKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
RK+I+PP LG+ + G +IPP +TL + +++ ++ P
Sbjct: 101 RKLIIPPALGYGKEGKG-----KIPPESTLIFNIDLLEIRNGP 138
>MIP_COXBU (P51752) Peptidyl-prolyl cis-trans isomerase Mip
precursor (EC 5.2.1.8) (PPIase) (Rotamase) (Macrophage
infectivity potentiator)
Length = 230
Score = 47.8 bits (112), Expect = 3e-05
Identities = 34/106 (32%), Positives = 54/106 (50%), Gaps = 8/106 (7%)
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
T + K TL NG++Y L+ G G SP D V ++ G++ + G VF ++++ + P
Sbjct: 111 TANKNKPGVKTLANGLQYKVLQAGQGQSPTLNDEVTVNYEGRLIN-GTVFDSSYKRGQ-P 168
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGA 222
+ K V +G + L MK G ++ VPPQL + E GA
Sbjct: 169 ATFPL------KSVIKGWQEALTRMKPGAIWEIYVPPQLAYGEQGA 208
>FKBP_NEIMB (P25138) FK506-binding protein (Peptidyl-prolyl
cis-trans isomerase) (PPIase) (EC 5.2.1.8) (Rotamase)
Length = 109
Score = 47.8 bits (112), Expect = 3e-05
Identities = 37/110 (33%), Positives = 58/110 (52%), Gaps = 10/110 (9%)
Query: 136 ELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIE 195
+L+ G G G + + G +E+ G F ++ + ++PL + +G KG EG
Sbjct: 8 DLQEGFGKEAVKGKEITVHYTGWLEN-GTKFDSSLDR-RQPLTITLGVGQVIKGWDEGFG 65
Query: 196 YVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
MK GGKRK+ +P ++G+ +GA G IPP ATL + VE+ KV
Sbjct: 66 ----GMKEGGKRKLTIPSEMGYGAHGA----GGVIPPHATLIFEVELLKV 107
>FKBY_HAEIN (P44760) Probable FKBP-type peptidyl-prolyl cis-trans
isomerase (EC 5.2.1.8) (PPIase) (Rotamase)
Length = 241
Score = 47.4 bits (111), Expect = 3e-05
Identities = 38/116 (32%), Positives = 60/116 (50%), Gaps = 12/116 (10%)
Query: 127 TLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPY 186
T +G+ Y G G + + D V + GK+ + G+VF ++ E + P+ +
Sbjct: 129 TTQSGLMYKIESAGKGDTIKSTDTVKVHYTGKLPN-GKVFDSSVERGQ-PVEFQLDQ--V 184
Query: 187 SKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG++ V K GGK + ++ P+LG+ E GA G IPP +TL + VEV
Sbjct: 185 IKGWTEGLQLV----KKGGKIQFVIAPELGYGEQGA----GASIPPNSTLIFDVEV 232
>FKBP_NEIMA (P56989) FK506-binding protein (Peptidyl-prolyl
cis-trans isomerase) (PPIase) (EC 5.2.1.8) (Rotamase)
Length = 109
Score = 47.4 bits (111), Expect = 3e-05
Identities = 37/110 (33%), Positives = 57/110 (51%), Gaps = 10/110 (9%)
Query: 136 ELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIE 195
+L+ G G G + + G +E+ G F ++ + ++PL + +G KG EG
Sbjct: 8 DLQEGFGKEAVKGKEITVHYTGWLEN-GTKFDSSLDR-RQPLTITLGVGQVIKGWDEGFG 65
Query: 196 YVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
MK GGKRK+ +P ++G+ GA G IPP ATL + VE+ KV
Sbjct: 66 ----GMKEGGKRKLTIPSEMGYGARGA----GGVIPPHATLIFEVELLKV 107
>FK14_MOUSE (P59024) FK506 binding protein 14 precursor (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase)
Length = 211
Score = 47.0 bits (110), Expect = 4e-05
