

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.10
(150 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
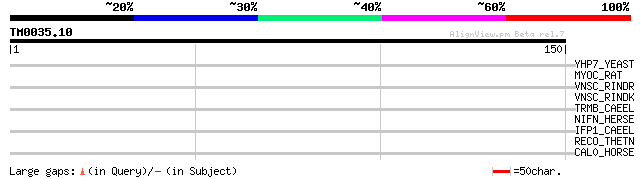
Score E
Sequences producing significant alignments: (bits) Value
YHP7_YEAST (P38809) Hypothetical 40.7 kDa protein in HXT5-NRK1 i... 32 0.68
MYOC_RAT (Q9R1J4) Myocilin precursor (Trabecular meshwork-induce... 30 2.6
VNSC_RINDR (Q03339) Nonstructural protein C 28 5.8
VNSC_RINDK (P35948) Nonstructural protein C 28 5.8
TRMB_CAEEL (Q23126) Probable tRNA (guanine-N(7)-)-methyltransfer... 28 7.5
NIFN_HERSE (O87627) Nitrogenase iron-molybdenum cofactor biosynt... 28 7.5
IFP1_CAEEL (Q09501) Intermediate filament protein ifp-1 (Interme... 28 7.5
RECO_THETN (Q8RB48) DNA repair protein recO (Recombination prote... 28 9.8
CAL0_HORSE (Q9N0V5) Calcitonin precursor 28 9.8
>YHP7_YEAST (P38809) Hypothetical 40.7 kDa protein in HXT5-NRK1
intergenic region
Length = 366
Score = 31.6 bits (70), Expect = 0.68
Identities = 15/57 (26%), Positives = 25/57 (43%), Gaps = 5/57 (8%)
Query: 35 KKNTTQHNDRDSSELRDQVSIHPSSS-----SFIITMLLGPSLHQCGWMDGCMPTRN 86
++ + H D+D + + + + P + +T L G S H G D C P RN
Sbjct: 104 RRTSHGHRDKDKQKSKSRTKVKPPKNVDTIDKMDVTGLFGGSFHHDGPFDACTPQRN 160
>MYOC_RAT (Q9R1J4) Myocilin precursor (Trabecular meshwork-induced
glucocorticoid response protein)
Length = 502
Score = 29.6 bits (65), Expect = 2.6
Identities = 20/58 (34%), Positives = 24/58 (40%), Gaps = 2/58 (3%)
Query: 70 PSLHQCGWMDGCMPTRNSYDTSGSLSHADTRATSAKATCLLSWFLLHQSLSRSLTYPQ 127
PS C D M S+ HAD +T A+ L S LLHQ S +T Q
Sbjct: 55 PSESSCPREDQAMSAIQDLQRDSSIQHADLESTKARVRSLES--LLHQMTSGGVTGTQ 110
>VNSC_RINDR (Q03339) Nonstructural protein C
Length = 177
Score = 28.5 bits (62), Expect = 5.8
Identities = 19/55 (34%), Positives = 22/55 (39%), Gaps = 7/55 (12%)
Query: 81 CMPTRNSYDTSGSLSHADTRATSAKATCLLSWF-------LLHQSLSRSLTYPQC 128
C PTR S SHA + AKA CL + +S SL PQC
Sbjct: 41 CPPTRPKQTIRISASHASQQLDQAKAACLAVTIKDLEEATAVMRSWEHSLVTPQC 95
>VNSC_RINDK (P35948) Nonstructural protein C
Length = 177
Score = 28.5 bits (62), Expect = 5.8
Identities = 19/55 (34%), Positives = 22/55 (39%), Gaps = 7/55 (12%)
Query: 81 CMPTRNSYDTSGSLSHADTRATSAKATCLLSWF-------LLHQSLSRSLTYPQC 128
C PTR S SHA + AKA CL + +S SL PQC
Sbjct: 41 CPPTRPKQTIRISASHASQQLDQAKAACLAVTIRDLEEATAVMRSWEHSLVTPQC 95
>TRMB_CAEEL (Q23126) Probable tRNA
(guanine-N(7)-)-methyltransferase (EC 2.1.1.33)
(tRNA(m7G46)-methyltransferase)
Length = 256
Score = 28.1 bits (61), Expect = 7.5
Identities = 12/42 (28%), Positives = 25/42 (58%), Gaps = 1/42 (2%)
Query: 4 LVPSLTRMKSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDRD 45
