

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0034b.2
(494 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
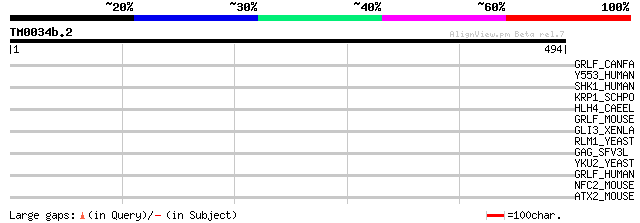
Score E
Sequences producing significant alignments: (bits) Value
GRLF_CANFA (P83509) Glucocorticoid receptor DNA binding factor 1... 34 0.99
Y553_HUMAN (Q9UKJ3) Hypothetical protein KIAA0553 33 1.3
SHK1_HUMAN (Q9Y566) SH3 and multiple ankyrin repeat domains prot... 32 2.9
KRP1_SCHPO (Q09175) Dibasic processing endoprotease precursor (E... 32 2.9
HLH4_CAEEL (P34555) Helix-loop-helix protein 4 32 2.9
GRLF_MOUSE (Q91YM2) Glucocorticoid receptor DNA binding factor 1... 32 2.9
GLI3_XENLA (Q91660) Zinc finger protein GLI3 (Neural specific DN... 32 2.9
RLM1_YEAST (Q12224) Transcription factor RLM1 32 4.9
GAG_SFV3L (P27400) Gag polyprotein (Core polyprotein) 32 4.9
YKU2_YEAST (P36042) Hypothetical 21.2 kDa protein in TOR2-MNN4 i... 31 6.4
GRLF_HUMAN (Q9NRY4) Glucocorticoid receptor DNA binding factor 1... 31 6.4
NFC2_MOUSE (Q60591) Nuclear factor of activated T-cells, cytopla... 31 8.4
ATX2_MOUSE (O70305) Ataxin-2 (Spinocerebellar ataxia type 2 prot... 31 8.4
>GRLF_CANFA (P83509) Glucocorticoid receptor DNA binding factor 1 (Rho
GAP p190A) (p190-A)
Length = 1500
Score = 33.9 bits (76), Expect = 0.99
Identities = 25/71 (35%), Positives = 35/71 (49%), Gaps = 12/71 (16%)
Query: 66 TSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSP------ 119
T I LF++ F FHNRPI + P AT + + S +P ++T S SP
Sbjct: 1424 TIIELFIQQCPFFFHNRPI---SEPPGATPSSPSAVASTVPFLTSTPVTSQPSPPQSPPP 1480
Query: 120 ---SPIKTLLP 127
SP++ LLP
Sbjct: 1481 TPQSPMQALLP 1491
>Y553_HUMAN (Q9UKJ3) Hypothetical protein KIAA0553
Length = 1089
Score = 33.5 bits (75), Expect = 1.3
Identities = 36/139 (25%), Positives = 61/139 (42%), Gaps = 28/139 (20%)
Query: 52 LYPPHVAAPRDASVTSISLF---LRNRH-------FPFHNRPIVRSNSPLHATHLPKTIM 101
L P H+ P AS TSI+ + H H P + + +H H+P+ +
Sbjct: 943 LQPIHIQQPATASATSITTVQHAILQHHAAAAAAAIGIHPHPHPQPLAQVH--HIPQPHL 1000
Query: 102 T----SNLPHASNTRHPSSFSPSPIKTLLP----HSHPFL------RPLFTSLIAYQTPS 147
T S+L H+ HP++F S ++P H PF L+ +L+A + +
Sbjct: 1001 TPISLSHLTHSIIPGHPATFLASHPIHIIPASAIHPGPFTFHPVPHAALYPTLLAPRPAA 1060
Query: 148 AHFRSGAVAYHPFRYPLFT 166
A + A+ HP +P+F+
Sbjct: 1061 A--AATALHLHPLLHPIFS 1077
>SHK1_HUMAN (Q9Y566) SH3 and multiple ankyrin repeat domains protein 1
(Shank1) (Somatostatin receptor interacting protein)
(SSTR interacting protein) (SSTRIP)
Length = 2161
Score = 32.3 bits (72), Expect = 2.9
Identities = 20/53 (37%), Positives = 25/53 (46%), Gaps = 11/53 (20%)
Query: 200 PTSPNHSSSSSGSD-----------PDATAAGGLRLEAPEIPPYRLKTYATLP 241
PTSP S + +G+D P A AGG PE+PP L T + LP
Sbjct: 1782 PTSPTVSVTGAGTDGLLALRACSGPPTAGVAGGPVAVEPEVPPVPLPTASCLP 1834