Identities = 32/103 (31%), Positives = 55/103 (53%), Gaps = 10/103 (9%)
Query: 148 GDLVVIDIMGKVESTGEVFVNTFEGDK-KPLALVMGSRPYSKGVCEGIEYVLKTMKAGGK 206
GDL+++ G +E G +F +T + + +P+ +G KG +G LK M G K
Sbjct: 45 GDLMLVHYEGYLEKDGSLFHSTHKHNNGQPVWFTLGILEVLKGWDQG----LKGMCVGEK 100
Query: 207 RKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
RK+ VPP LG+ + G +IPP +TL + +++ ++ P
Sbjct: 101 RKLTVPPALGYGKEGKG-----KIPPESTLIFNIDLLEIRNGP 138
>FKB1_ARATH (Q9LM71) Probable FKBP-type peptidyl-prolyl cis-trans
isomerase 1, chloroplast precursor (EC 5.2.1.8) (PPIase)
(Rotamase)
Length = 232
Score = 45.8 bits (107), Expect = 1e-04
Identities = 43/171 (25%), Positives = 72/171 (41%), Gaps = 34/171 (19%)
Query: 108 LEVSQQEANTRDVEK----EEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTG 163
L S EA R K EE T P G+++ +++ G G G + + S
Sbjct: 64 LTPSSSEARERRSRKVIPLEEYSTGPEGLKFYDIEEGKGPVATEGSTAQVHFDCRYRSIT 123
Query: 164 EVFVNTFE---GDK---KPLALVMGSRPYSKGVCEGIE-------------------YVL 198
+ + G++ +P +GS P + E ++ ++
Sbjct: 124 AISTRESKLLAGNRSIAQPYEFKVGSTPGKERKREFVDNPNGLFSAQAAPKPPPAMYFIT 183
Query: 199 KTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
+ MK GGKR VIVPP+ G+ + G + +IPP AT E +E+ +V+ P
Sbjct: 184 EGMKVGGKRTVIVPPEAGYGQKGMN-----EIPPGATFELNIELLRVTPPP 229
>MIP_LEGMI (P31106) Outer membrane protein MIP precursor (Macrophage
infectivity potentiator) (Peptidyl-prolyl cis-trans
isomerase) (EC 5.2.1.8) (PPIase) (Rotamase)
Length = 243
Score = 45.4 bits (106), Expect = 1e-04
Identities = 40/131 (30%), Positives = 62/131 (46%), Gaps = 14/131 (10%)
Query: 97 FGVVSEQIKTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIM 156
F +E+ K++ E E + KE V+LP+G++Y L+ G GA P D+V ++
Sbjct: 106 FNKKAEENKSKGEAFLNENKS----KEGVVSLPSGLQYNILERGDGAKPTKDDVVTVEYT 161
Query: 157 GKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLG 216
GK+ G+VF +T E KP + V G L+ M AG +V +P L
Sbjct: 162 GKL-IDGQVFDST-EKTGKPATFKVSQ------VIPGWTEALQLMPAGSTWEVYIPSNLA 213
Query: 217 F--RENGADLG 225
+ R G +G
Sbjct: 214 YGPRSVGGPIG 224
>FKBP_STRCH (P28725) FK506-binding protein (Peptidyl-prolyl
cis-trans isomerase) (PPIase) (EC 5.2.1.8) (Rotamase)
Length = 124
Score = 45.1 bits (105), Expect = 2e-04
Identities = 35/126 (27%), Positives = 60/126 (46%), Gaps = 13/126 (10%)
Query: 122 KEEEVTLPNGIRYCELKV-----GGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
++ EV P G +L + G G + G V + +G STGE F ++ P
Sbjct: 4 EKPEVDFPGGEPPADLAIKDIWEGDGPVAQAGQTVSVHYVGVAFSTGEEFDASWNRGT-P 62
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATL 236
L +G+ G +G++ MK GG+R++I+P L + + GA G +I P TL
Sbjct: 63 LQFQLGAGQVISGWDQGVQ----GMKVGGRRELIIPAHLAYGDRGA---GGGKIAPGETL 115
Query: 237 EYIVEV 242
++ ++
Sbjct: 116 IFVCDL 121
>FKB7_MOUSE (O54998) FK506-binding protein 7 precursor (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) (FKBP-23)
