

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
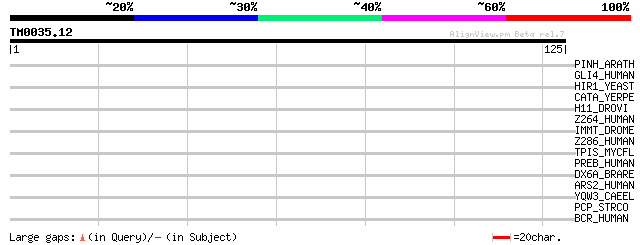
Query= TM0035.12
(125 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
PINH_ARATH (Q9XGW1) PINHEAD protein (ZWILLE protein) 29 1.6
GLI4_HUMAN (P10075) Zinc finger protein GLI4 (Krueppel-related z... 29 2.1
HIR1_YEAST (P32479) Histone transcription regulator 1 28 2.8
CATA_YERPE (Q9X6B0) Peroxidase/catalase (EC 1.11.1.6) (Catalase-... 28 2.8
H11_DROVI (Q24704) Histone H1.1 28 3.6
Z264_HUMAN (O43296) Zinc finger protein 264 28 4.7
IMMT_DROME (P91928) Putative mitochondrial inner membrane protein 28 4.7
Z286_HUMAN (Q9HBT8) Zinc finger protein 286 27 6.2
TPIS_MYCFL (P48779) Triosephosphate isomerase (EC 5.3.1.1) (TIM)... 27 6.2
PREB_HUMAN (Q9HCU5) Prolactin regulatory element-binding protein... 27 6.2
DX6A_BRARE (Q98877) Homeobox protein Dlx6a (DLX-6) 27 6.2
ARS2_HUMAN (Q9BXP5) Arsenite-resistance protein 2 27 6.2
YQW3_CAEEL (Q09556) Hypothetical protein F33H1.3 from chromosome II 27 8.0
PCP_STRCO (Q9RL48) Pyrrolidone-carboxylate peptidase (EC 3.4.19.... 27 8.0
BCR_HUMAN (P11274) Breakpoint cluster region protein (EC 2.7.1.-) 27 8.0
>PINH_ARATH (Q9XGW1) PINHEAD protein (ZWILLE protein)
Length = 988
Score = 29.3 bits (64), Expect = 1.6
Identities = 17/44 (38%), Positives = 24/44 (53%), Gaps = 1/44 (2%)
Query: 31 VLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIK 74
++A + L+K +DL RR A DG KSL T E W + +K
Sbjct: 177 IIAELVRLYKESDLGRRLPAYDGRKSL-YTAGELPFTWKEFSVK 219
>GLI4_HUMAN (P10075) Zinc finger protein GLI4 (Krueppel-related zinc
finger protein 4) (HKR4 protein)
Length = 376
Score = 28.9 bits (63), Expect = 2.1
Identities = 17/64 (26%), Positives = 30/64 (46%)
Query: 11 SHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNK 70
SH F H+ + P++ + +S++L R +R G+K +Q KA IW+
Sbjct: 249 SHSSHFTQHLRIHNGEKPYKCGECGQAFSQSSNLVRHQRLHTGEKPYACSQCGKAFIWSS 308
Query: 71 TQIK 74
I+
Sbjct: 309 VLIE 312
>HIR1_YEAST (P32479) Histone transcription regulator 1
Length = 788
Score = 28.5 bits (62), Expect = 2.8
Identities = 15/59 (25%), Positives = 31/59 (52%)
Query: 12 HRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNK 70
+R ++ T+++ P EV F P LF+ N ++K+ D + +L ++K ++K
Sbjct: 212 NRGTWDTNVSLIGHDAPTEVARFNPRLFERNAGVKQKKDDDPENALVGQNDDKVHHFDK 270
>CATA_YERPE (Q9X6B0) Peroxidase/catalase (EC 1.11.1.6)
(Catalase-peroxidase) (Antigen 5)
Length = 737
Score = 28.5 bits (62), Expect = 2.8
Identities = 30/109 (27%), Positives = 44/109 (39%), Gaps = 16/109 (14%)
Query: 2 IHNPGQHHHSHRRSFF--THINWAP------DQHPHEVLAFMPSLFKSNDLARRKRASDG 53
+ NP + + R++F TH + P D+ P E + L +N K
Sbjct: 392 LENPEEFKMAFARAWFKLTHRDMGPAARYLGDEVPKETFIWQDPLPAAN----YKMIDSA 447
Query: 54 DKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGAAVR 102
D S K + K + + IKT ASTF T R G +GA +R
