

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0030.7
(504 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
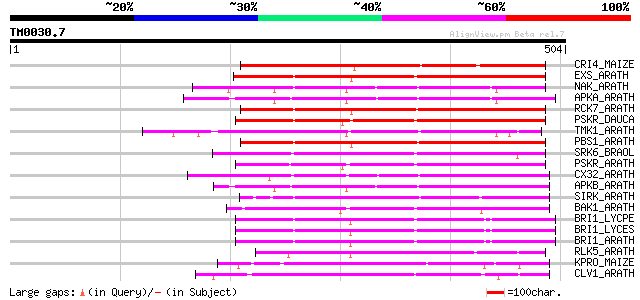
Score E
Sequences producing significant alignments: (bits) Value
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 219 2e-56
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 214 5e-55
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 210 8e-54
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 209 1e-53
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 207 4e-53
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 206 9e-53
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 205 2e-52
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 204 6e-52
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 199 2e-50
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 194 3e-49
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 192 1e-48
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 192 1e-48
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 191 3e-48
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 191 5e-48
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 186 2e-46
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 186 2e-46
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 183 8e-46
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 179 2e-44
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 173 8e-43
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 167 6e-41
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 219 bits (557), Expect = 2e-56
Identities = 116/281 (41%), Positives = 180/281 (63%), Gaps = 7/281 (2%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRA-KREHFESLRTEFSSEVELLA 268
++ +AT FSE Q+G+G F V+K L DG VVAVKRA K + EF +E++LL+
Sbjct: 497 ELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKKSSKEFHNELDLLS 556
Query: 269 KIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKI---LDFNQRLEIAIDVA 325
+++H +L+ LLG+ + G+ER+L+ E++ +G+L +HL G + L++ +R+ IA+ A
Sbjct: 557 RLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRLNWARRVTIAVQAA 616
Query: 326 HGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTV 385
G+ YLH YA +IHRD+KSSNIL+ E A+VADFG + LGP + T +S GT+
Sbjct: 617 RGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADSG-TPLSELPAGTL 675
Query: 386 GYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSV 445
GYLDPEY + H LT KSDVYSFG++LLEIL+GR+ ++++ +E + WA G +
Sbjct: 676 GYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQ--FEEGNIVEWAVPLIKAGDI 733
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
++DP++ + + L K+ ++ +C DRP+M V
Sbjct: 734 FAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKV 774
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 214 bits (544), Expect = 5e-55
Identities = 119/286 (41%), Positives = 174/286 (60%), Gaps = 6/286 (2%)
Query: 204 LNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSE 263
L + L +V AT +FS+ IG GGFGTVYKA L VAVK+ + R EF +E
Sbjct: 903 LKVRLGDIVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNR-EFMAE 961
Query: 264 VELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG--KILDFNQRLEIA 321
+E L K+ H NLV LLG+ E++L+ EY+ NG+L L G ++LD+++RL+IA
Sbjct: 962 METLGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIA 1021
Query: 322 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKV 381
+ A GL +LH IIHRD+K+SNILL KVADFG ARL ++ ++H+ST +
Sbjct: 1022 VGAARGLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARL--ISACESHVSTVI 1079
Query: 382 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELK-KAADERVTLRWAFRKY 440
GT GY+ PEY ++ + T K DVYSFG++LLE++TG+ P K ++ + WA +K
Sbjct: 1080 AGTFGYIPPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVGWAIQKI 1139
Query: 441 NEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
N+G V+++DPL+ + +++L ++ C A RPNM V
Sbjct: 1140 NQGKAVDVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDV 1185
