

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
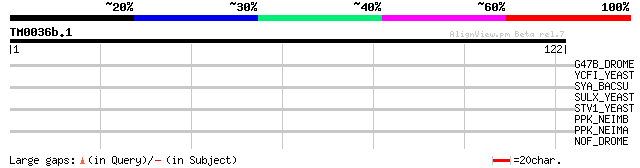
Query= TM0036b.1
(122 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
G47B_DROME (P58961) Putative gustatory receptor 47b 30 0.73
YCFI_YEAST (P39109) Metal resistance protein YCF1 (Yeast cadmium... 30 0.96
SYA_BACSU (O34526) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 30 1.3
SULX_YEAST (P53394) Putative sulfate transporter YPR003C 28 2.8
STV1_YEAST (P37296) Vacuolar ATP synthase 101 kDa subunit (V-ATP... 28 2.8
PPK_NEIMB (Q9JXS9) Polyphosphate kinase (EC 2.7.4.1) (Polyphosph... 27 8.1
PPK_NEIMA (Q9JW42) Polyphosphate kinase (EC 2.7.4.1) (Polyphosph... 27 8.1
NOF_DROME (P16320) 120.7 kDa protein in NOF-FB transposable element 27 8.1
>G47B_DROME (P58961) Putative gustatory receptor 47b
Length = 414
Score = 30.4 bits (67), Expect = 0.73
Identities = 25/92 (27%), Positives = 40/92 (43%), Gaps = 7/92 (7%)
Query: 8 YEPNDSLRSDRFIVSLKDHSSTCNF--SELLEILSRHVVAGIFYKGLNI-----VLCILI 60
Y+ +DS I L + S C+F +EL + AGIFY + V+C +
Sbjct: 218 YQADDSSEVHERIAYLFEMSKRCSFLLAELNGVFGFAAAAGIFYDFTIMTCFVYVICQKL 277
Query: 61 LQKDKLIRTHTCTLLHQLMGKSKVFDANCYSY 92
L+++ + LLH + KV + Y Y
Sbjct: 278 LEREPWDPEYVYMLLHVAIHTYKVVITSTYGY 309
>YCFI_YEAST (P39109) Metal resistance protein YCF1 (Yeast cadmium
factor 1)
Length = 1515
Score = 30.0 bits (66), Expect = 0.96
Identities = 19/70 (27%), Positives = 33/70 (47%), Gaps = 8/70 (11%)
Query: 5 LDWYEPNDSLRSDRFIVSLKDHSSTCNFSELLEILSRHVVAGIFYKG--------LNIVL 56
L W E + S+ ++ ++ + NF++L+ IL RH GI+Y G ++
Sbjct: 120 LHWIEYDRSVVANTVLLFYWLFETFGNFAKLINILIRHTYEGIWYSGQTGFILTLFQVIT 179
Query: 57 CILILQKDKL 66
C IL + L
Sbjct: 180 CASILLLEAL 189
>SYA_BACSU (O34526) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 878
Score = 29.6 bits (65), Expect = 1.3
Identities = 21/67 (31%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Query: 14 LRSDRFIVSLKDHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILILQKDKLIRTHTCT 73
LRS++ +V +KD N + E + V +G KGL++ + + +I+ HT T
Sbjct: 513 LRSEQAVVRIKDVQKAPNGQHVHEGV---VESGTVQKGLHVTAEVEDHMRSGVIKNHTAT 569
Query: 74 -LLHQLM 79
LLHQ +
Sbjct: 570 HLLHQAL 576
>SULX_YEAST (P53394) Putative sulfate transporter YPR003C
Length = 754
Score = 28.5 bits (62), Expect = 2.8
Identities = 21/62 (33%), Positives = 32/62 (50%), Gaps = 6/62 (9%)
Query: 14 LRSDRFIVSLKDHSSTCNFSELLEIL-----SRHVVAGIFYKGLNIVLCILILQKDKLIR 68
L+ D+F+VSL H T F ++L ++ H+ IF IVL + L K KL++
Sbjct: 255 LKLDKFLVSLPQHYHT-PFEKILFLIDYAPAQYHIPTAIFSGCCLIVLFLTRLLKRKLMK 313
Query: 69 TH 70
H
Sbjct: 314 YH 315
>STV1_YEAST (P37296) Vacuolar ATP synthase 101 kDa subunit (V-ATPase
subunit AC115)
Length = 890
Score = 28.5 bits (62), Expect = 2.8
Identities = 16/57 (28%), Positives = 26/57 (45%), Gaps = 1/57 (1%)
Query: 44 VAGIFYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSL 100
V +F + +CIL+ + H L H + SK F+ Y+Y P FR++
Sbjct: 833 VVFLFAMWFVLTVCILVFMEGTSAMLHALRL-HWVEAMSKFFEGEGYAYEPFSFRAI 888
>PPK_NEIMB (Q9JXS9) Polyphosphate kinase (EC 2.7.4.1)
(Polyphosphoric acid kinase) (ATP-polyphosphate
phosphotransferase)
Length = 685
Score = 26.9 bits (58), Expect = 8.1
Identities = 11/46 (23%), Positives = 28/46 (59%)
Query: 25 DHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILILQKDKLIRTH 70
D ++ N+++ LE HVV G+F ++ + ++I ++D +++ +
Sbjct: 407 DEANNVNWAKQLEEAGAHVVYGVFGYKVHAKMALVIRREDGVLKRY 452
>PPK_NEIMA (Q9JW42) Polyphosphate kinase (EC 2.7.4.1)
(Polyphosphoric acid kinase) (ATP-polyphosphate
phosphotransferase)
Length = 685
Score = 26.9 bits (58), Expect = 8.1
Identities = 11/46 (23%), Positives = 28/46 (59%)
Query: 25 DHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILILQKDKLIRTH 70
D ++ N+++ LE HVV G+F ++ + ++I ++D +++ +
Sbjct: 407 DEANNVNWAKQLEEAGAHVVYGVFGYKVHAKMALVIRREDGVLKRY 452
>NOF_DROME (P16320) 120.7 kDa protein in NOF-FB transposable element
Length = 1056
Score = 26.9 bits (58), Expect = 8.1
Identities = 12/30 (40%), Positives = 16/30 (53%)
Query: 23 LKDHSSTCNFSELLEILSRHVVAGIFYKGL 52
L +TC F L+EILS + +YK L
Sbjct: 790 LLQFKNTCPFDSLVEILSTAYIDNFYYKSL 819
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,964,629
Number of Sequences: 164201
Number of extensions: 481209
Number of successful extensions: 1185
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1181
Number of HSP's gapped (non-prelim): 8
length of query: 122
length of database: 59,974,054
effective HSP length: 98
effective length of query: 24
effective length of database: 43,882,356
effective search space: 1053176544
effective search space used: 1053176544
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0036b.1