Sbjct: 448 DISELKDKILKTGLSDTKLIKTAWASASTFRGTDFRGGD----NGARIR 492
>H11_DROVI (Q24704) Histone H1.1
Length = 250
Score = 28.1 bits (61), Expect = 3.6
Identities = 23/70 (32%), Positives = 36/70 (50%), Gaps = 5/70 (7%)
Query: 56 SLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFR--SGAAVRFVILGFAERRE 113
S NK + KAS K KT AS ATK +S T + +GAA + + A ++
Sbjct: 119 SANKDAKPKASAVEK---KTKKVNASAARATKSKSSTSTTKKAAGAADKKLSKSAAAKKN 175
Query: 114 MEEEEKRKQK 123
+E+++ K+K
Sbjct: 176 VEKKKADKEK 185
>Z264_HUMAN (O43296) Zinc finger protein 264
Length = 627
Score = 27.7 bits (60), Expect = 4.7
Identities = 14/64 (21%), Positives = 28/64 (42%)
Query: 11 SHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNK 70
S+R+ H + + P+E + + + + L R KR G+K + K+ W+
Sbjct: 465 SNRKDLIRHFSIHTGEKPYECVECGKAFTRMSGLTRHKRIHSGEKPYECVECGKSFCWST 524
Query: 71 TQIK 74
I+
Sbjct: 525 NLIR 528
>IMMT_DROME (P91928) Putative mitochondrial inner membrane protein
Length = 841
Score = 27.7 bits (60), Expect = 4.7
Identities = 24/98 (24%), Positives = 45/98 (45%), Gaps = 13/98 (13%)
Query: 35 MPSLFKS------NDLARRKRASDGDKSLNKTQEEKASIW------NKTQIKTPLNKAST 82
+P+LF S +D R RAS+ ++ + TQ+ A W K Q ++ +
Sbjct: 2 LPTLFPSKHKPKLSDQIRVARASETKRTTSSTQDSCACAWECGEWKRKQQHWPAIHPPAL 61
Query: 83 FEATKLRSGTCIFRSGAA-VRFVILGFAERREMEEEEK 119
++++LR+ IF S ++F + A R ++ K
Sbjct: 62 TQSSQLRTNWAIFNSRRRDLKFTVAEGAGARAEQDHSK 99
>Z286_HUMAN (Q9HBT8) Zinc finger protein 286
Length = 521
Score = 27.3 bits (59), Expect = 6.2
Identities = 16/72 (22%), Positives = 32/72 (44%), Gaps = 3/72 (4%)
Query: 2 IHNPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGD---KSLN 58
+ N + H R FTH+N ++ H+ + S + + L R G +L
Sbjct: 162 LENQQGNQERHLREMFTHMNSLSEETDHKHDVYWKSFNQKSVLITEDRVPKGSYAFHTLE 221
Query: 59 KTQEEKASIWNK 70
K+ ++K+++ K
Sbjct: 222 KSLKQKSNLMKK 233
>TPIS_MYCFL (P48779) Triosephosphate isomerase (EC 5.3.1.1) (TIM)
(Triose-phosphate isomerase)
Length = 242
Score = 27.3 bits (59), Expect = 6.2
Identities = 25/100 (25%), Positives = 43/100 (43%), Gaps = 3/100 (3%)
Query: 21 NWAPDQHPHEVLAFMPS--LFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLN 78
NW ++ HE F+ +F S +L + K D + + T S ++K
Sbjct: 8 NWKMNKTVHETRDFIQKFDIFYSKNLGKIKDNLDFAIAPSFTSLALISTSKIDKLKVAAQ 67
Query: 79 KASTFEATKLRSG-TCIFRSGAAVRFVILGFAERREMEEE 117
S F++ + AV +VI+G +ERRE+ +E
Sbjct: 68 NLSQFDSGAFTGEISAKMLQDLAVHYVIIGHSERREIFKE 107
>PREB_HUMAN (Q9HCU5) Prolactin regulatory element-binding protein
(Mammalian guanine nucleotide exchange factor mSec12)
Length = 417
Score = 27.3 bits (59), Expect = 6.2
Identities = 23/69 (33%), Positives = 32/69 (46%), Gaps = 7/69 (10%)
Query: 30 EVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLN---KASTFEAT 86
+VL F + DLA DG K + ++ KAS+W K Q+ T L+ TF +T
