

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0037a.10
(168 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
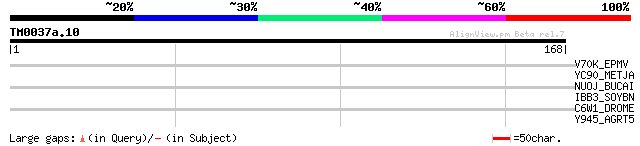
Score E
Sequences producing significant alignments: (bits) Value
V70K_EPMV (P20129) 70 kDa protein 29 4.4
YC90_METJA (Q58686) Hypothetical protein MJ1290 29 5.8
NUOJ_BUCAI (P57260) NADH-quinone oxidoreductase chain J (EC 1.6.... 28 7.6
IBB3_SOYBN (P01064) Bowman-Birk type proteinase inhibitor D-II p... 28 7.6
C6W1_DROME (Q9V9L1) Probable cytochrome P450 6w1 (EC 1.14.-.-) (... 28 7.6
Y945_AGRT5 (Q8UGU0) Hypothetical UPF0061 protein Atu0945/AGR_C_1725 28 9.9
>V70K_EPMV (P20129) 70 kDa protein
Length = 649
Score = 29.3 bits (64), Expect = 4.4
Identities = 18/57 (31%), Positives = 29/57 (50%), Gaps = 9/57 (15%)
Query: 70 SNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFYPCF 126
S+ L + DS C + LPPS +P++HS +SF CT+ + +PC+
Sbjct: 548 SSLSLRTFLDSAVISCDSSPVLPPSPSPSSHSSSSFQ-SCTSPPR--------FPCY 595
>YC90_METJA (Q58686) Hypothetical protein MJ1290
Length = 312
Score = 28.9 bits (63), Expect = 5.8
Identities = 18/62 (29%), Positives = 30/62 (48%), Gaps = 5/62 (8%)
Query: 36 SNSEEPYLFE--GLLEIRKFYSLQGMNWFLLKYQGCSNFDLFIYNDSIEEVCYLTRSLPP 93
+ + PY + G++ K Y G N+ L Y+ SN +F YN S++++ Y P
Sbjct: 74 NGTNNPYHIKSAGIILYEKIY---GYNYSNLLYRNSSNSLIFYYNFSVDKINYTINITIP 130
Query: 94 SI 95
I
Sbjct: 131 QI 132
>NUOJ_BUCAI (P57260) NADH-quinone oxidoreductase chain J (EC
1.6.99.5) (NADH dehydrogenase I, chain J) (NDH-1, chain
J)
Length = 170
Score = 28.5 bits (62), Expect = 7.6
Identities = 15/50 (30%), Positives = 26/50 (52%)
Query: 119 EWIFYPCFFTTRLSPNQVSSSQLVMNQFCYVLMTNESVYGIFVSLNAELA 168
E++FY C F +S V + + Y++++ S+ G+F SL A A
Sbjct: 2 EFVFYVCSFAAVVSTFFVIIQKNAVYSLVYLIISLLSIAGVFFSLGAFFA 51
>IBB3_SOYBN (P01064) Bowman-Birk type proteinase inhibitor D-II
precursor (IV)
Length = 83
Score = 28.5 bits (62), Expect = 7.6
Identities = 23/85 (27%), Positives = 32/85 (37%), Gaps = 16/85 (18%)
Query: 64 LKYQGCSNFDLFIYNDSIEEVCYLTRSLPPSITPTTHSHTSFHGECTTGSKSVTEEWIFY 123
LK S++D Y+ ++C TRS+PP + S H +C +
Sbjct: 7 LKSDQSSSYDDDEYSKPCCDLCMCTRSMPPQCSCEDIRLNSCHSDCKS------------ 54
Query: 124 PCFFTTRLSPNQVSSSQLVMNQFCY 148
TR P Q L N FCY
Sbjct: 55 --CMCTRSQPGQCRC--LDTNDFCY 75
>C6W1_DROME (Q9V9L1) Probable cytochrome P450 6w1 (EC 1.14.-.-)
(CYPVIW1)
Length = 514
Score = 28.5 bits (62), Expect = 7.6
Identities = 15/41 (36%), Positives = 23/41 (55%)
Query: 18 LQWKLVDPFGSVHLVSYESNSEEPYLFEGLLEIRKFYSLQG 58
L+ ++VD FG +SYE E PYL + + E + Y + G
Sbjct: 331 LRQEIVDFFGDEDHISYERIQEMPYLSQVVNETLRKYPIVG 371
>Y945_AGRT5 (Q8UGU0) Hypothetical UPF0061 protein Atu0945/AGR_C_1725
Length = 487
Score = 28.1 bits (61), Expect = 9.9
Identities = 16/44 (36%), Positives = 22/44 (49%), Gaps = 1/44 (2%)
Query: 106 HGECTTGSKSVTEEWI-FYPCFFTTRLSPNQVSSSQLVMNQFCY 148
HG T + +V+ E I F PC F PN+V SS ++ Y
Sbjct: 241 HGVMNTDNMAVSGETIDFGPCAFLDEYHPNKVFSSIDAQGRYAY 284
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.136 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,450,146
Number of Sequences: 164201
Number of extensions: 759144
Number of successful extensions: 1590
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1588
Number of HSP's gapped (non-prelim): 6
length of query: 168
length of database: 59,974,054
effective HSP length: 102
effective length of query: 66
effective length of database: 43,225,552
effective search space: 2852886432
effective search space used: 2852886432
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0037a.10