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 210 bits (534), Expect = 8e-54
Identities = 127/339 (37%), Positives = 198/339 (57%), Gaps = 23/339 (6%)
Query: 167 DASVSASVSGKIPASPLRVPSSPSRFSMSPK-----LSRLHTLNLNLSQVVRATRNFSET 221
D + S +S K + SS + FS P+ L + N +LS++ ATRNF
Sbjct: 12 DIASSTWLSSKFLSRDGSKGSSTASFSYMPRTEGEILQNANLKNFSLSELKSATRNFRPD 71
Query: 222 LQIGQGGFGTVYKANLED----------GLVVAVKRAKREHFESLRTEFSSEVELLAKID 271
+G+GGFG V+K +++ G+V+AVKR +E F+ R E+ +E+ L ++D
Sbjct: 72 SVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGFQGHR-EWLAEINYLGQLD 130
Query: 272 HRNLVKLLGFIDKGNERILITEYVPNGTLREHL--DGLRGKILDFNQRLEIAIDVAHGLT 329
H NLVKL+G+ + R+L+ E++ G+L HL G + L +N R+ +A+ A GL
Sbjct: 131 HPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPLSWNTRVRMALGAARGLA 190
Query: 330 YLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLD 389
+LH A+ Q+I+RD K+SNILL + AK++DFG AR GP+ GD +H+ST+V GT GY
Sbjct: 191 FLH-NAQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPM-GDNSHVSTRVMGTQGYAA 248
Query: 390 PEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY--NEGSVVE 447
PEY+ T L+ KSDVYSFG++LLE+L+GRR ++ + E + WA R Y N+ ++
Sbjct: 249 PEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHNLVDWA-RPYLTNKRRLLR 307
Query: 448 LMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
+MDP ++ + +K+ L+ C + RP M +
Sbjct: 308 VMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEI 346
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 209 bits (533), Expect = 1e-53
Identities = 127/352 (36%), Positives = 205/352 (58%), Gaps = 24/352 (6%)
Query: 159 SMDSATSFDASVSASVSGKIPASPLRV-PSSPSRFSMSPKLSRLHTLNLNLSQVVRATRN 217
S ++T +DA S+ K + +R P + SP L + + +++ ATRN
Sbjct: 13 SSGASTKYDAKDIGSLGSKASSVSVRPSPRTEGEILQSPNLK-----SFSFAELKSATRN 67
Query: 218 FSETLQIGQGGFGTVYKANLED----------GLVVAVKRAKREHFESLRTEFSSEVELL 267
F +G+GGFG V+K +++ GLV+AVK+ ++ ++ E+ +EV L
Sbjct: 68 FRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQDGWQG-HQEWLAEVNYL 126
Query: 268 AKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHL--DGLRGKILDFNQRLEIAIDVA 325
+ HR+LVKL+G+ + R+L+ E++P G+L HL GL + L + RL++A+ A
Sbjct: 127 GQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPLSWKLRLKVALGAA 186
Query: 326 HGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTV 385
GL +LH +E ++I+RD K+SNILL AK++DFG A+ GP+ GD++H+ST+V GT
Sbjct: 187 KGLAFLHS-SETRVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPI-GDKSHVSTRVMGTH 244
Query: 386 GYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY--NEG 443
GY PEY+ T LT KSDVYSFG++LLE+L+GRR V+ + + ER + WA + Y N+
Sbjct: 245 GYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGERNLVEWA-KPYLVNKR 303
Query: 444 SVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRA 495
+ ++D +++ + + K+ LS +C RPNM V L I++
Sbjct: 304 KIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHLEHIQS 355
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 207 bits (528), Expect = 4e-53
Identities = 117/281 (41%), Positives = 175/281 (61%), Gaps = 6/281 (2%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLED-GLVVAVKRAKREHFESLRTEFSSEVELLA 268
++ AT NF +G+GGFG V+K +E VVA+K+ R + +R EF EV L+
Sbjct: 95 ELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIR-EFVVEVLTLS 153
Query: 269 KIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG--KILDFNQRLEIAIDVAH 326
DH NLVKL+GF +G++R+L+ EY+P G+L +HL L K LD+N R++IA A
Sbjct: 154 LADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKIAAGAAR 213
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH +I+RD+K SNILL E + K++DFG A++GP +GD+TH+ST+V GT G
Sbjct: 214 GLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGP-SGDKTHVSTRVMGTYG 272
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE-GSV 445
Y P+Y T QLT KSD+YSFG++LLE++TGR+ ++ K ++ + WA + + +
Sbjct: 273 YCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARPLFKDRRNF 332
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
+++DPL++ L + L +S C T RP + V