Length = 218
Score = 42.4 bits (98), Expect = 0.001
Identities = 34/107 (31%), Positives = 49/107 (45%), Gaps = 12/107 (11%)
Query: 146 RPGDLVVIDIMGKVESTGEVFV---NTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMK 202
R GDL+ G + G F EG K L +G V +G++ + M
Sbjct: 47 RKGDLLNAHYDGYLAKDGSKFYCSRTQDEGHPKWFVLGVGH------VIKGLDIAMMDMC 100
Query: 203 AGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
G KRKVI+PP + + G G +IPP ATL + +E+ V+ P
Sbjct: 101 PGEKRKVIIPPSFAYGKEGYAEG---KIPPNATLMFEIELYAVTKGP 144
>FKBA_BUCAP (Q8K943) FKBP-type peptidyl-prolyl cis-trans isomerase
fkpA (EC 5.2.1.8) (PPIase) (Rotamase)
Length = 252
Score = 42.0 bits (97), Expect = 0.001
Identities = 46/163 (28%), Positives = 76/163 (46%), Gaps = 32/163 (19%)
Query: 101 SEQIKT---RLEVSQQEANTRDVEKEEEVTLPNGIRYCEL--KVGGGASPRPGDLVVIDI 155
+E+I T +LE + + A + EK E+ L G Y + ++ G + G L +ID
Sbjct: 95 NEEISTILQKLEKNLKNAAKIEFEKSEKENLIQGKLYMKKFSEMKGVSKTSSGLLYIIDK 154
Query: 156 MGKVES----TGEV-------FVNTFEGDK-----KPLALVMGSRPYSKGVCEGIEYVLK 199
+G+ E E+ +N E D KP+ L++ K V G + LK
Sbjct: 155 LGEGEEIKTKNAEITVHYKGSLINGTEFDSSYKRGKPITLML------KDVILGWQEGLK 208
Query: 200 TMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
+K GGK K+I+PP LG+ N + +IP + L + +E+
Sbjct: 209 YIKKGGKIKLIIPPNLGYGSNRIN-----EIPANSILIFDIEL 246
>FKB2_MOUSE (P45878) FK506-binding protein 2 precursor (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase)
(Rotamase) (13 kDa FKBP) (FKBP-13)
Length = 140
Score = 41.6 bits (96), Expect = 0.002
Identities = 35/125 (28%), Positives = 63/125 (50%), Gaps = 16/125 (12%)
Query: 121 EKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALV 180
+++ ++ + + +C +K R GD++ + GK+E G F ++ +P
Sbjct: 26 KRKLQIGVKKRVDHCPIK------SRKGDVLHMHYTGKLED-GTEFDSSLP-QNQPFVFS 77
Query: 181 MGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV 240
+G+ KG +G L M G KRK+++P +LG+ E GA +IP ATL + V
Sbjct: 78 LGTGQVIKGWDQG----LLGMCEGEKRKLVIPSELGYGERGAP----PKIPGGATLVFEV 129
Query: 241 EVDKV 245
E+ K+
Sbjct: 130 ELLKI 134
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,906,729
Number of Sequences: 164201
Number of extensions: 1372330
Number of successful extensions: 8238
Number of sequences better than 10.0: 214
Number of HSP's better than 10.0 without gapping: 102
Number of HSP's successfully gapped in prelim test: 120
Number of HSP's that attempted gapping in prelim test: 7626
Number of HSP's gapped (non-prelim): 591
length of query: 250
length of database: 59,974,054
effective HSP length: 108
effective length of query: 142
effective length of database: 42,240,346
effective search space: 5998129132
effective search space used: 5998129132
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0042.13


