

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.15
(145 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
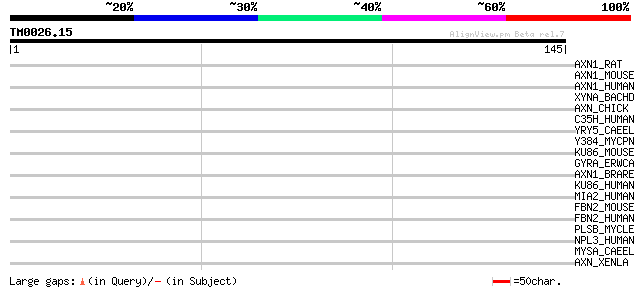
Score E
Sequences producing significant alignments: (bits) Value
AXN1_RAT (O70239) Axin 1 protein (Axis inhibition protein 1) (rA... 31 0.80
AXN1_MOUSE (O35625) Axin 1 (Axis inhibition protein 1) (Fused pr... 31 0.80
AXN1_HUMAN (O15169) Axin 1 (Axis inhibition protein 1) (hAxin) 31 1.0
XYNA_BACHD (P07528) Endo-1,4-beta-xylanase A precursor (EC 3.2.1... 30 1.4
AXN_CHICK (O42400) Axin (Axis inhibition protein) 30 1.4
C35H_HUMAN (Q9UGN4) CMRF35-H antigen precursor (CMRF35-H9) (CMRF... 29 3.0
YRY5_CAEEL (Q09355) Hypothetical protein T15H9.5 in chromosome II 29 4.0
Y384_MYCPN (P75215) Probable GTP-binding protein MG384 homolog 29 4.0
KU86_MOUSE (P27641) ATP-dependent DNA helicase II, 80 kDa subuni... 29 4.0
GYRA_ERWCA (P41513) DNA gyrase subunit A (EC 5.99.1.3) 29 4.0
AXN1_BRARE (P57094) Axin 1 (Axis inhibition protein 1) 29 4.0
KU86_HUMAN (P13010) ATP-dependent DNA helicase II, 80 kDa subuni... 28 5.2
MIA2_HUMAN (Q96PC5) Melanoma inhibitory activity protein 2 precu... 28 6.8
FBN2_MOUSE (Q61555) Fibrillin 2 precursor 28 6.8
FBN2_HUMAN (P35556) Fibrillin 2 precursor 28 6.8
PLSB_MYCLE (Q9X7B0) Glycerol-3-phosphate acyltransferase (EC 2.3... 28 8.9
NPL3_HUMAN (Q99457) Nucleosome assembly protein 1-like 3 28 8.9
MYSA_CAEEL (P12844) Myosin heavy chain A (MHC A) 28 8.9
AXN_XENLA (Q9YGY0) Axin (Axis inhibition protein) (xAxin) 28 8.9
>AXN1_RAT (O70239) Axin 1 protein (Axis inhibition protein 1)
(rAxin)
Length = 827
Score = 31.2 bits (69), Expect = 0.80
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELV-------PDDENEHYV 104
E+ +NGRV LP+ PR++ +P ++ EP+K EL+ E E +
Sbjct: 358 ESVQVNGRVPLPHIPRTYRMP--------KEIRVEPQKFAEELIHRLEAVQRTREAEEKL 409
Query: 105 DSKLMSKPMEEE 116
+ +L MEEE
Sbjct: 410 EERLKRVRMEEE 421
>AXN1_MOUSE (O35625) Axin 1 (Axis inhibition protein 1) (Fused
protein)
Length = 863
Score = 31.2 bits (69), Expect = 0.80
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELV-------PDDENEHYV 104
E+ +NGRV LP+ PR++ +P ++ EP+K EL+ E E +
Sbjct: 358 ESIQVNGRVPLPHIPRTYRMP--------KEIRVEPQKFAEELIHRLEAVQRTREAEEKL 409
Query: 105 DSKLMSKPMEEE 116
+ +L MEEE
Sbjct: 410 EERLKRVRMEEE 421
>AXN1_HUMAN (O15169) Axin 1 (Axis inhibition protein 1) (hAxin)
Length = 862
Score = 30.8 bits (68), Expect = 1.0
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELV-------PDDENEHYV 104
E+ +NGRV LP+ PR++ +P ++ EP+K EL+ E E +