Sbjct: 333 PKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDV 373
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 206 bits (525), Expect = 9e-53
Identities = 116/285 (40%), Positives = 174/285 (60%), Gaps = 9/285 (3%)
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVE 265
L+L ++++T +F++ IG GGFG VYKA L DG VA+KR + + R EF +EVE
Sbjct: 731 LSLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDR-EFQAEVE 789
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTL----REHLDGLRGKILDFNQRLEIA 321
L++ H NLV LLG+ + N+++LI Y+ NG+L E +DG LD+ RL IA
Sbjct: 790 TLSRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPS--LDWKTRLRIA 847
Query: 322 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKV 381
A GL YLH E I+HRD+KSSNILL+++ A +ADFG ARL + TH++T +
Sbjct: 848 RGAAEGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARL--ILPYDTHVTTDL 905
Query: 382 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYN 441
GT+GY+ PEY + T K DVYSFG++LLE+LTGRRP+++ K R + W +
Sbjct: 906 VGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKT 965
Query: 442 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
E E+ DP + + +A+ ++ +L+++ +C RP + +
Sbjct: 966 EKRESEIFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQL 1010
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 205 bits (522), Expect = 2e-52
Identities = 147/384 (38%), Positives = 208/384 (53%), Gaps = 31/384 (8%)
Query: 121 VGIFAGGAL-LACCAVLCPCFYGKKRKA-----TAHAVLAKDPISMDSATSFDASV---S 171
VG GG L + +L C+Y K++K +++AV+ S S +V S
Sbjct: 485 VGSVLGGLLSIFLIGLLVFCWYKKRQKRFSGSESSNAVVVHPRHSGSDNESVKITVAGSS 544
Query: 172 ASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLS-QVVRA-TRNFSETLQIGQGGF 229
SV G L P + + + N+ +S QV+R+ T NFS +G GGF
Sbjct: 545 VSVGGISDTYTL-----PGTSEVGDNIQMVEAGNMLISIQVLRSVTNNFSSDNILGSGGF 599
Query: 230 GTVYKANLEDGLVVAVKRAKREHFESLR-TEFSSEVELLAKIDHRNLVKLLGFIDKGNER 288
G VYK L DG +AVKR + EF SE+ +L K+ HR+LV LLG+ GNE+
Sbjct: 600 GVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAVLTKVRHRHLVTLLGYCLDGNEK 659
Query: 289 ILITEYVPNGTLREHL-----DGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRD 343
+L+ EY+P GTL HL +GL K L + QRL +A+DVA G+ YLH A + IHRD
Sbjct: 660 LLVYEYMPQGTLSRHLFEWSEEGL--KPLLWKQRLTLALDVARGVEYLHGLAHQSFIHRD 717
Query: 344 VKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSD 403
+K SNILL + MRAKVADFG RL P + I T++ GT GYL PEY T ++T K D
Sbjct: 718 LKPSNILLGDDMRAKVADFGLVRLAPEG--KGSIETRIAGTFGYLAPEYAVTGRVTTKVD 775
Query: 404 VYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY--NEGSVVELMDPL--MEEAVNA 459
VYSFG++L+E++TGR+ ++ + + + W R Y E S + +D ++E A
Sbjct: 776 VYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYINKEASFKKAIDTTIDLDEETLA 835
Query: 460 DVLMKMLDLSFQCAAPIRTDRPNM 483
V + +L+ C A RP+M
Sbjct: 836 SV-HTVAELAGHCCAREPYQRPDM 858
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 204 bits (518), Expect = 6e-52
Identities = 116/281 (41%), Positives = 170/281 (60%), Gaps = 6/281 (2%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLED-GLVVAVKRAKREHFESLRTEFSSEVELLA 268
++ AT NF +G+GGFG VYK L+ G VVAVK+ R + R EF EV +L+
Sbjct: 78 ELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNR-EFLVEVLMLS 136
Query: 269 KIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG--KILDFNQRLEIAIDVAH 326
+ H NLV L+G+ G++R+L+ E++P G+L +HL L + LD+N R++IA A
Sbjct: 137 LLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAGAAK 196
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL +LH A +I+RD KSSNILL E K++DFG A+LGP GD++H+ST+V GT G
Sbjct: 197 GLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPT-GDKSHVSTRVMGTYG 255
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE-GSV 445
Y PEY T QLT KSDVYSFG++ LE++TGR+ ++ + E+ + WA +N+
Sbjct: 256 YCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFNDRRKF 315
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
++L DP ++ L + L ++ C RP + V
Sbjct: 316 IKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADV 356