Sbjct: 188 KVLEFKAHEGEIEDLAL---GPDG-KLVTVGRDLKASVWQKDQLVTQLHWQENGPTFSST 243
Query: 87 KLRSGTCIF 95
R C F
Sbjct: 244 PYRYQACRF 252
>DX6A_BRARE (Q98877) Homeobox protein Dlx6a (DLX-6)
Length = 247
Score = 27.3 bits (59), Expect = 6.2
Identities = 20/80 (25%), Positives = 37/80 (46%), Gaps = 16/80 (20%)
Query: 2 IHNPGQHHHSHRRSFFTHIN-------WAPDQHPHEVLAFMPSLFKSNDLARRKRASDGD 54
+H+ HHH H S ++ N ++ HPH ++PS + SN AS
Sbjct: 47 LHSGSHHHHQHDSSPYSGSNSYNRSLPYSYVSHPHH-SPYLPS-YHSN-------ASGTQ 97
Query: 55 KSLNKTQEEKASIWNKTQIK 74
L+ T+++K ++ +I+
Sbjct: 98 TRLDATEQQKTTVIENGEIR 117
>ARS2_HUMAN (Q9BXP5) Arsenite-resistance protein 2
Length = 876
Score = 27.3 bits (59), Expect = 6.2
Identities = 22/83 (26%), Positives = 32/83 (38%), Gaps = 7/83 (8%)
Query: 41 SNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGAA 100
SND +K DGDK K EK + + + + +F+ + SG A
Sbjct: 320 SNDDKTKKSEGDGDKEEKKEDSEKEAKKSSKKRNRKHSGDDSFDEGSVSESESESESGQA 379
Query: 101 VRFVILGFAERREMEEEEKRKQK 123
E+ E EE K K+K
Sbjct: 380 EE-------EKEEAEEALKEKEK 395
>YQW3_CAEEL (Q09556) Hypothetical protein F33H1.3 from chromosome II
Length = 359
Score = 26.9 bits (58), Expect = 8.0
Identities = 17/57 (29%), Positives = 26/57 (44%), Gaps = 2/57 (3%)
Query: 8 HHHSHRRSFFTHINWAPDQHPHE--VLAFMPSLFKSNDLARRKRASDGDKSLNKTQE 62
HHH H +H N Q+ HE V++ P + +SN+ A S + N +E
Sbjct: 248 HHHHHHPHASSHYNPMGFQNHHEEAVISSAPQINRSNEPAAPATLSAAPELRNLRRE 304
>PCP_STRCO (Q9RL48) Pyrrolidone-carboxylate peptidase (EC 3.4.19.3)
(5-oxoprolyl-peptidase) (Pyroglutamyl-peptidase I)
(PGP-I) (Pyrase)
Length = 216
Score = 26.9 bits (58), Expect = 8.0
Identities = 10/25 (40%), Positives = 13/25 (52%)
Query: 12 HRRSFFTHINWAPDQHPHEVLAFMP 36
H R F H+ WAP+Q P +P
Sbjct: 160 HLRGGFAHVPWAPEQVPDGTAPALP 184
>BCR_HUMAN (P11274) Breakpoint cluster region protein (EC 2.7.1.-)
Length = 1271
Score = 26.9 bits (58), Expect = 8.0
Identities = 20/83 (24%), Positives = 37/83 (44%), Gaps = 11/83 (13%)
Query: 42 NDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGAAV 101
+D+ R KRA+ G K+ T+ K + + + ++ + F R+G +
Sbjct: 796 SDIQREKRANKGSKA---TERLKKKLSEQESLLLLMSPSMAFRVHS--------RNGKSY 844
Query: 102 RFVILGFAERREMEEEEKRKQKQ 124
F+I ER E E + +QK+
Sbjct: 845 TFLISSDYERAEWRENIREQQKK 867
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.129 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,407,622
Number of Sequences: 164201
Number of extensions: 493154
Number of successful extensions: 1854
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1846
Number of HSP's gapped (non-prelim): 17
length of query: 125
length of database: 59,974,054
effective HSP length: 101
effective length of query: 24
effective length of database: 43,389,753
effective search space: 1041354072
effective search space used: 1041354072
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0035.12