Sbjct: 358 ESVQVNGRVPLPHIPRTYRVP--------KEVRVEPQKFAEELIHRLEAVQRTREAEEKL 409
Query: 105 DSKLMSKPMEEE 116
+ +L MEEE
Sbjct: 410 EERLKRVRMEEE 421
>XYNA_BACHD (P07528) Endo-1,4-beta-xylanase A precursor (EC 3.2.1.8)
(Xylanase A) (1,4-beta-D-xylan xylanohydrolase A)
Length = 396
Score = 30.4 bits (67), Expect = 1.4
Identities = 13/29 (44%), Positives = 18/29 (61%), Gaps = 1/29 (3%)
Query: 101 EHYVDSKLMSKPMEEEELPPLEGELWNWE 129
+H+ +S + M+ E L P EGE WNWE
Sbjct: 85 KHHYNSLVAENAMKPESLQPREGE-WNWE 112
>AXN_CHICK (O42400) Axin (Axis inhibition protein)
Length = 841
Score = 30.4 bits (67), Expect = 1.4
Identities = 22/72 (30%), Positives = 34/72 (46%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELV-------PDDENEHYV 104
E+ NGRV LP+ PR++ +P ++ EPEK EL+ + E E +
Sbjct: 358 ESAKANGRVPLPHIPRTYRMP--------KDIHVEPEKFAAELINRLEEVQKEREAEEKL 409
Query: 105 DSKLMSKPMEEE 116
+ +L EEE
Sbjct: 410 EERLKRVRAEEE 421
>C35H_HUMAN (Q9UGN4) CMRF35-H antigen precursor (CMRF35-H9)
(CMRF-35-H9) (Inhibitory receptor protein 60) (IRp60)
(IRC1/IRC2) (NK inhibitory receptor)
Length = 299
Score = 29.3 bits (64), Expect = 3.0
Identities = 22/77 (28%), Positives = 35/77 (44%), Gaps = 8/77 (10%)
Query: 67 RSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGEL- 125
R F+ I G H L+ + EL HY + +L+ P++E+ PP E E+
Sbjct: 202 RMFQKWIKAGDHSELSQNPKQAATQSEL-------HYANLELLMWPLQEKPAPPREVEVE 254
Query: 126 WNWEDPPDDELRARSFV 142
++ P +EL S V
Sbjct: 255 YSTVASPREELHYASVV 271
>YRY5_CAEEL (Q09355) Hypothetical protein T15H9.5 in chromosome II
Length = 173
Score = 28.9 bits (63), Expect = 4.0
Identities = 22/71 (30%), Positives = 34/71 (46%), Gaps = 9/71 (12%)
Query: 63 PYAPRSFELPI-VVGSHQLLNMKSEPEKDPLELVPDD--------ENEHYVDSKLMSKPM 113
P +P + PI V+ +L + K E +K+P E V D NE V + ++K
Sbjct: 64 PVSPEKKKSPIKVLSEKKLKSKKKEEDKEPDEKVEKDVKKEVKADNNEPLVKNLKIAKKE 123
Query: 114 EEEELPPLEGE 124
+EEE P + E
Sbjct: 124 QEEENPKTDLE 134
>Y384_MYCPN (P75215) Probable GTP-binding protein MG384 homolog
Length = 433
Score = 28.9 bits (63), Expect = 4.0
Identities = 12/56 (21%), Positives = 25/56 (44%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSK 107
+ + ++ + + F LP+ + H + SE + DPL + D +V+ K
Sbjct: 323 QVFALHQKTLAQFGANKFHLPMEMEKHYVFEQASETDHDPLNIERDALGRWHVECK 378
>KU86_MOUSE (P27641) ATP-dependent DNA helicase II, 80 kDa subunit
(Ku autoantigen protein p86 homolog) (Ku80) (CTC box
binding factor 85 kDa subunit) (CTCBF) (CTC85) (Nuclear
factor IV) (DNA-repair protein XRCC5)
Length = 732
Score = 28.9 bits (63), Expect = 4.0
Identities = 20/84 (23%), Positives = 36/84 (42%), Gaps = 4/84 (4%)