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 199 bits (505), Expect = 2e-50
Identities = 119/311 (38%), Positives = 183/311 (58%), Gaps = 12/311 (3%)
Query: 185 VPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVA 244
V SS FS K L + + VV+AT NFS ++GQGGFG VYK L DG +A
Sbjct: 495 VLSSKREFSGEYKFEELELPLIEMETVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIA 554
Query: 245 VKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHL 304
VKR + + EF +EV L+A++ H NLV++LG +G+E++LI EY+ N +L +L
Sbjct: 555 VKRLSKTSVQGT-DEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYL 613
Query: 305 DG-LRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFG 363
G R L++N+R +I VA GL YLH + +IIHRD+K SNILL ++M K++DFG
Sbjct: 614 FGKTRRSKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFG 673
Query: 364 FARLGPVNGDQTHIST-KVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVE 422
AR+ D+T +T KV GT GY+ PEY + KSDV+SFG+++LEI++G++
Sbjct: 674 MARI--FERDETEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRG 731
Query: 423 LKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNA-------DVLMKMLDLSFQCAAP 475
E L + + ++ EG +E++DP++ +++++ ++K + + C
Sbjct: 732 FYNLDYENDLLSYVWSRWKEGRALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQE 791
Query: 476 IRTDRPNMKSV 486
+ RP M SV
Sbjct: 792 LAEHRPAMSSV 802
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 194 bits (494), Expect = 3e-49
Identities = 113/285 (39%), Positives = 167/285 (57%), Gaps = 9/285 (3%)
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVE 265
L+ ++ +T +F + IG GGFG VYKA L DG VA+K+ + + + EF +EVE
Sbjct: 722 LSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQ-IEREFEAEVE 780
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTL----REHLDGLRGKILDFNQRLEIA 321
L++ H NLV L GF N+R+LI Y+ NG+L E DG +L + RL IA
Sbjct: 781 TLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDG--PALLKWKTRLRIA 838
Query: 322 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKV 381
A GL YLH + I+HRD+KSSNILL E+ + +ADFG ARL ++ +TH+ST +
Sbjct: 839 QGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARL--MSPYETHVSTDL 896
Query: 382 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYN 441
GT+GY+ PEY + T K DVYSFG++LLE+LT +RPV++ K R + W + +
Sbjct: 897 VGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKH 956
Query: 442 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
E E+ DPL+ N + ++L+++ C + RP + +
Sbjct: 957 ESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQL 1001
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 192 bits (489), Expect = 1e-48
Identities = 118/341 (34%), Positives = 186/341 (53%), Gaps = 16/341 (4%)
Query: 162 SATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNL-NLSQVVRATRNFSE 220
+ T F ++ A+ + + S S S KL L + N + AT+NF
Sbjct: 29 NGTEFSSTTGATTNSSVGQQSQFSDISTGIISDSGKLLESPNLKVYNFLDLKTATKNFKP 88
Query: 221 TLQIGQGGFGTVYK----------ANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKI 270
+GQGGFG VY+ + + G++VA+KR E + E+ SEV L +
Sbjct: 89 DSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQGF-AEWRSEVNFLGML 147
Query: 271 DHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTY 330
HRNLVKLLG+ + E +L+ E++P G+L HL R ++ R++I I A GL +
Sbjct: 148 SHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFR-RNDPFPWDLRIKIVIGAARGLAF 206
Query: 331 LHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDP 390
LH ++++I+RD K+SNILL + AK++DFG A+LGP + +++H++T++ GT GY P
Sbjct: 207 LHSL-QREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPAD-EKSHVTTRIMGTYGYAAP 264
Query: 391 EYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKY-NEGSVVELM 449
EYM T L KSDV++FG++LLEI+TG K+ + + W + N+ V ++M
Sbjct: 265 EYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQESLVDWLRPELSNKHRVKQIM 324
Query: 450 DPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
D ++ V +M ++ C P +RP+MK V E L
Sbjct: 325 DKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVL 365