>KRP1_SCHPO (Q09175) Dibasic processing endoprotease precursor (EC
3.4.21.-) (KEX2-related protease)
Length = 709
Score = 32.3 bits (72), Expect = 2.9
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 5/65 (7%)
Query: 81 NRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSL 140
N + S++ L T T T S T P+S S PI T+LP + P L P + +
Sbjct: 615 NSDLTNSSTLLSPTSTSFTSYT-----VSATATPTSTSHIPIPTVLPPTQPVLEPSYREI 669
Query: 141 IAYQT 145
+A+ T
Sbjct: 670 VAFIT 674
>HLH4_CAEEL (P34555) Helix-loop-helix protein 4
Length = 205
Score = 32.3 bits (72), Expect = 2.9
Identities = 18/51 (35%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Query: 212 SDPDATAAGGLRLEAPEIPPYRLKTYATLPLPVQIAPTHEAIDQPSIFQNS 262
S PD + +R AP +P R +TY P+ AP H +D S F +S
Sbjct: 90 SKPDTVISQPIRPLAPVLP--RHETYLPAPIAATCAPDHSLVDYRSTFASS 138
>GRLF_MOUSE (Q91YM2) Glucocorticoid receptor DNA binding factor 1
(Fragment)
Length = 1337
Score = 32.3 bits (72), Expect = 2.9
Identities = 23/74 (31%), Positives = 38/74 (51%), Gaps = 5/74 (6%)
Query: 66 TSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTL 125
T I LF++ F F+NRPI + P A + M +P ++T P++ PSP ++
Sbjct: 1261 TIIELFIQQCPFFFYNRPI---SEPPGAAPGSPSAMAPTVPFLTST--PATSQPSPPQSP 1315
Query: 126 LPHSHPFLRPLFTS 139
P ++PL +S
Sbjct: 1316 PPTPQSPMQPLLSS 1329
>GLI3_XENLA (Q91660) Zinc finger protein GLI3 (Neural specific DNA
binding protein XGLI3) (XGLI-3)
Length = 1569
Score = 32.3 bits (72), Expect = 2.9
Identities = 37/145 (25%), Positives = 59/145 (40%), Gaps = 33/145 (22%)
Query: 34 PYSGEAFRAFP-------YFKTLLNLYPP-HVAAPRDASVTSISLFLRNRHFPFHNRPIV 85
PY G F P ++++ L+L+P H P DA R+ +H P
Sbjct: 103 PYRGTVFAMDPRNGYIDPHYRSDLDLFPAFHPPVPIDA---------RHHEGRYHYEP-- 151
Query: 86 RSNSPLHATHLPKTIMTS----NLP--HASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTS 139
+P+ H+P + +S +LP S R+P+S S SP HP++ P
Sbjct: 152 ---TPIPPLHVPTALASSPTYPDLPFFRISPHRNPASASDSPFSP----PHPYISPYMDY 204
Query: 140 LIA-YQTPSAHFRSGAVAYHPFRYP 163
+ + + +PS S A P P
Sbjct: 205 IRSLHSSPSLSMISAARGLSPTDVP 229
>RLM1_YEAST (Q12224) Transcription factor RLM1
Length = 676
Score = 31.6 bits (70), Expect = 4.9
Identities = 16/81 (19%), Positives = 40/81 (48%)
Query: 199 NPTSPNHSSSSSGSDPDATAAGGLRLEAPEIPPYRLKTYATLPLPVQIAPTHEAIDQPSI 258
N + P HSS + + P ++++ ++ P++P ++ ++ P P+ I+P + +
Sbjct: 213 NISRPYHSSMYNLNQPSSSSSSPSTMDFPKLPSFQNSSFNGRPPPISISPNKFSKPFTNA 272
Query: 259 FQNSDLIVQFVNSLGGLSNDD 279
+ +N+ G +ND+
Sbjct: 273 SSRTPKQEHKINNSGSNNNDN 293
>GAG_SFV3L (P27400) Gag polyprotein (Core polyprotein)
Length = 643
Score = 31.6 bits (70), Expect = 4.9
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 11/82 (13%)
Query: 193 LLKMASNPTSPNHSSSSSGSDPDATAAGGLRLEAPEIPPYRLKTYATLPLPVQIAPTHEA 252
L ++A NP P S S GS P P PP L + +LP PV P +