Query: 55 LMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPE--KD--PLELVPDDENEHYVDSKLMS 110
L N + C P + + ++ S L+ E + +D P +P+ E + L
Sbjct: 438 LKNNKKCTPTEAQLSAIDDLIDSMSLVKKNEEEDIVEDLFPASKIPNPEFQRLYQCLLHR 497
Query: 111 KPMEEEELPPLEGELWNWEDPPDD 134
+E LPP++ + N DPP +
Sbjct: 498 ALHLQERLPPIQQHILNIWDPPTE 521
>GYRA_ERWCA (P41513) DNA gyrase subunit A (EC 5.99.1.3)
Length = 878
Score = 28.9 bits (63), Expect = 4.0
Identities = 16/44 (36%), Positives = 27/44 (61%), Gaps = 3/44 (6%)
Query: 94 LVPDDENEHYVDSKLMSKPMEEEELP---PLEGELWNWEDPPDD 134
L+ E+EH V + +++P+E+EEL +EGE+ +D DD
Sbjct: 823 LIRTAEDEHVVGLQRVAEPVEDEELDGVVKVEGEVAEDDDAIDD 866
>AXN1_BRARE (P57094) Axin 1 (Axis inhibition protein 1)
Length = 835
Score = 28.9 bits (63), Expect = 4.0
Identities = 21/72 (29%), Positives = 35/72 (48%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVP-------DDENEHYV 104
E+ +NGRV LP+ PR+ +P ++ EPEK EL+ + E + +
Sbjct: 361 ESAKVNGRVPLPHIPRTNRIP--------KDIHVEPEKFAAELISRLEGVLREREAQEKL 412
Query: 105 DSKLMSKPMEEE 116
+ +L +EEE
Sbjct: 413 EERLKRVRLEEE 424
>KU86_HUMAN (P13010) ATP-dependent DNA helicase II, 80 kDa subunit
(Lupus Ku autoantigen protein p86) (Ku86) (Ku80) (86 kDa
subunit of Ku antigen) (Thyroid-lupus autoantigen)
(TLAA) (CTC box binding factor 85 kDa subunit) (CTCBF)
(CTC85) (Nuclear factor I
Length = 731
Score = 28.5 bits (62), Expect = 5.2
Identities = 18/76 (23%), Positives = 30/76 (38%), Gaps = 7/76 (9%)
Query: 64 YAPRSFELPIVVGSHQLLNMKSEPEKD-------PLELVPDDENEHYVDSKLMSKPMEEE 116
YAP +L V +++ + EK P +P+ + L E
Sbjct: 443 YAPTEAQLNAVDALIDSMSLAKKDEKTDTLEDLFPTTKIPNPRFQRLFQCLLHRALHPRE 502
Query: 117 ELPPLEGELWNWEDPP 132
LPP++ +WN +PP
Sbjct: 503 PLPPIQQHIWNMLNPP 518
>MIA2_HUMAN (Q96PC5) Melanoma inhibitory activity protein 2
precursor
Length = 541
Score = 28.1 bits (61), Expect = 6.8
Identities = 16/53 (30%), Positives = 32/53 (60%), Gaps = 2/53 (3%)
Query: 81 LNMKSEPEKDP-LELVPDDENEHYVDSKLMSKPMEEEE-LPPLEGELWNWEDP 131
L + +PEK+ +E + E E +D K +S+ +E + +P L+ L+N+++P
Sbjct: 414 LTSELDPEKEQEIETIKIIETEDQIDKKPVSEKTDESDTIPYLKKFLYNFDNP 466
>FBN2_MOUSE (Q61555) Fibrillin 2 precursor
Length = 2907
Score = 28.1 bits (61), Expect = 6.8
Identities = 19/64 (29%), Positives = 26/64 (39%), Gaps = 5/64 (7%)
Query: 42 RSFDQRGRWCETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENE 101
R FD R +C T NG+ +P A + + M E DP EL P D+
Sbjct: 2088 RCFDTRQSFCFTNFENGKCSVPKAFNTTKAKCCCS-----KMPGEGWGDPCELCPKDDEV 2142
Query: 102 HYVD 105
+ D
Sbjct: 2143 AFQD 2146
>FBN2_HUMAN (P35556) Fibrillin 2 precursor