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 192 bits (489), Expect = 1e-48
Identities = 113/318 (35%), Positives = 182/318 (56%), Gaps = 21/318 (6%)
Query: 186 PSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLED------ 239
P + SP L + +++ ATRNF +G+GGFG+V+K +++
Sbjct: 42 PRTEGEILQSPNLK-----SFTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTAS 96
Query: 240 ----GLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYV 295
G+V+AVK+ ++ ++ E+ +EV L + H NLVKL+G+ + R+L+ E++
Sbjct: 97 KPGTGVVIAVKKLNQDGWQG-HQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFM 155
Query: 296 PNGTLREHL--DGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTE 353
P G+L HL G + L + RL++A+ A GL +LH AE +I+RD K+SNILL
Sbjct: 156 PRGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLH-NAETSVIYRDFKTSNILLDS 214
Query: 354 SMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLE 413
AK++DFG A+ GP GD++H+ST++ GT GY PEY+ T LT KSDVYS+G++LLE
Sbjct: 215 EYNAKLSDFGLAKDGPT-GDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLE 273
Query: 414 ILTGRRPVELKKAADERVTLRWAFRKY-NEGSVVELMDPLMEEAVNADVLMKMLDLSFQC 472
+L+GRR V+ + E+ + WA N+ + ++D +++ + + K+ L+ +C
Sbjct: 274 VLSGRRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRC 333
Query: 473 AAPIRTDRPNMKSVGEQL 490
RPNM V L
Sbjct: 334 LTFEIKLRPNMNEVVSHL 351
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 191 bits (486), Expect = 3e-48
Identities = 114/283 (40%), Positives = 164/283 (57%), Gaps = 8/283 (2%)
Query: 209 SQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLA 268
S+VV T NF IG+GGFG VY + +G VAVK E + + EF +EV+LL
Sbjct: 567 SEVVNITNNFERV--IGKGGFGKVYHGVI-NGEQVAVKVLSEESAQGYK-EFRAEVDLLM 622
Query: 269 KIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGL 328
++ H NL L+G+ ++ N +LI EY+ N L ++L G R IL + +RL+I++D A GL
Sbjct: 623 RVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGKRSFILSWEERLKISLDAAQGL 682
Query: 329 TYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYL 388
YLH + I+HRDVK +NILL E ++AK+ADFG +R V G IST V G++GYL
Sbjct: 683 EYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEG-SGQISTVVAGSIGYL 741
Query: 389 DPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRK-YNEGSVVE 447
DPEY T Q+ KSDVYS G++LLE++TG+ + K E+V + R G +
Sbjct: 742 DPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKT--EKVHISDHVRSILANGDIRG 799
Query: 448 LMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
++D + E + KM +++ C RP M V +L
Sbjct: 800 IVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMEL 842
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 191 bits (484), Expect = 5e-48
Identities = 112/299 (37%), Positives = 175/299 (58%), Gaps = 12/299 (4%)
Query: 198 LSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLR 257
L +L +L QV A+ NFS +G+GGFG VYK L DG +VAVKR K E +
Sbjct: 271 LGQLKRFSLRELQV--ASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERTQGGE 328
Query: 258 TEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGT----LREHLDGLRGKILD 313
+F +EVE+++ HRNL++L GF ER+L+ Y+ NG+ LRE + LD
Sbjct: 329 LQFQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPES--QPPLD 386
Query: 314 FNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGD 373
+ +R IA+ A GL YLH + + +IIHRDVK++NILL E A V DFG A+L ++
Sbjct: 387 WPKRQRIALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKL--MDYK 444
Query: 374 QTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAA--DERV 431
TH++T V+GT+G++ PEY+ T + + K+DV+ +G++LLE++TG+R +L + A D+ +
Sbjct: 445 DTHVTTAVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVM 504
Query: 432 TLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
L W E + L+D ++ + + +++ ++ C +RP M V L
Sbjct: 505 LLDWVKGLLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRML 563
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 186 bits (471), Expect = 2e-46
Identities = 113/295 (38%), Positives = 177/295 (59%), Gaps = 10/295 (3%)
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVE 265