+ P+LT M+ +H+P C + Q+K +++ ++ H+D D
Sbjct: 3 ITPALTEMEL-DHKPTCETVPGLPQKKHYRQRAHSNPHSDHD 43
>NIFN_HERSE (O87627) Nitrogenase iron-molybdenum cofactor
biosynthesis protein nifN
Length = 443
Score = 28.1 bits (61), Expect = 7.5
Identities = 19/65 (29%), Positives = 30/65 (45%), Gaps = 8/65 (12%)
Query: 45 DSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCMPTRNSYDTSGSLSHADTRATSA 104
D E+RD V SF + ++ P + W+DG +P S + G + A+ RA A
Sbjct: 178 DIEEMRDIVQ------SFGLEPIVLPDVSS--WLDGHLPDNFSPTSMGGTTLAEMRALGA 229
Query: 105 KATCL 109
C+
Sbjct: 230 SIVCI 234
>IFP1_CAEEL (Q09501) Intermediate filament protein ifp-1
(Intermediate filament protein E1) (IF-E1) (Cel IF E1)
Length = 776
Score = 28.1 bits (61), Expect = 7.5
Identities = 28/101 (27%), Positives = 43/101 (41%), Gaps = 5/101 (4%)
Query: 6 PSLTRMKSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFII- 64
PS + S R S I+ H R ++T H RDSS R +IH + + I
Sbjct: 375 PSTQVIASALSRQNYSSISDHHSRTSSGIVQHTAHHAVRDSSSSRTGTTIHETITHTPIP 434
Query: 65 ---TMLLGPSLHQCGWMDGCMPTRNSYDTSGSLSHADTRAT 102
T+ + P + + + P R SY S AD+R++
Sbjct: 435 IPATLPVQP-IREYTYSRDASPIRPSYTPYQQESRADSRSS 474
>RECO_THETN (Q8RB48) DNA repair protein recO (Recombination protein
O)
Length = 247
Score = 27.7 bits (60), Expect = 9.8
Identities = 24/82 (29%), Positives = 38/82 (46%), Gaps = 8/82 (9%)
Query: 2 WRLVPSLTRMKSGNHR-PCCSYITLHMQRKLKKKKKNT-----TQHNDRDSSELRDQVSI 55
W LV + ++ + SYI+ + R L++K+KNT T+H+ L + SI
Sbjct: 73 WELVKNFENIEKDLKKFALASYISETISRVLEEKQKNTKLYFFTKHSLEAVESLNVETSI 132
Query: 56 HPSSSSFIITMLLG--PSLHQC 75
S + + LLG P L C
Sbjct: 133 FLFSYTLKLISLLGYMPVLDSC 154
>CAL0_HORSE (Q9N0V5) Calcitonin precursor
Length = 140
Score = 27.7 bits (60), Expect = 9.8
Identities = 22/64 (34%), Positives = 32/64 (49%), Gaps = 5/64 (7%)
Query: 89 DTSG-SLSHADTRATSAKATCLLSWFLLHQSLSRSLTYPQCK--VQTPKKYQVVAVGLEM 145
+T G SL + S +TC+L + Q L++ T+PQ V P K +V+A GLE
Sbjct: 70 ETGGASLDSPRAKRCSNLSTCVLGTYT--QDLNKFHTFPQTAIGVGAPGKKRVMARGLER 127
Query: 146 GSAP 149
P
Sbjct: 128 DHGP 131
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.128 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,119,244
Number of Sequences: 164201
Number of extensions: 593927
Number of successful extensions: 1342
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1342
Number of HSP's gapped (non-prelim): 9
length of query: 150
length of database: 59,974,054
effective HSP length: 100
effective length of query: 50
effective length of database: 43,553,954
effective search space: 2177697700
effective search space used: 2177697700
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0035.10