Length = 2911
Score = 28.1 bits (61), Expect = 6.8
Identities = 19/64 (29%), Positives = 26/64 (39%), Gaps = 5/64 (7%)
Query: 42 RSFDQRGRWCETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENE 101
R FD R +C T NG+ +P A + + M E DP EL P D+
Sbjct: 2094 RCFDTRQSFCFTNFENGKCSVPKAFNTTKAKCCCS-----KMPGEGWGDPCELCPKDDEV 2148
Query: 102 HYVD 105
+ D
Sbjct: 2149 AFQD 2152
>PLSB_MYCLE (Q9X7B0) Glycerol-3-phosphate acyltransferase (EC
2.3.1.15) (GPAT)
Length = 775
Score = 27.7 bits (60), Expect = 8.9
Identities = 16/55 (29%), Positives = 27/55 (49%), Gaps = 2/55 (3%)
Query: 67 RSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPL 121
R FE I +Q+ M++ E P L+ + Y+D ++ M+E LPP+
Sbjct: 237 RGFEPEIDYDEYQVAAMRAALEAHPAVLL--FSHRSYIDGAVVPVAMQENRLPPV 289
>NPL3_HUMAN (Q99457) Nucleosome assembly protein 1-like 3
Length = 506
Score = 27.7 bits (60), Expect = 8.9
Identities = 17/53 (32%), Positives = 27/53 (50%), Gaps = 3/53 (5%)
Query: 72 PIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGE 124
P+ Q++N + EP ++ E +DE E D ++ E+PPLEGE
Sbjct: 142 PLYDRRFQIINAEYEPTEEECEWNSEDE-EFSSDEEVQDNT--PSEMPPLEGE 191
>MYSA_CAEEL (P12844) Myosin heavy chain A (MHC A)
Length = 1969
Score = 27.7 bits (60), Expect = 8.9
Identities = 18/59 (30%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Query: 79 QLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDELR 137
Q L KSE EK L+ +E++H DS++ S+ E+ L +E + + D++ R
Sbjct: 1224 QKLKAKSEAEKSKLQR-DLEESQHATDSEVRSRQDLEKALKTIEVQYSELQTKADEQSR 1281
>AXN_XENLA (Q9YGY0) Axin (Axis inhibition protein) (xAxin)
Length = 842
Score = 27.7 bits (60), Expect = 8.9
Identities = 20/67 (29%), Positives = 31/67 (45%), Gaps = 15/67 (22%)
Query: 57 NGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVP-------DDENEHYVDSKLM 109
NGR LP+ PR++ +P ++ +PEK EL+ D E E ++ +L
Sbjct: 363 NGRGPLPHIPRTYHMP--------KDIHVDPEKFAAELISRLEGVLRDREAEQKLEERLK 414
Query: 110 SKPMEEE 116
EEE
Sbjct: 415 RVRAEEE 421
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.137 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,825,640
Number of Sequences: 164201
Number of extensions: 838691
Number of successful extensions: 2160
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 2157
Number of HSP's gapped (non-prelim): 21
length of query: 145
length of database: 59,974,054
effective HSP length: 100
effective length of query: 45
effective length of database: 43,553,954
effective search space: 1959927930
effective search space used: 1959927930
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0026.15