L + ++ AT F +G GGFG VYKA L+DG VVA+K+ + R EF++E+E
Sbjct: 876 LTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDR-EFTAEME 934
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLR--GKILDFNQRLEIAID 323
+ KI HRNLV LLG+ G ER+L+ EY+ G+L + L + G L++ R +IAI
Sbjct: 935 TIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIG 994
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIS-TKVK 382
A GL +LH IIHRD+KSSN+LL E++ A+V+DFG ARL ++ TH+S + +
Sbjct: 995 AARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARL--MSAMDTHLSVSTLA 1052
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE 442
GT GY+ PEY ++ + + K DVYS+G++LLE+LTG++P + D + + W + + +
Sbjct: 1053 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNL-VGWV-KLHAK 1110
Query: 443 GSVVELMD-PLMEEAVNADV-LMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRA 495
G + ++ D L++E + ++ L++ L ++ C RP M V I+A
Sbjct: 1111 GKITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFKEIQA 1165
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 186 bits (471), Expect = 2e-46
Identities = 113/295 (38%), Positives = 177/295 (59%), Gaps = 10/295 (3%)
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVE 265
L + ++ AT F +G GGFG VYKA L+DG VVA+K+ + R EF++E+E
Sbjct: 876 LTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDR-EFTAEME 934
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLR--GKILDFNQRLEIAID 323
+ KI HRNLV LLG+ G ER+L+ EY+ G+L + L + G L++ R +IAI
Sbjct: 935 TIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIG 994
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIS-TKVK 382
A GL +LH IIHRD+KSSN+LL E++ A+V+DFG ARL ++ TH+S + +
Sbjct: 995 AARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARL--MSAMDTHLSVSTLA 1052
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE 442
GT GY+ PEY ++ + + K DVYS+G++LLE+LTG++P + D + + W + + +
Sbjct: 1053 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNL-VGWV-KLHAK 1110
Query: 443 GSVVELMD-PLMEEAVNADV-LMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRA 495
G + ++ D L++E + ++ L++ L ++ C RP M V I+A
Sbjct: 1111 GKITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFKEIQA 1165
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 183 bits (465), Expect = 8e-46
Identities = 114/295 (38%), Positives = 178/295 (59%), Gaps = 10/295 (3%)
Query: 206 LNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVE 265
L + +++AT F IG GGFG VYKA L+DG VA+K+ + R EF +E+E
Sbjct: 871 LTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQGDR-EFMAEME 929
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLR--GKILDFNQRLEIAID 323
+ KI HRNLV LLG+ G+ER+L+ E++ G+L + L + G L+++ R +IAI
Sbjct: 930 TIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIAIG 989
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHIS-TKVK 382
A GL +LH IIHRD+KSSN+LL E++ A+V+DFG ARL ++ TH+S + +
Sbjct: 990 SARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARL--MSAMDTHLSVSTLA 1047
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE 442
GT GY+ PEY ++ + + K DVYS+G++LLE+LTG+RP + D + + W +++ +
Sbjct: 1048 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSPDFGDNNL-VGWV-KQHAK 1105
Query: 443 GSVVELMDP-LMEEAVNADV-LMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRA 495
+ ++ DP LM+E ++ L++ L ++ C RP M V I+A
Sbjct: 1106 LRISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFKEIQA 1160
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 179 bits (453), Expect = 2e-44
Identities = 103/275 (37%), Positives = 165/275 (59%), Gaps = 15/275 (5%)
Query: 224 IGQGGFGTVYKANLEDGLVVAVKRAKRE--------HFESL-RTEFSSEVELLAKIDHRN 274
IG G G VYK L G VVAVK+ + +SL R F++EVE L I H++
Sbjct: 689 IGFGSSGKVYKVELRGGEVVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGTIRHKS 748
Query: 275 LVKLLGFIDKGNERILITEYVPNGTLREHLDGLR--GKILDFNQRLEIAIDVAHGLTYLH 332
+V+L G+ ++L+ EY+PNG+L + L G R G +L + +RL IA+D A GL+YLH
Sbjct: 749 IVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEGLSYLH 808