Sbjct: 227 LSRVAYNPFLPGSSDGSGGSIP----------VQPSAPPAVLPSLPSLPAPVS-QPIIQY 275
Query: 253 IDQPSIFQNSDLIVQFVNSLGG 274
+ QP + + +Q + ++ G
Sbjct: 276 VAQPPVPAPQAIPIQHIRAVTG 297
>YKU2_YEAST (P36042) Hypothetical 21.2 kDa protein in TOR2-MNN4
intergenic region
Length = 193
Score = 31.2 bits (69), Expect = 6.4
Identities = 26/66 (39%), Positives = 34/66 (51%), Gaps = 9/66 (13%)
Query: 83 PIVRSNSPLHA-----THLPKTIMTSNLPHASNTRHPSSFSPSPIKT---LLPHSHPFLR 134
P++RS+S LH HL + LP +S+ PSSFSP P + LLP L
Sbjct: 80 PLLRSSSFLHLHSSYFLHLHSSSFLL-LPASSSLLLPSSFSPLPPASSFLLLPSFSSLLL 138
Query: 135 PLFTSL 140
P F+SL
Sbjct: 139 PSFSSL 144
>GRLF_HUMAN (Q9NRY4) Glucocorticoid receptor DNA binding factor 1
(Glucocorticoid receptor repression factor 1) (GRF-1)
(Rho GAP p190A) (p190-A)
Length = 1513
Score = 31.2 bits (69), Expect = 6.4
Identities = 24/82 (29%), Positives = 39/82 (47%), Gaps = 5/82 (6%)
Query: 66 TSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTL 125
T I LF++ F F+NRPI P A + + S +P ++T P + PSP ++
Sbjct: 1423 TIIELFIQQCPFFFYNRPI---TEPPGARPSSPSAVASTVPFLTST--PVTSQPSPPQSP 1477
Query: 126 LPHSHPFLRPLFTSLIAYQTPS 147
P ++PL S + P+
Sbjct: 1478 PPTPQSPMQPLLPSPLHPHRPT 1499
>NFC2_MOUSE (Q60591) Nuclear factor of activated T-cells,
cytoplasmic 2 (T cell transcription factor NFAT1) (NFAT
pre-existing subunit) (NF-ATp)
Length = 1064
Score = 30.8 bits (68), Expect = 8.4
Identities = 27/115 (23%), Positives = 46/115 (39%), Gaps = 3/115 (2%)
Query: 203 PNHSSSSSGSDPDATAAGGLRLEAPEIPPYRLKTYATLPLPVQIAPTHEAIDQPSIFQNS 262
P SSS G+ + A L P P R ++ + P P +AP ++I P+ + +
Sbjct: 243 PASRSSSPGAKRRHSCAEALVAPLPAASPQRSRSPSPQPSP-HVAPQDDSI--PAGYPPT 299
Query: 263 DLIVQFVNSLGGLSNDDEFSAKLRIWICSPGDRPWLHKPDGDVKRSHFFFAYEYM 317
+++L L+ D +IW SP P P H + E++
Sbjct: 300 AGSAVLMDALNTLATDSPCGIPSKIWKTSPDPTPVSTAPSKAGLARHIYPTVEFL 354
>ATX2_MOUSE (O70305) Ataxin-2 (Spinocerebellar ataxia type 2 protein
homolog)
Length = 1285
Score = 30.8 bits (68), Expect = 8.4
Identities = 18/53 (33%), Positives = 27/53 (49%), Gaps = 1/53 (1%)
Query: 108 ASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTS-LIAYQTPSAHFRSGAVAYHP 159
ASNT+ P S P+ +T+ ++P +T+ P AH +SG V HP
Sbjct: 1181 ASNTQSPQSSFPAAQQTVFTIHPSHVQPAYTTPPHMAHVPQAHVQSGMVPSHP 1233
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,195,790
Number of Sequences: 164201
Number of extensions: 2799236
Number of successful extensions: 6163
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 6156
Number of HSP's gapped (non-prelim): 19
length of query: 494
length of database: 59,974,054
effective HSP length: 114
effective length of query: 380
effective length of database: 41,255,140
effective search space: 15676953200
effective search space used: 15676953200
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0034b.2