Query: 333 LYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQT-HISTKVKGTVGYLDPE 391
I+HRDVKSSNILL AKVADFG A++G ++G +T + + G+ GY+ PE
Sbjct: 809 HDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAKVGQMSGSKTPEAMSGIAGSCGYIAPE 868
Query: 392 YMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDP 451
Y+ T ++ KSD+YSFG++LLE++TG++P + ++ +W ++ + ++DP
Sbjct: 869 YVYTLRVNEKSDIYSFGVVLLELVTGKQPTD--SELGDKDMAKWVCTALDKCGLEPVIDP 926
Query: 452 LMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 486
++ ++ K++ + C +P+ +RP+M+ V
Sbjct: 927 KLDLKFKEEI-SKVIHIGLLCTSPLPLNRPSMRKV 960
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 173 bits (439), Expect = 8e-43
Identities = 116/315 (36%), Positives = 173/315 (54%), Gaps = 19/315 (6%)
Query: 189 PSRFSMSPKLSRLHTLNL---NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAV 245
PS S K + T N + ++V+ATR F +++G+G GTVYK LED VAV
Sbjct: 504 PSELWASEKGYKAMTSNFRRYSYRELVKATRKFK--VELGRGESGTVYKGVLEDDRHVAV 561
Query: 246 KRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLD 305
K K E+ + F +E+ ++ +I+H NLV++ GF +G+ R+L++EYV NG+L L
Sbjct: 562 K--KLENVRQGKEVFQAELSVIGRINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILF 619
Query: 306 GLRGKIL-DFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGF 364
G IL D+ R IA+ VA GL YLH + +IH DVK NILL ++ K+ DFG
Sbjct: 620 SEGGNILLDWEGRFNIALGVAKGLAYLHHECLEWVIHCDVKPENILLDQAFEPKITDFGL 679
Query: 365 ARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELK 424
+L G ++S V+GT+GY+ PE++ + +T K DVYS+G++LLE+LTG R EL
Sbjct: 680 VKLLNRGGSTQNVS-HVRGTLGYIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELV 738
Query: 425 KAADE-----RVTLRWAFRKYNEGSVVELMDPLMEEAVNADV----LMKMLDLSFQCAAP 475
DE R +R K EG +D ++ +N V ++ L+ C
Sbjct: 739 GGTDEVHSMLRKLVRMLSAKL-EGEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEE 797
Query: 476 IRTDRPNMKSVGEQL 490
R+ RP M+ + L
Sbjct: 798 DRSKRPTMEHAVQTL 812
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 167 bits (423), Expect = 6e-41
Identities = 109/332 (32%), Positives = 177/332 (52%), Gaps = 18/332 (5%)
Query: 169 SVSASVSGKIPASPL--RVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQ 226
+V A+++G I S ++ ++ S++ KL+ L+ V+ + E IG+
Sbjct: 644 TVIAAITGLILISVAIRQMNKKKNQKSLAWKLTAFQKLDFKSEDVLECLK---EENIIGK 700
Query: 227 GGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGN 286
GG G VY+ ++ + + VA+KR F++E++ L +I HR++V+LLG++ +
Sbjct: 701 GGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGYVANKD 760
Query: 287 ERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKS 346
+L+ EY+PNG+L E L G +G L + R +A++ A GL YLH I+HRDVKS
Sbjct: 761 TNLLLYEYMPNGSLGELLHGSKGGHLQWETRHRVAVEAAKGLCYLHHDCSPLILHRDVKS 820
Query: 347 SNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYS 406
+NILL A VADFG A+ V+G + + + G+ GY+ PEY T ++ KSDVYS
Sbjct: 821 NNILLDSDFEAHVADFGLAKF-LVDGAASECMSSIAGSYGYIAPEYAYTLKVDEKSDVYS 879
Query: 407 FGILLLEILTGRRPV-ELKKAADERVTLRWAFRKYNE-------GSVVELMDPLMEEAVN 458
FG++LLE++ G++PV E + D +RW E VV ++DP +
Sbjct: 880 FGVVLLELIAGKKPVGEFGEGVD---IVRWVRNTEEEITQPSDAAIVVAIVDPRLTGYPL 936
Query: 459 ADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
V+ + ++ C RP M+ V L
Sbjct: 937 TSVI-HVFKIAMMCVEEEAAARPTMREVVHML 967
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,553,006
Number of Sequences: 164201
Number of extensions: 2285780
Number of successful extensions: 10381
Number of sequences better than 10.0: 1673
Number of HSP's better than 10.0 without gapping: 1137
Number of HSP's successfully gapped in prelim test: 536
Number of HSP's that attempted gapping in prelim test: 6647
Number of HSP's gapped (non-prelim): 1875
length of query: 504
length of database: 59,974,054
effective HSP length: 115
effective length of query: 389
effective length of database: 41,090,939
effective search space: 15984375271
effective search space used: 15984375271
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0030.7


